Classification rationale
1
2
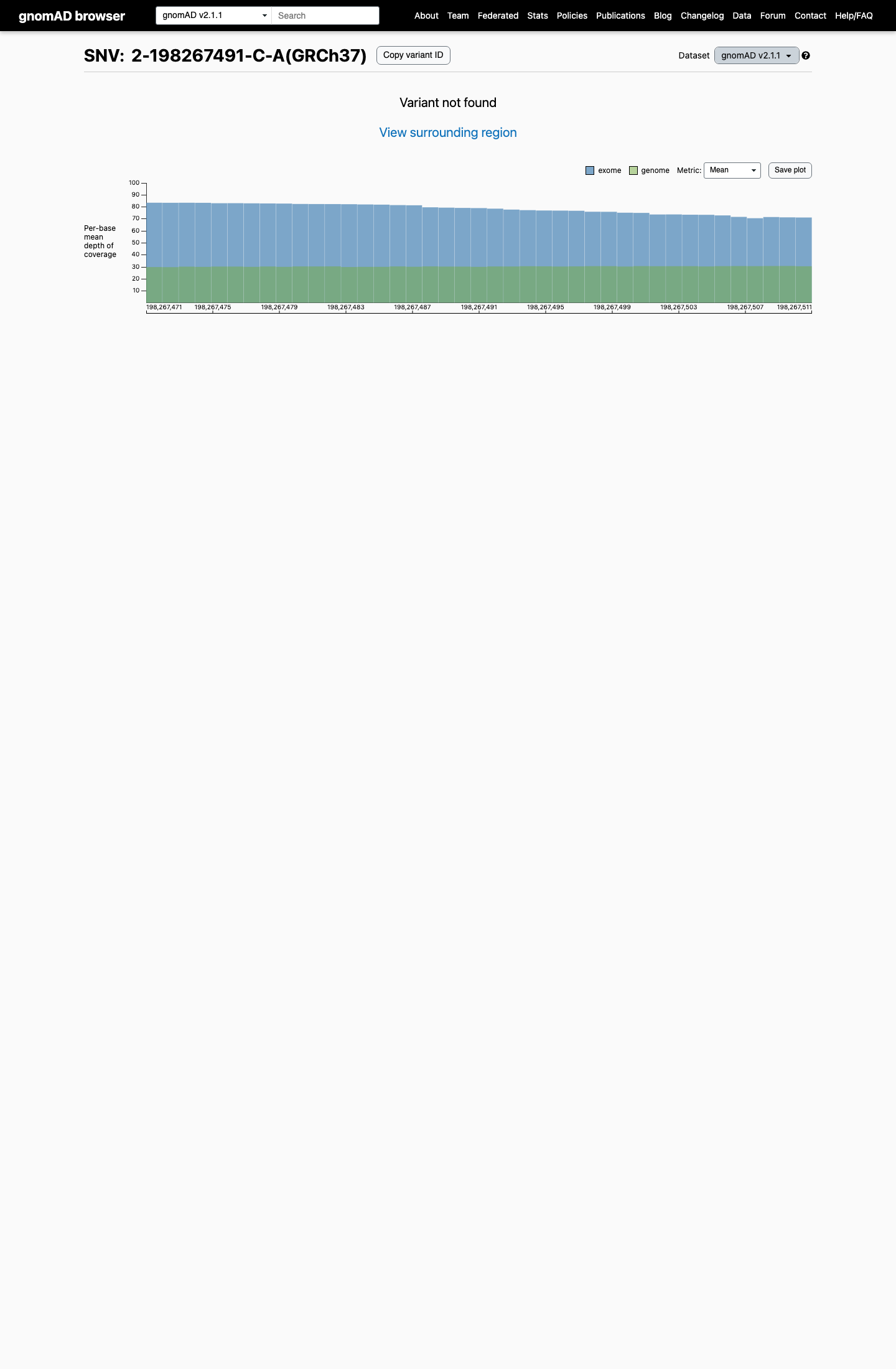
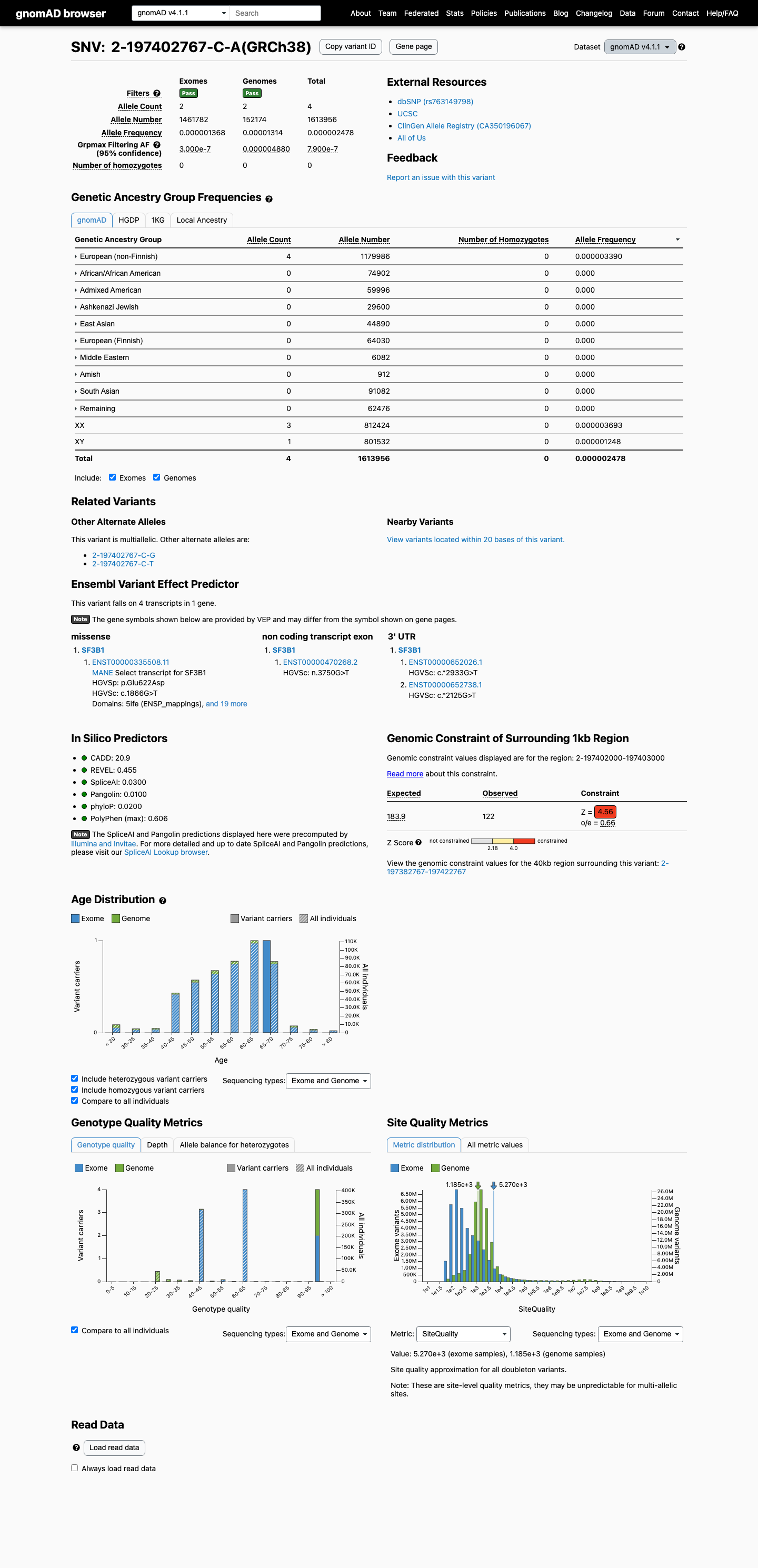
In population databases, this variant is absent from gnomAD v2.1 and is present at very low frequency in gnomAD v4.1 (AF 2.478382310298422e-06, 4/1613956 alleles), which is below the default PM2 threshold of 0.1%.
gnomad_v2 ↗ gnomad_v4 ↗3
Published studies show that disease-associated SF3B1 missense variants cluster in functionally important splicing regions and can disrupt RNA splicing, but no well-established functional assay specific to p.Glu622Asp was identified.
PMID:21909114 ↗ PMID:25428262 ↗4
Computational results are mixed: REVEL is 0.455, BayesDel is -0.0897007, and SpliceAI predicts a possible splice effect with a maximum delta score of 0.27, so in silico evidence does not clearly support either a damaging or benign interpretation.
spliceai ↗