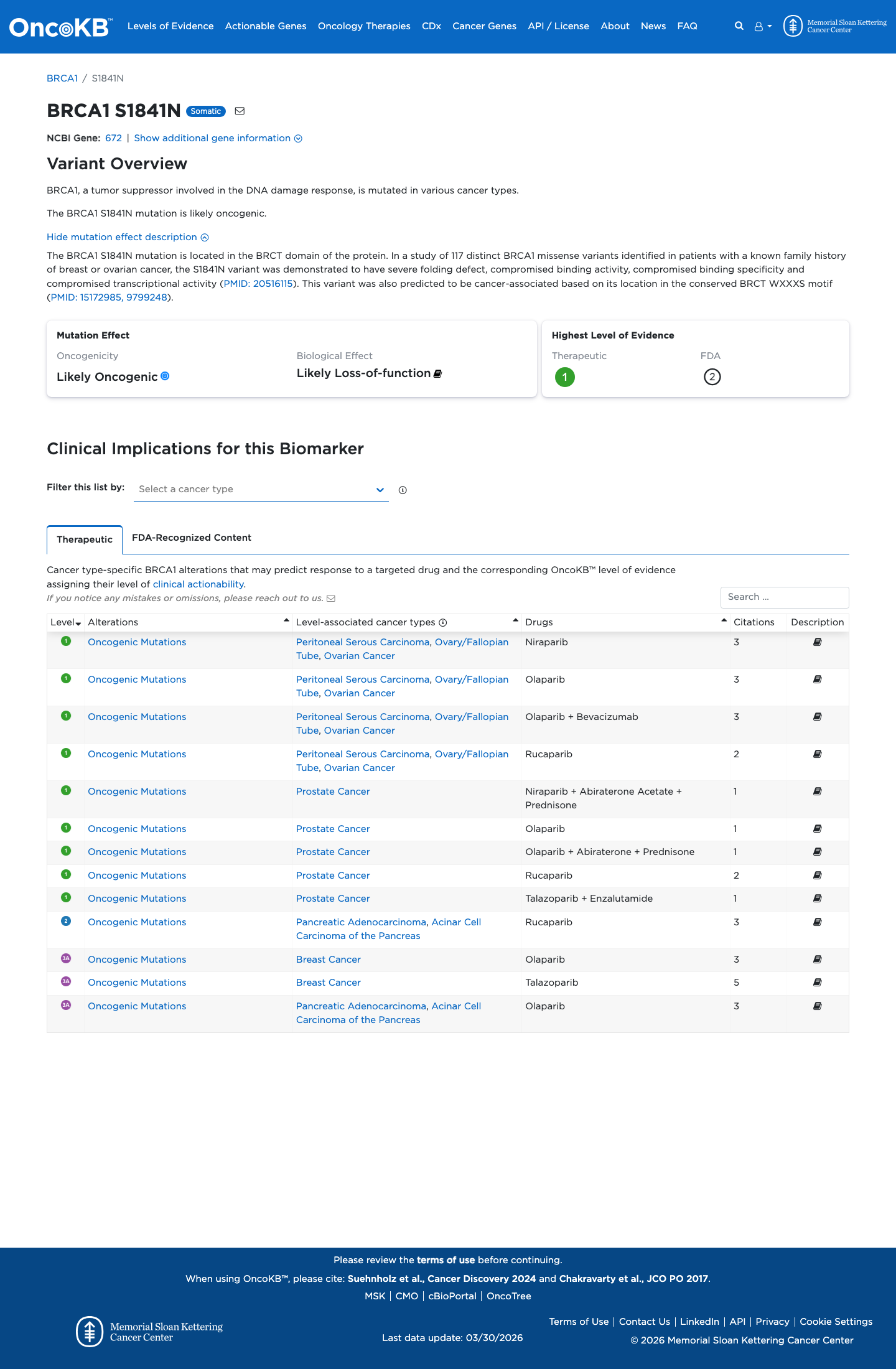
The BRCA1 c.5522G>A (p.Ser1841Asn; p.S1841N) variant has been reported in ClinVar with conflicting interpretations, including an expert-panel classification of uncertain significance.
