Classification rationale
1
2
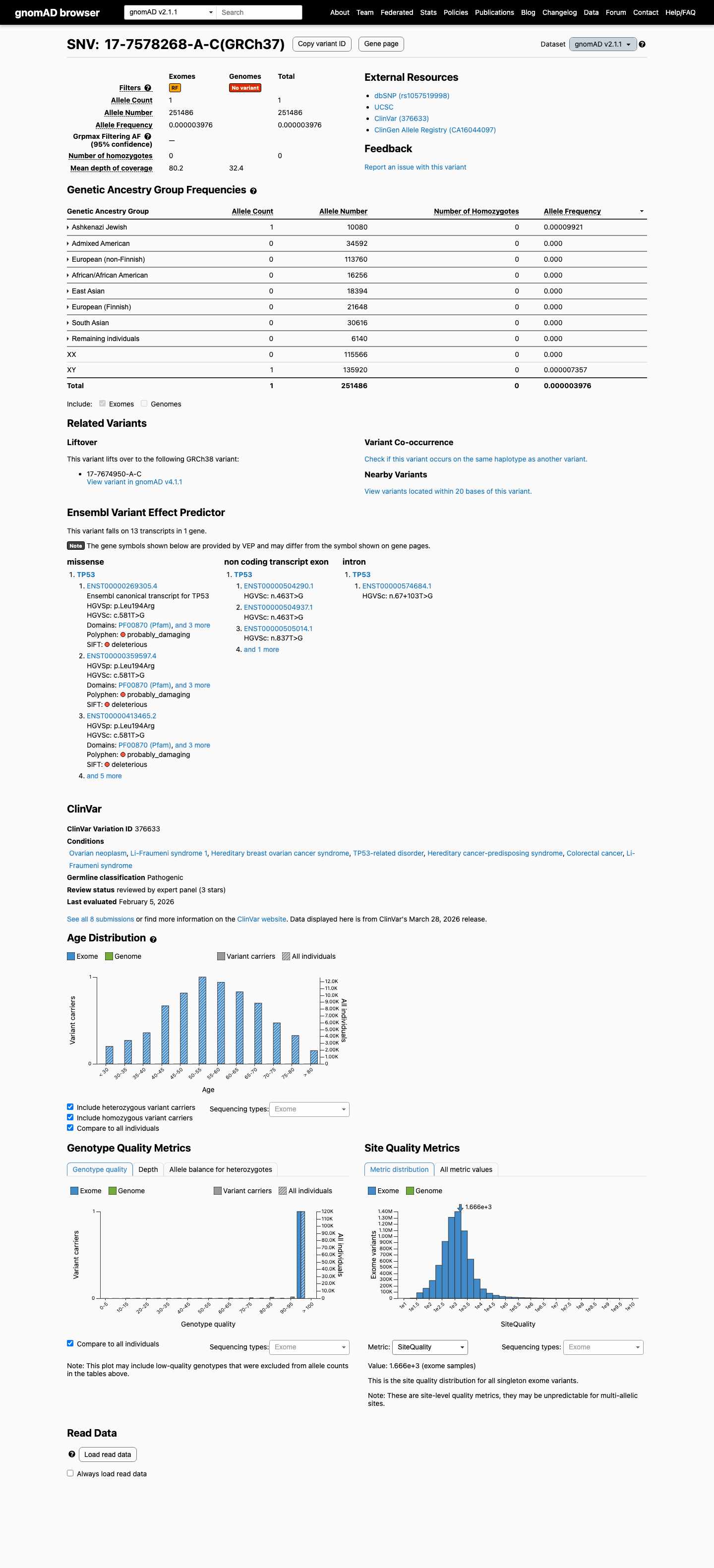
This variant is rare in population databases, with 1/1614188 alleles in gnomAD v4.1 (AF 6.20e-07) and 1/251486 alleles in gnomAD v2.1 (AF 3.98e-06), which is below the TP53 PM2 threshold of 0.00003.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗3
