Classification rationale
1
2
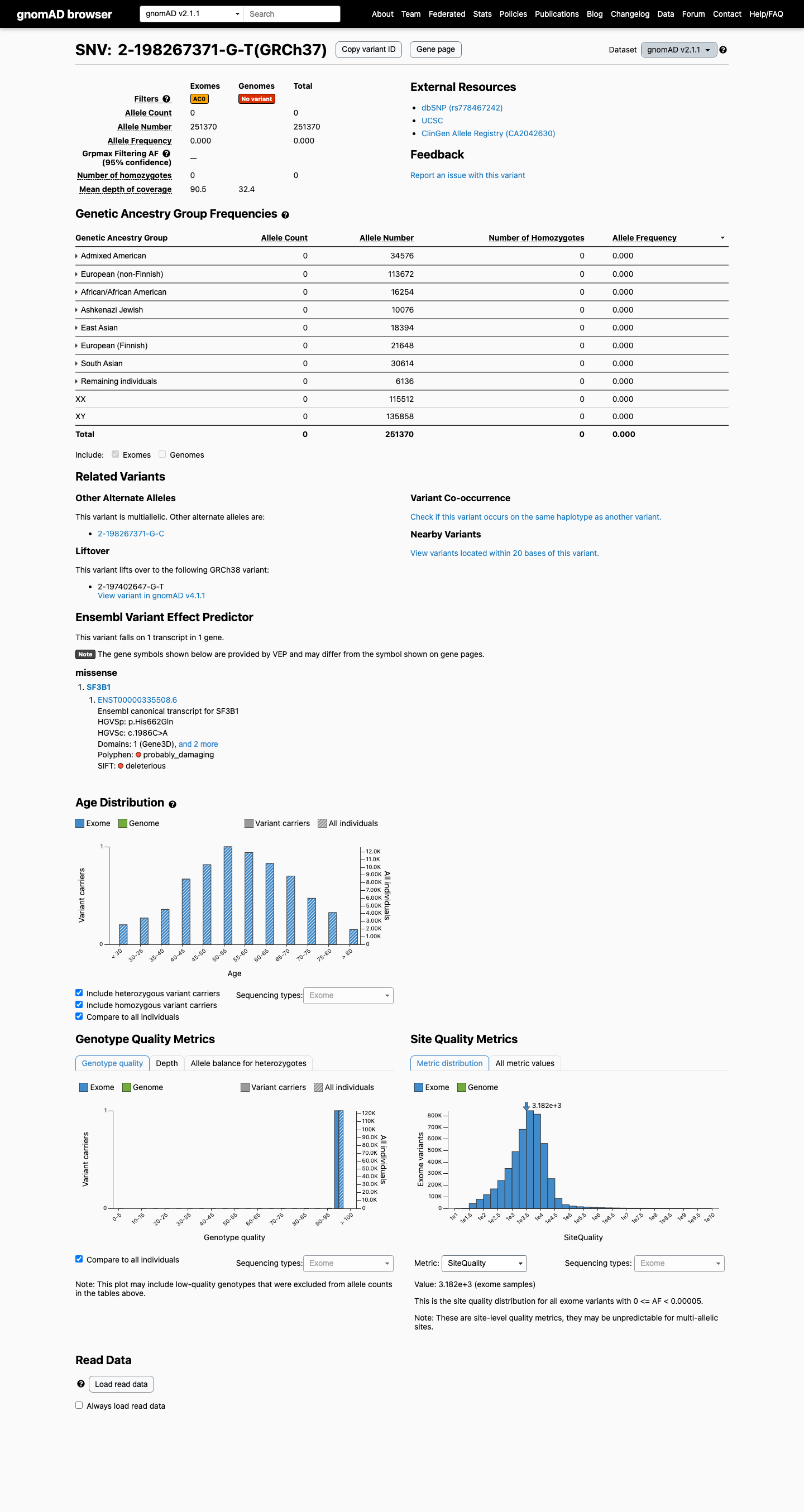
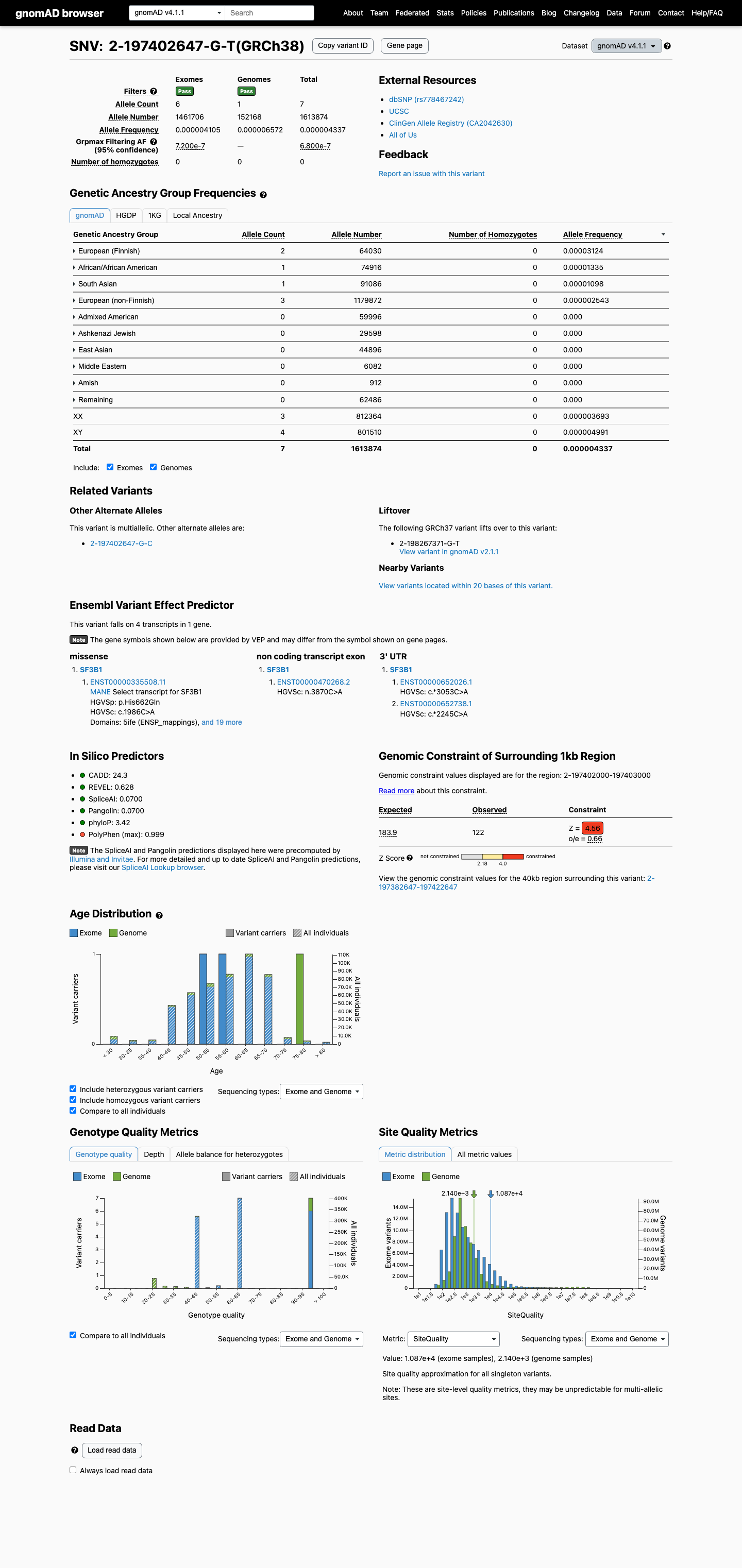
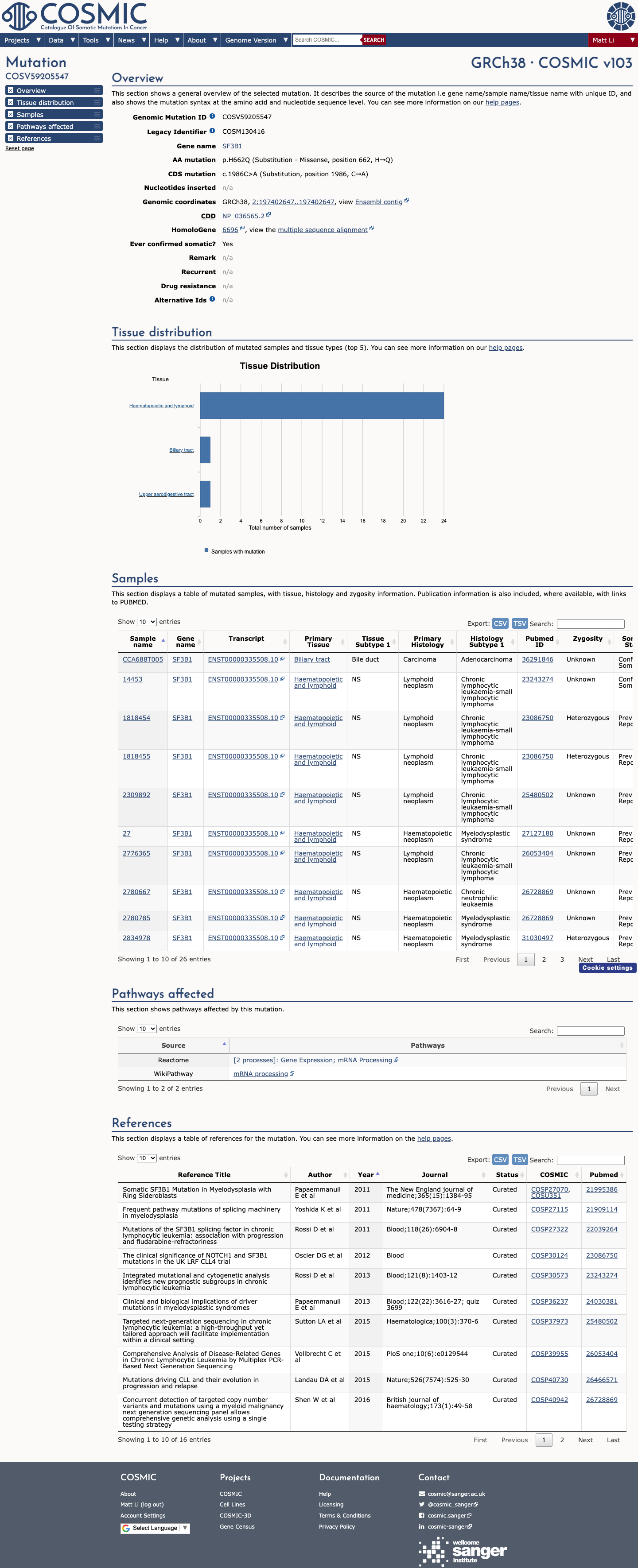
