Classification rationale
1
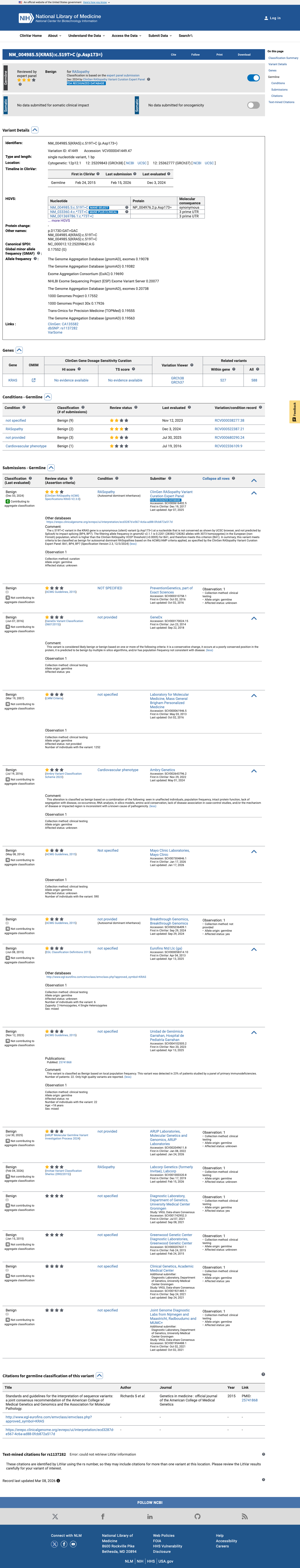
The KRAS c.519T>C (p.Asp173=; p.D173=) variant has been reported in ClinVar as Benign, including by the ClinGen RASopathy Variant Curation Expert Panel.
clinvar ↗2
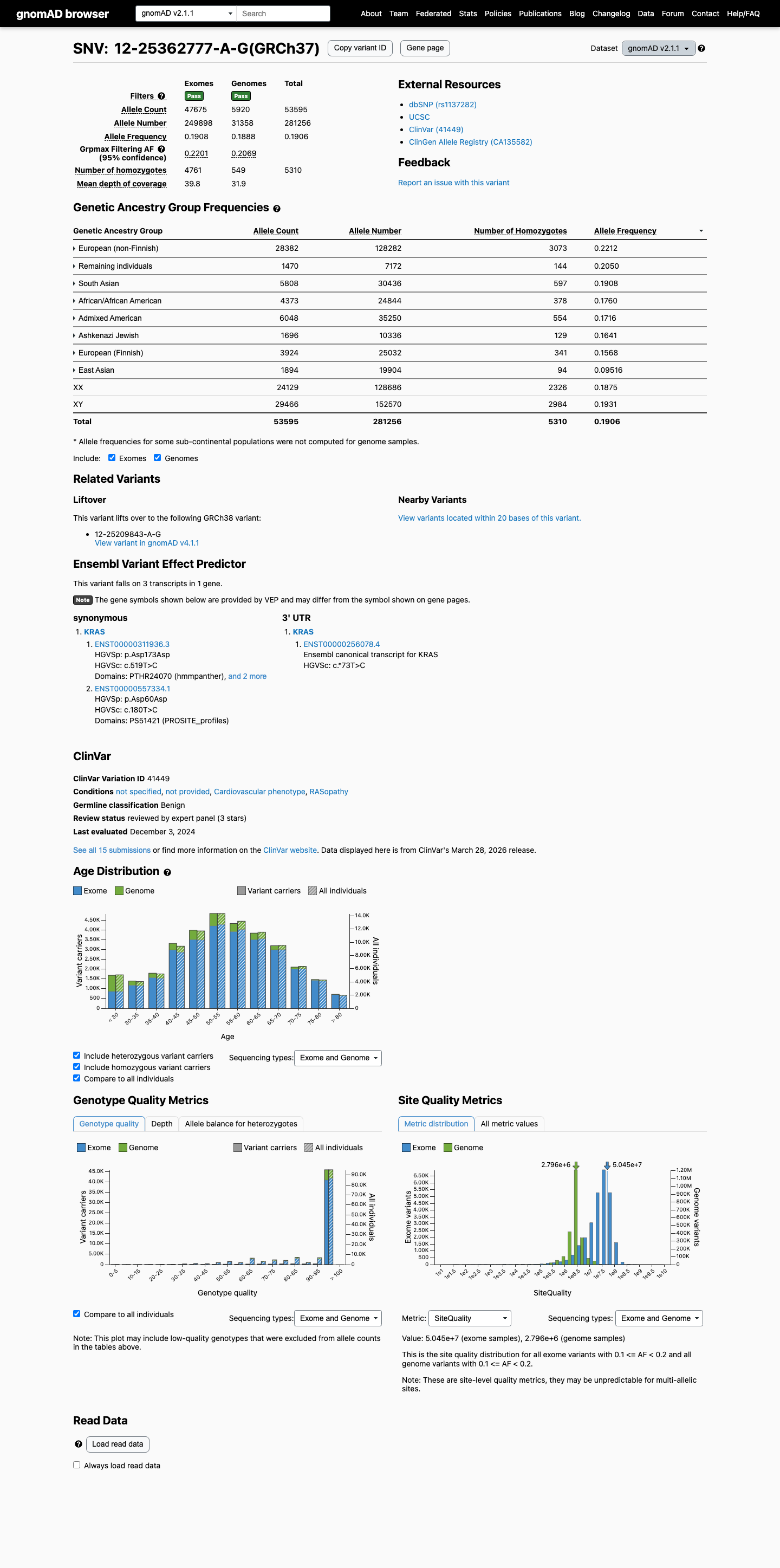