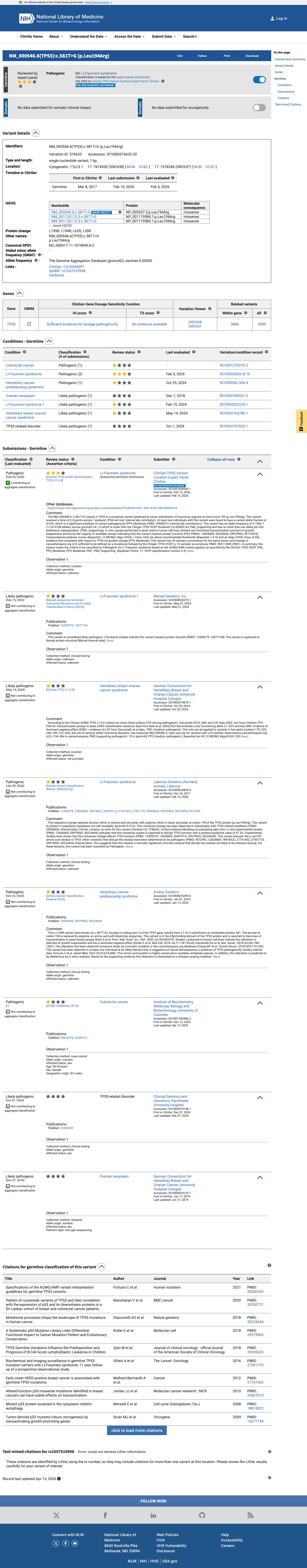
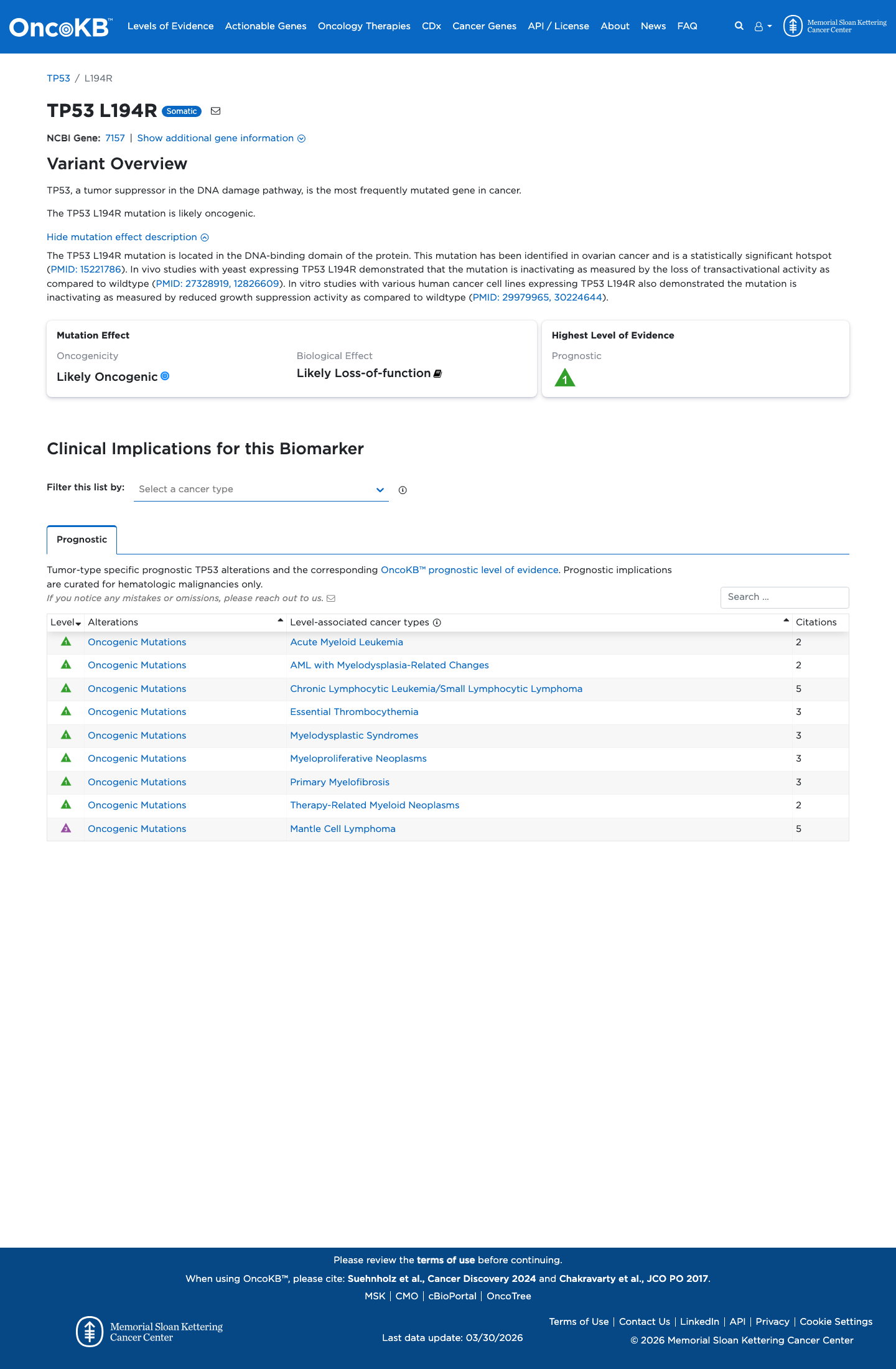
In the TP53 VCEP functional worksheet, p.Leu194Arg is listed as non-functional in the Kato assay and as loss-of-function in the other eligible assay categories reviewed, supporting PS3.