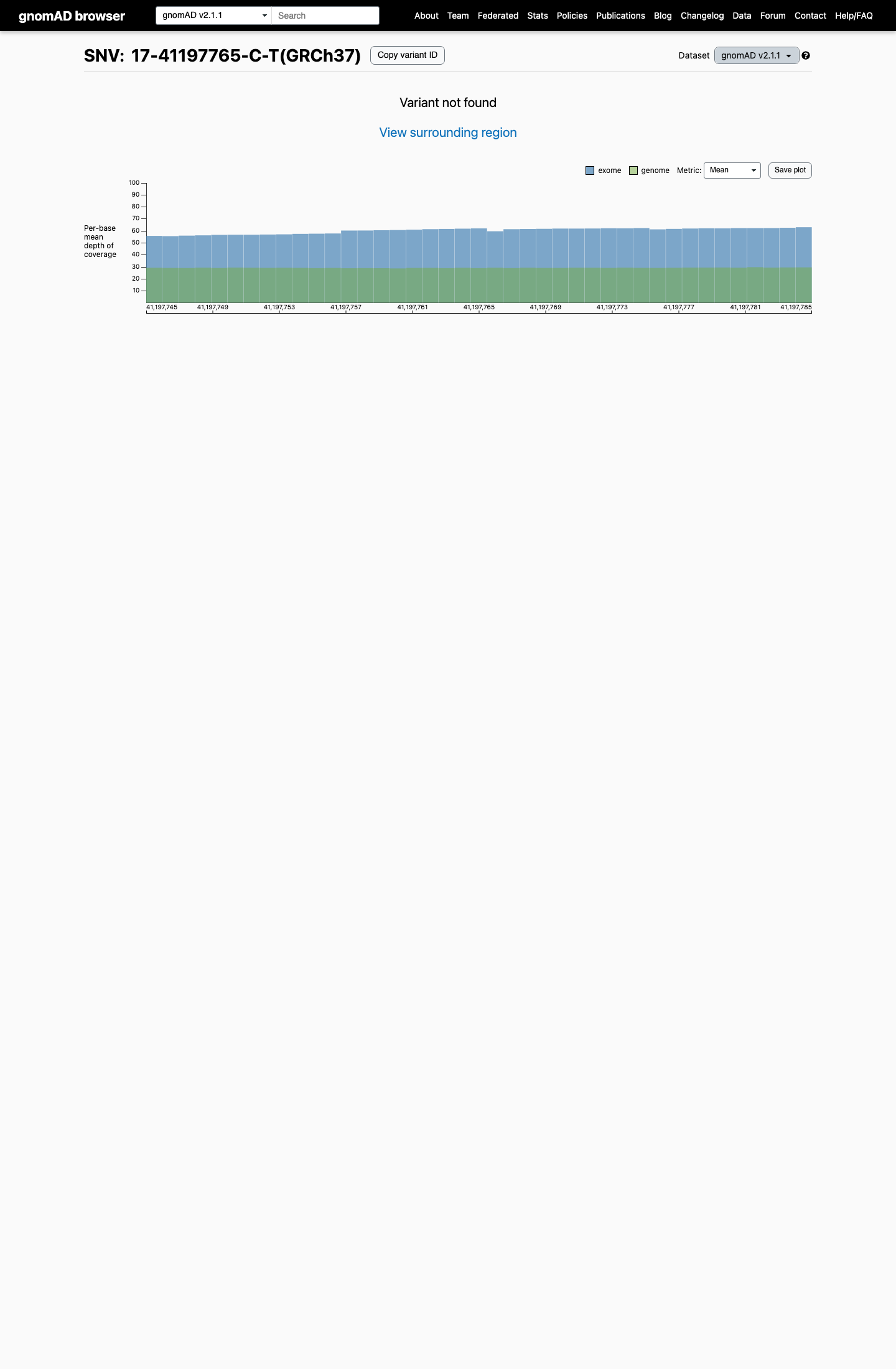
clinvar ↗This variant is absent from gnomAD v2.1 and present once in gnomAD v4.1 (1/1,614,094 alleles; AF 6.20e-07; highest South Asian AF 1.10e-05), so it is very rare but not absent from population databases.
gnomad_v2 ↗ gnomad_v4 ↗In ENIGMA-calibrated functional studies, results were discordant, ranging from loss-of-function to intermediate effects, and ENIGMA Table 9 does not assign PS3 or BS3 for this variant.
This missense change lies in the BRCA1 BRCT repeats, and the BRCA1 ENIGMA computational rule supports BP4 because BayesDel no-AF is 0.144191 (threshold <=0.15) and SpliceAI maximum delta is 0.00 (threshold <=0.1); REVEL is 0.626 but is not the deciding rule in this VCEP framework.
spliceai ↗