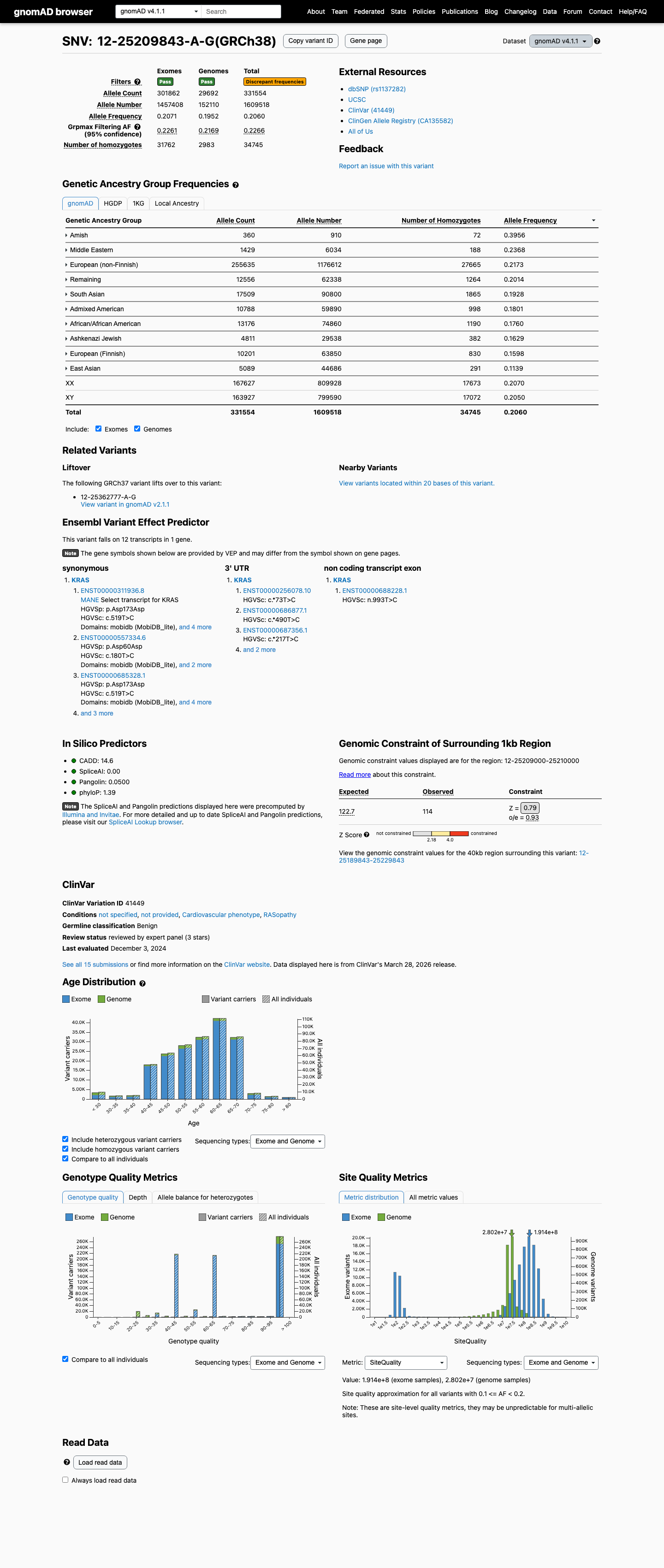
This variant is common in population databases, with an allele frequency of 19.06% in gnomAD v2.1 and 20.60% in gnomAD v4.1, which is far above the KRAS RASopathy VCEP BA1 benign threshold of 0.05%.
