Classification rationale
1
The BARD1 c.764A>G (p.Asn255Ser) variant has been reported in ClinVar predominantly as uncertain significance, with 11 uncertain significance submissions and 1 likely benign submission.
clinvar ↗2
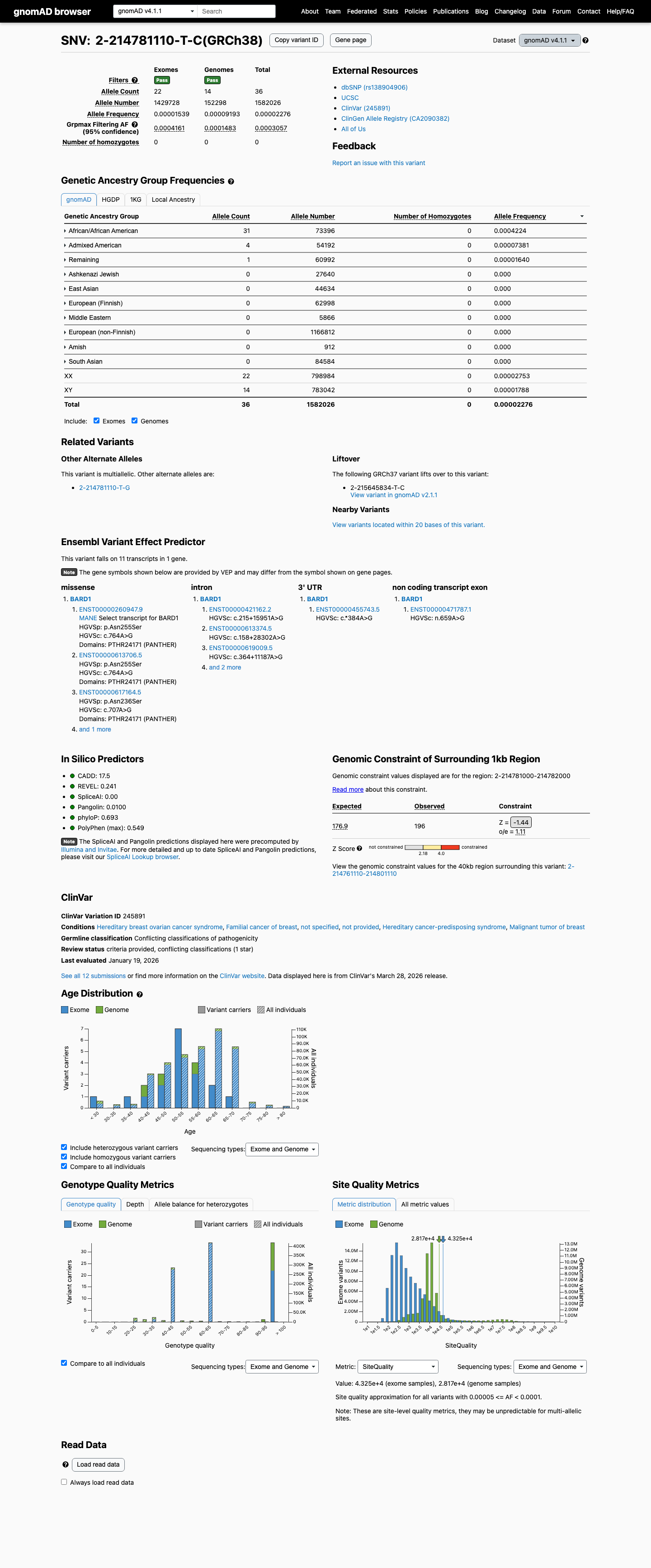
This variant is present at low frequency in population databases, with a total allele frequency of 0.00316% in gnomAD v2.1 and 0.00228% in gnomAD v4.1; the highest observed population frequency is 0.03280% in gnomAD v2.1 and 0.04224% in gnomAD v4.1 in the African/African American population, which remains below the 0.1% rarity threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
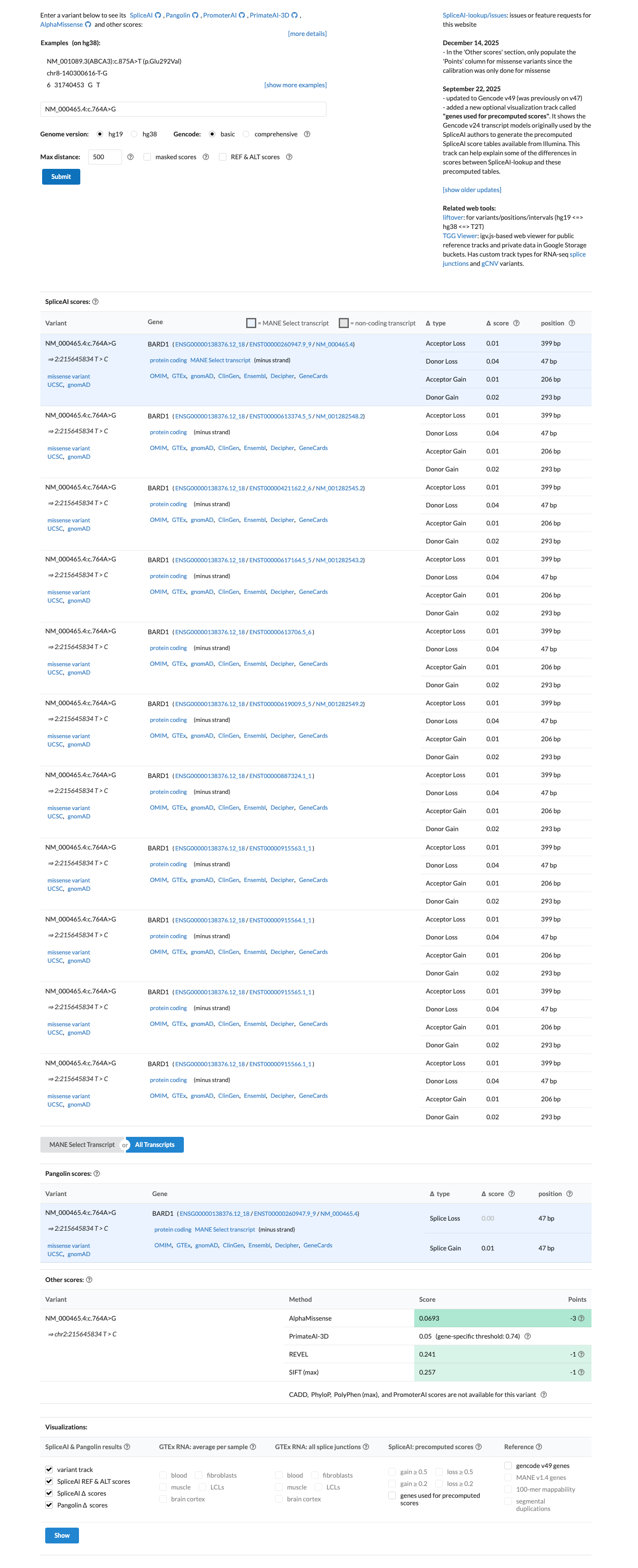
Available computational evidence is consistent with no significant functional impact, with SpliceAI predicting no significant splice effect (maximum delta score 0.04), REVEL 0.241, and BayesDel -0.546443.
spliceai ↗