This variant is absent from gnomAD v2.1 and remains very rare in gnomAD v4.1 at 7/1613874 alleles (0.00043%), with the highest observed population frequency of 0.00312% in Finnish individuals, which is below the 0.1% PM2 rarity threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
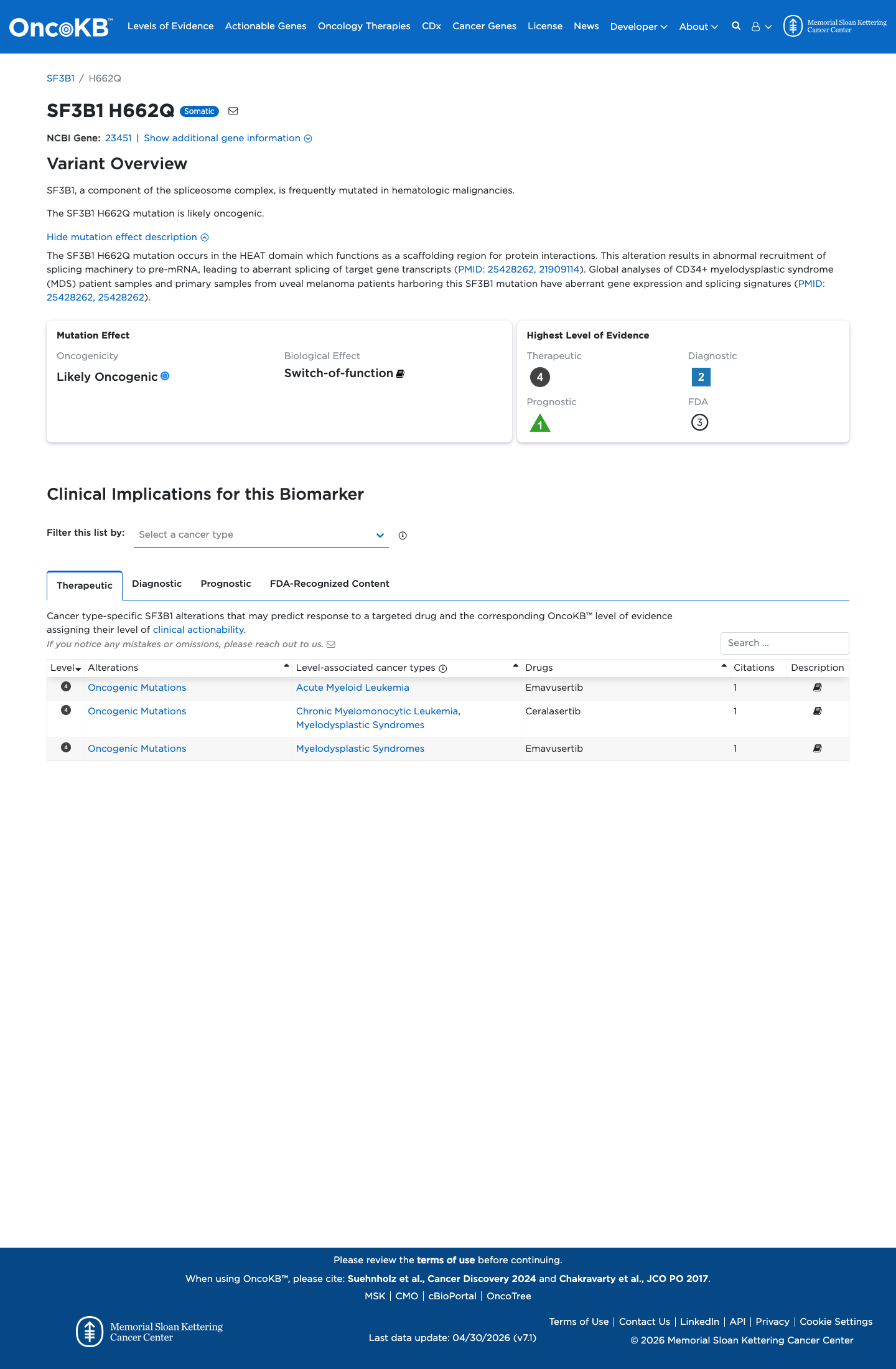
Published SF3B1 functional literature was identified for review, but the available evidence did not provide a validated assay directly testing p.(His662Gln), so functional criteria were not applied.