cspec ↗ gnomad_v2 ↗ gnomad_v4 ↗3
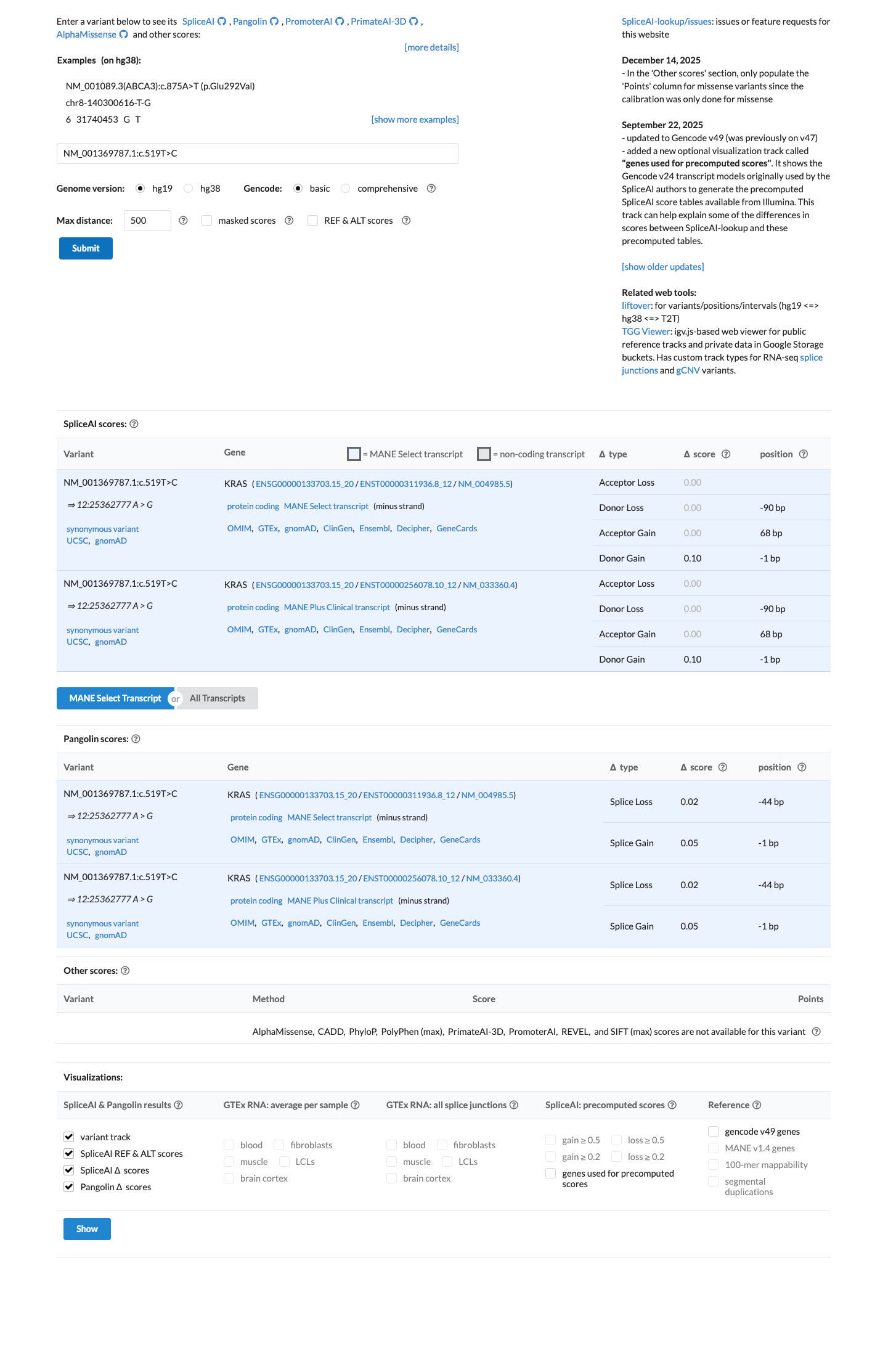
In silico splicing analysis predicts no significant splice impact, with a SpliceAI maximum delta score of 0.10.
spliceai ↗