Classification rationale
1
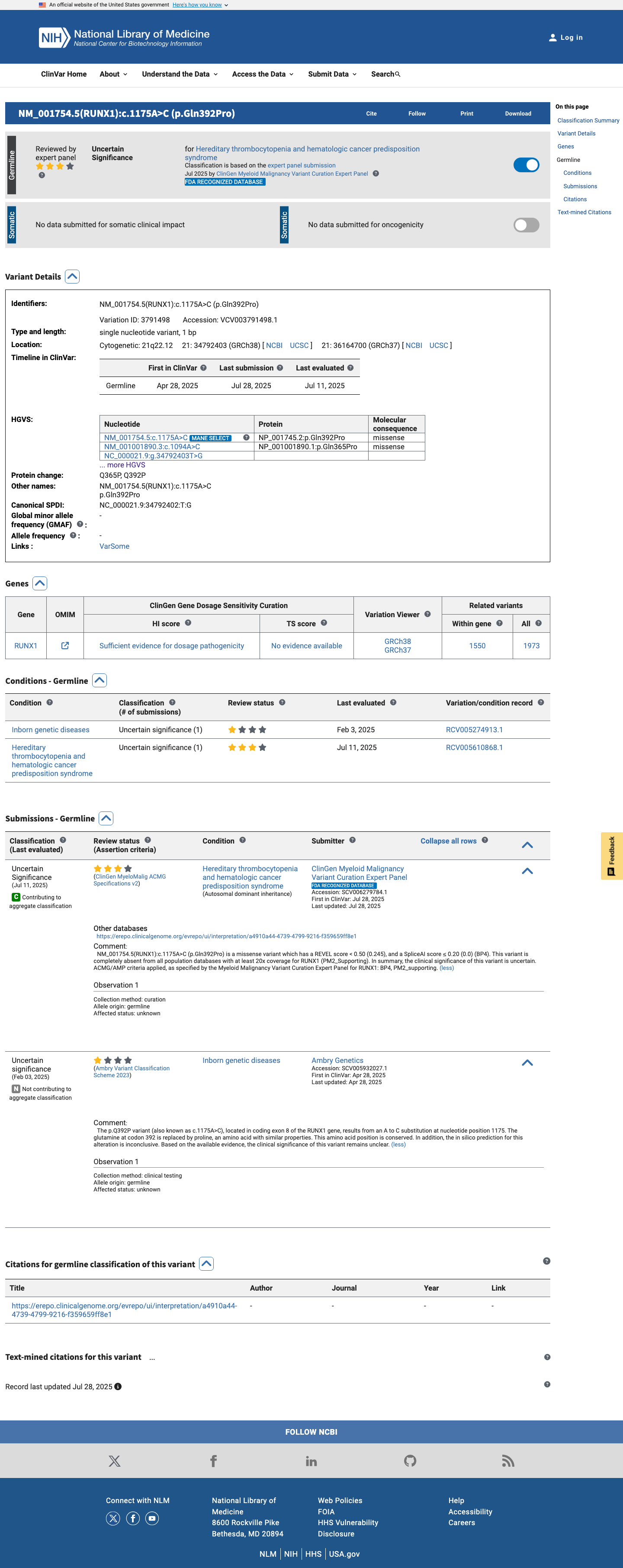
The RUNX1 NM_001001890.2:c.1094A>C (p.Gln365Pro, p.Q365P) variant has been reported in ClinVar and is currently classified there as a variant of uncertain significance by the ClinGen Myeloid Malignancy Variant Curation Expert Panel.
clinvar ↗2
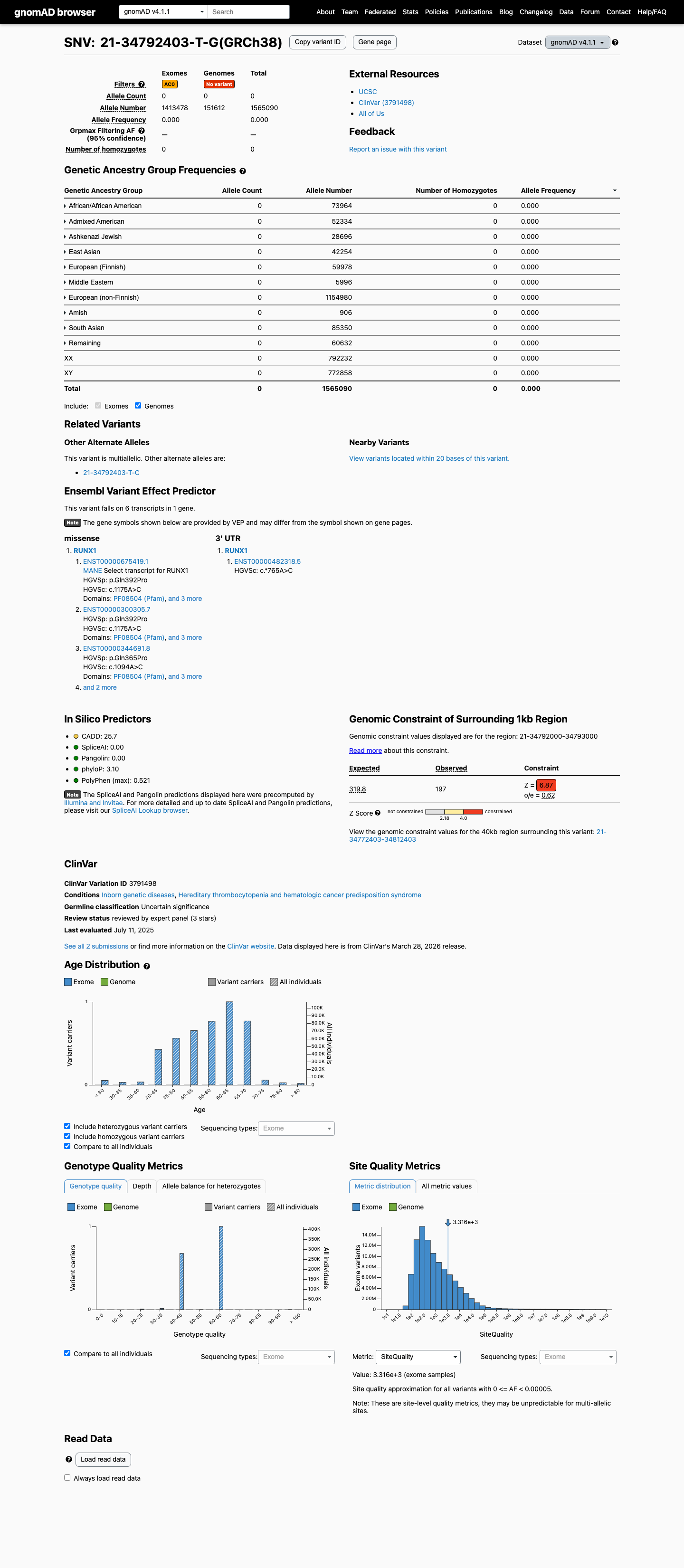
This variant is absent from gnomAD v2.1 and gnomAD v4.1, with 0/1565090 alleles observed in gnomAD v4.1, which supports very low population frequency for RUNX1 disease.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
Computational evidence does not support a damaging effect, with REVEL 0.245, SpliceAI maximum delta score 0.00, and BayesDel -0.122289; these findings support BP4 and do not support PP3.
spliceai ↗ cspec ↗