Classification rationale
1
2
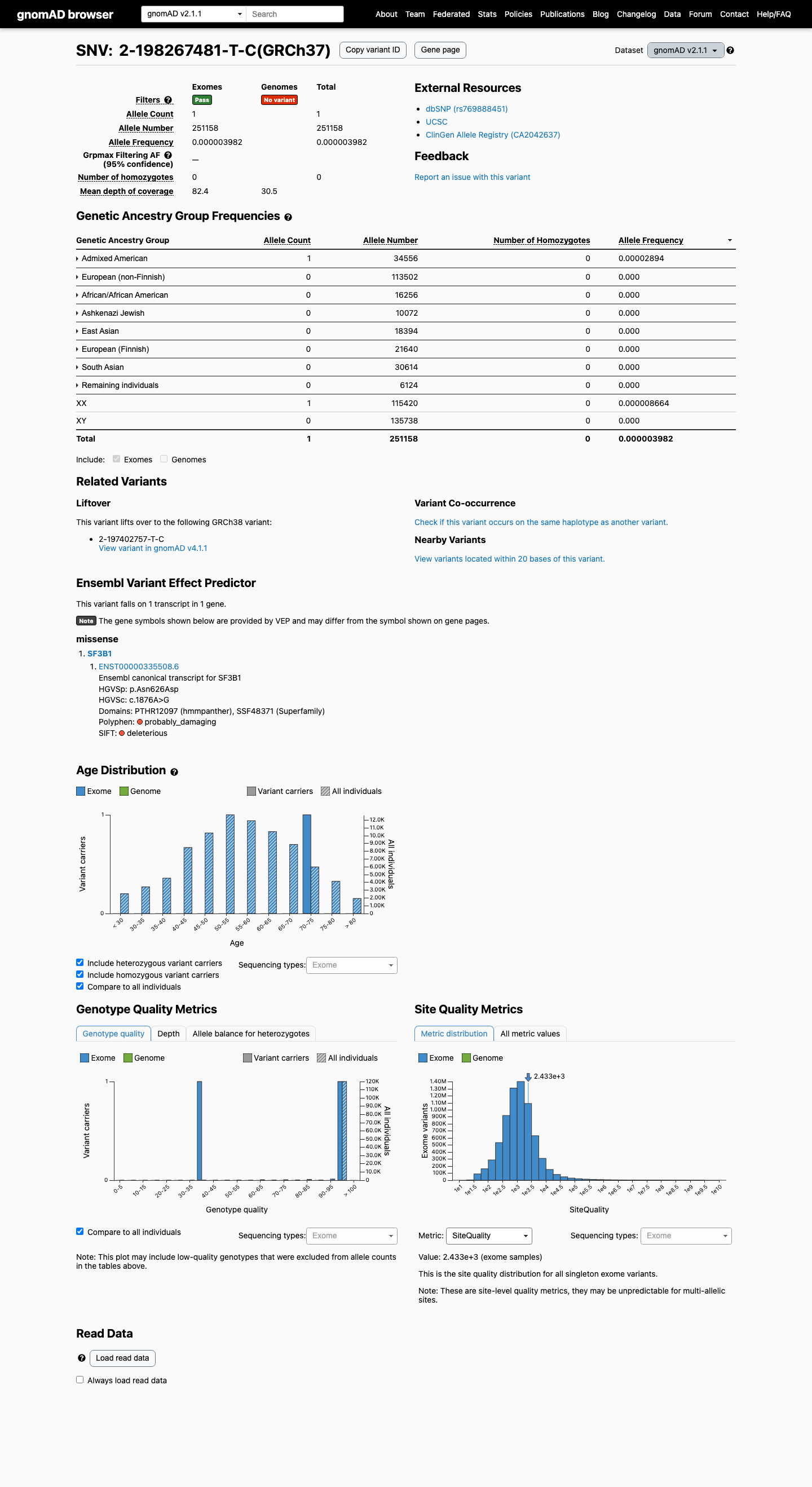
This variant is rare in population databases, with gnomAD v2.1 showing 1/251158 alleles (0.00040%) and gnomAD v4.1 showing 8/1613994 alleles (0.00050%; grpmax FAF 0.000124%), which is below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