PMID:12826609 ↗ PMID:29979965 ↗ PMID:30224644 ↗4
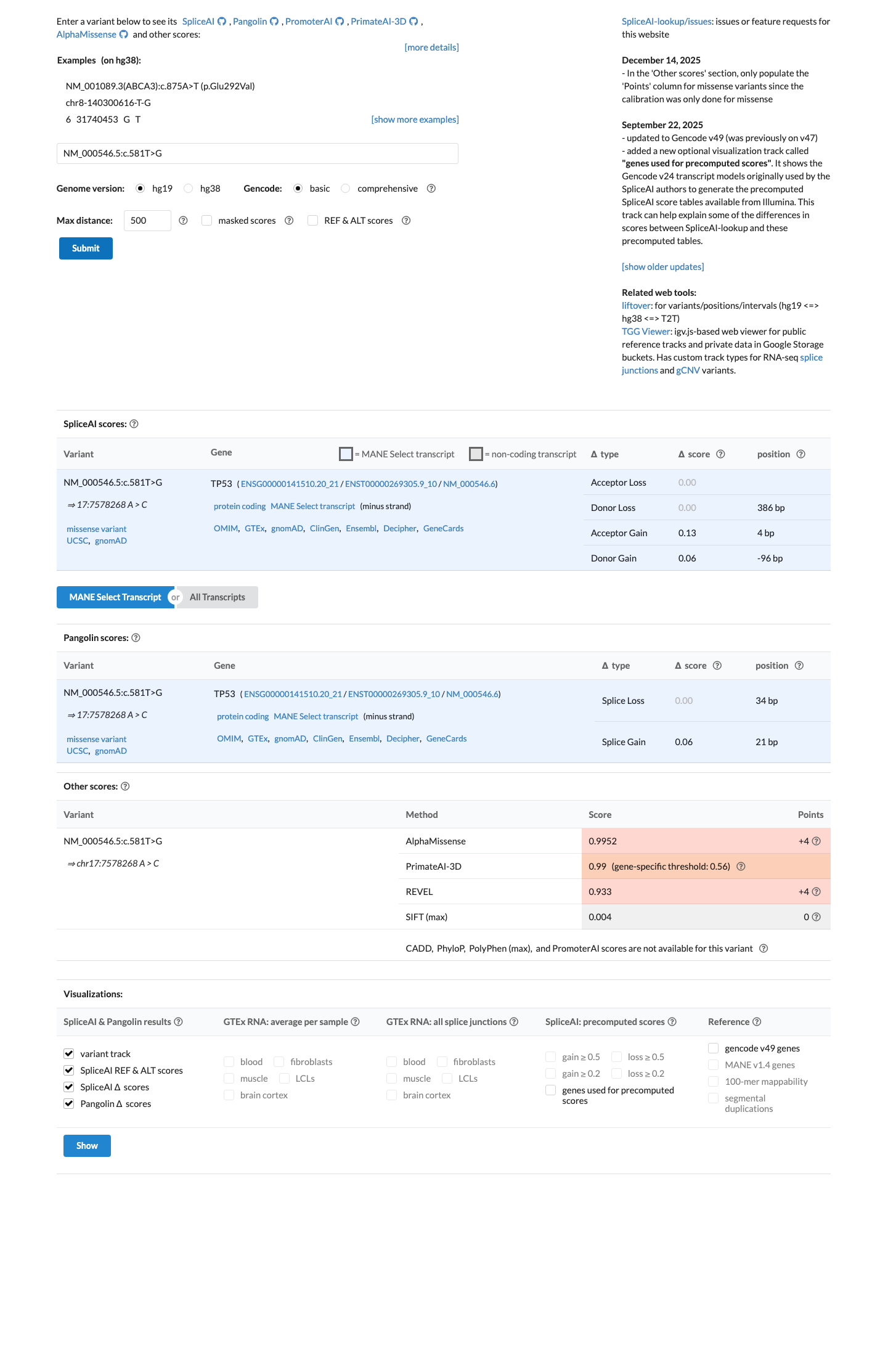
In the TP53 VCEP bioinformatic worksheet, c.581T>G is assigned PP3_moderate; BayesDel is 0.582962, REVEL is 0.933, and SpliceAI shows no significant predicted splice effect (max delta score 0.13), supporting a damaging missense effect without evidence for a splice-based mechanism.
spliceai ↗