oncokb ↗ PMID:21909114 ↗ PMID:25428262 ↗4
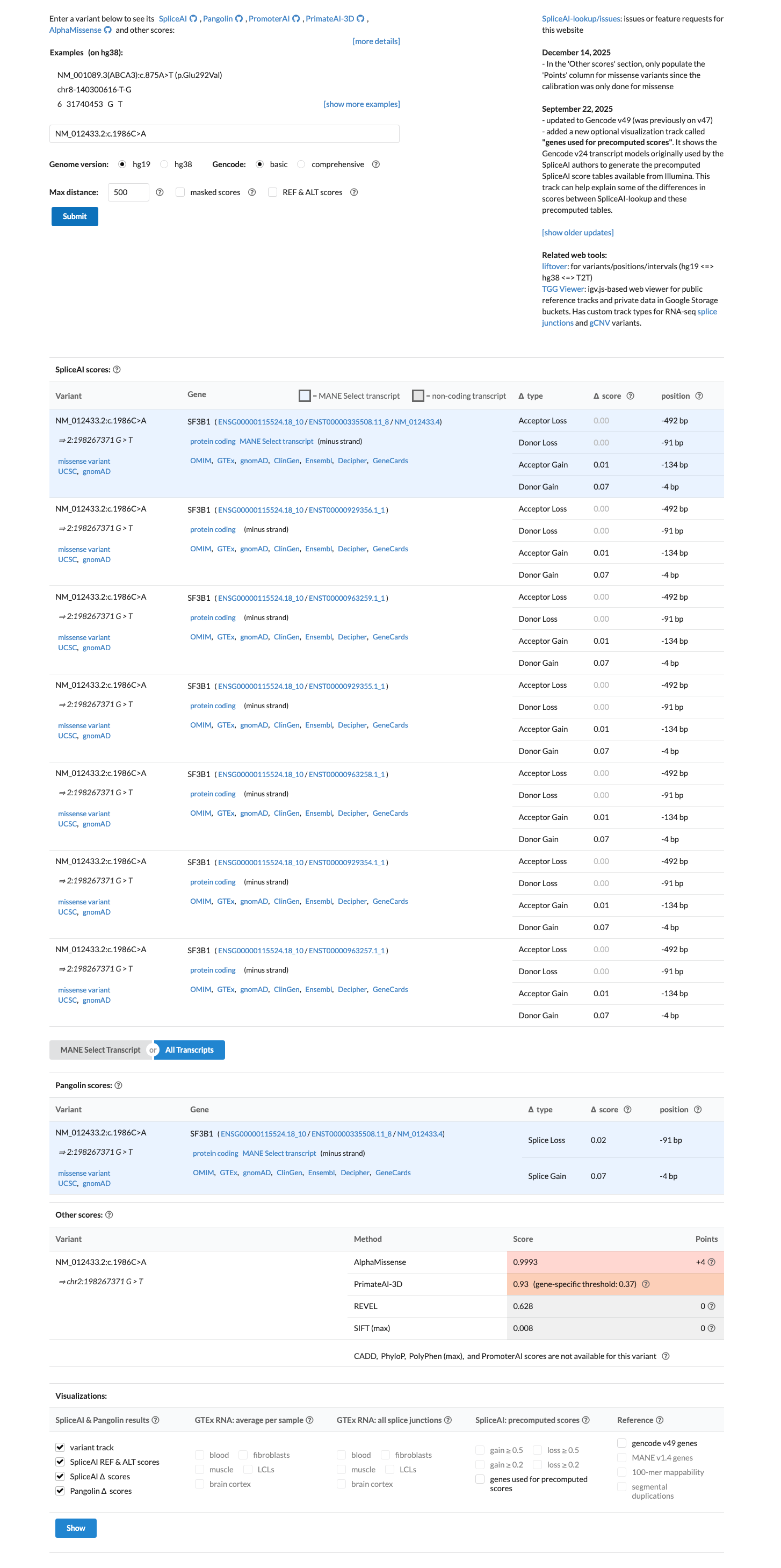
Computational evidence supports a deleterious missense effect, with REVEL 0.628 and BayesDel 0.0218, while SpliceAI shows a low maximum delta score of 0.07 and does not support a splice-altering effect.
spliceai ↗