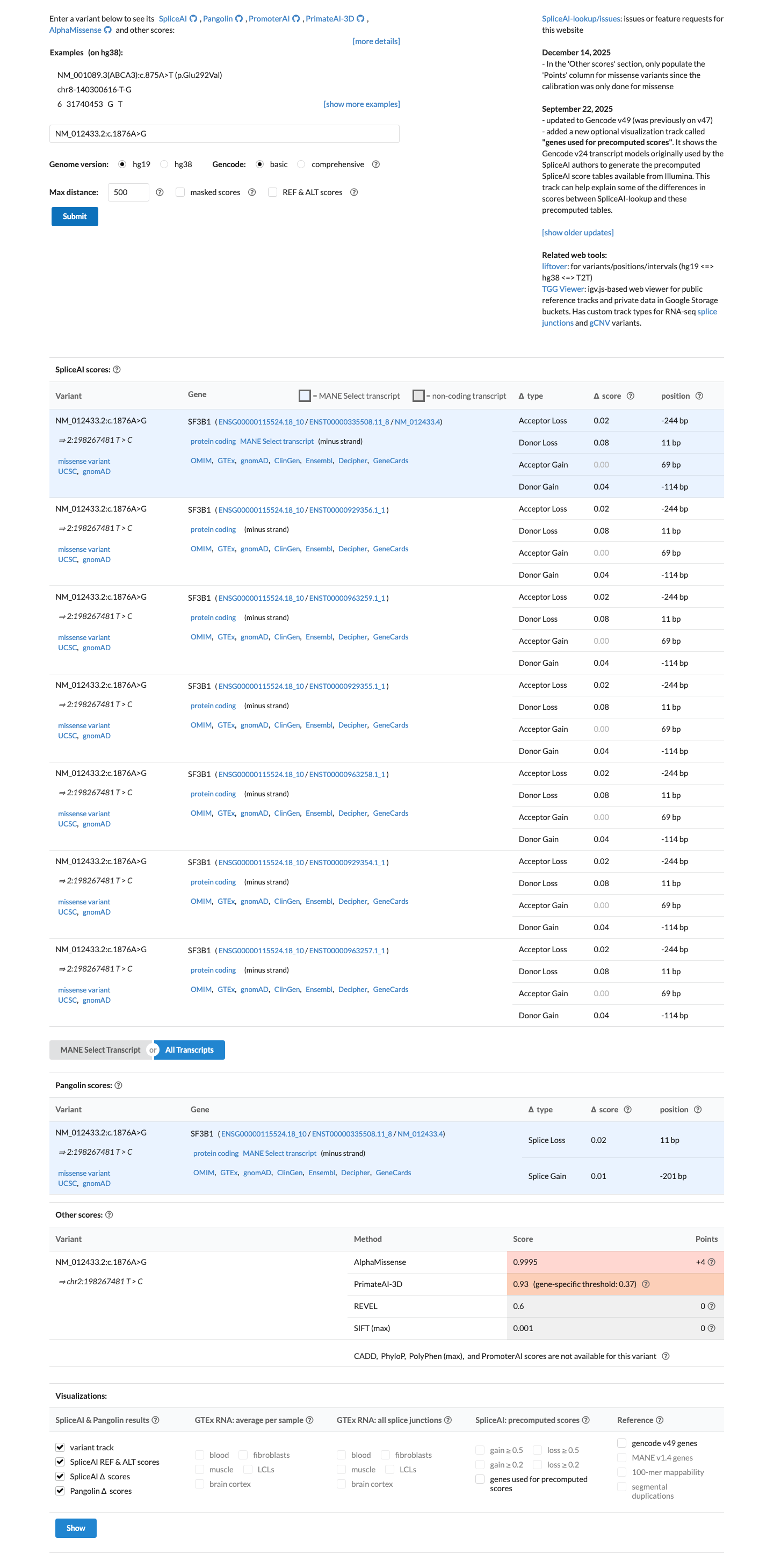
In silico results are mixed: SpliceAI predicts no significant splice impact (maximum delta score 0.08), while REVEL is 0.60 and BayesDel is -0.0733292, so computational evidence does not independently support PP3 or BP4.
spliceai ↗