Classification rationale
1
The PALB2 c.2027T>C (p.Ile676Thr) variant has been reported in ClinVar as Benign, including a reviewed expert panel classification.
clinvar ↗2
This variant is present in gnomAD v4.1 with an overall allele frequency of 0.01425% (230/1614160) and a highest observed population frequency of 0.37653% (226/60022 alleles in the Admixed American population), which is above the PALB2 BA1 threshold of 0.1% and the BS1 threshold of 0.01%.
gnomad_v4 ↗ cspec ↗3
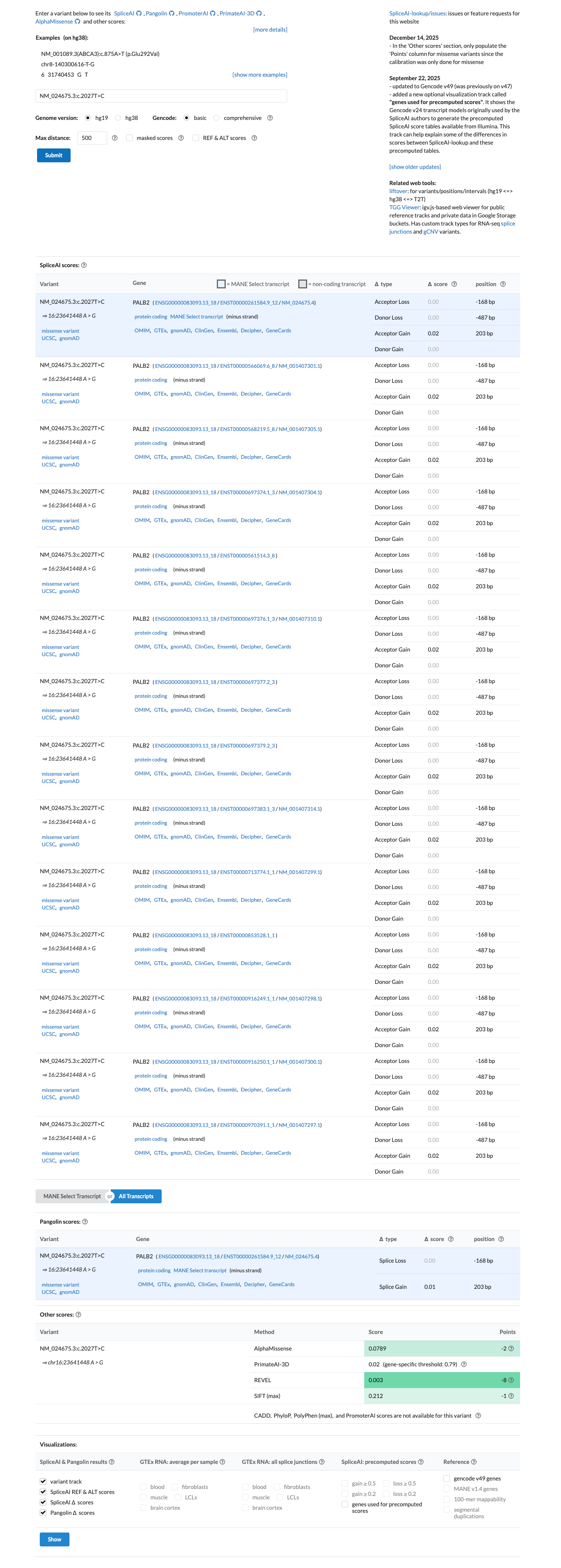
In silico data do not support a splice effect, with a SpliceAI maximum delta score of 0.02; REVEL is 0.003 and BayesDel is -0.705338, but the PALB2 VCEP does not use PP3 or BP4 for missense variants.
spliceai ↗ cspec ↗