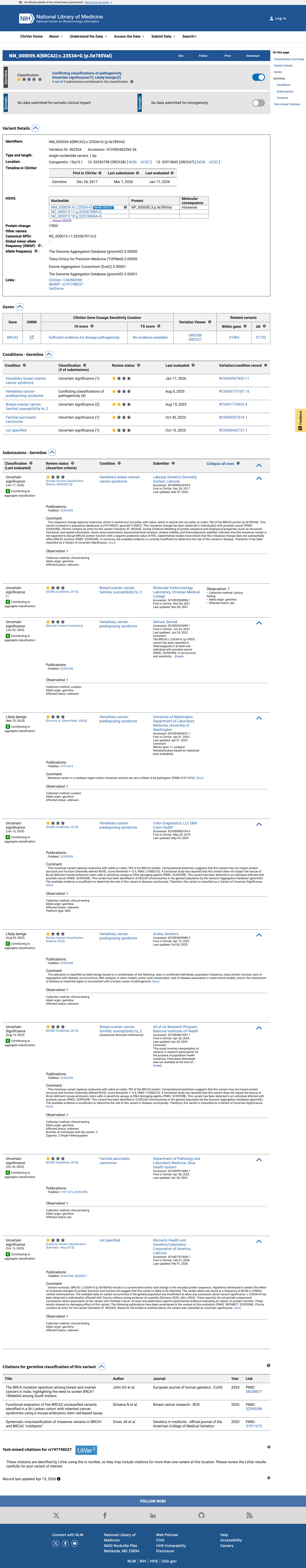
The BRCA2 NM_000059.4:c.2353A>G (p.(Ile785Val), p.(I785V)) variant has been reported in ClinVar, with predominantly uncertain significance submissions and one likely benign submission.
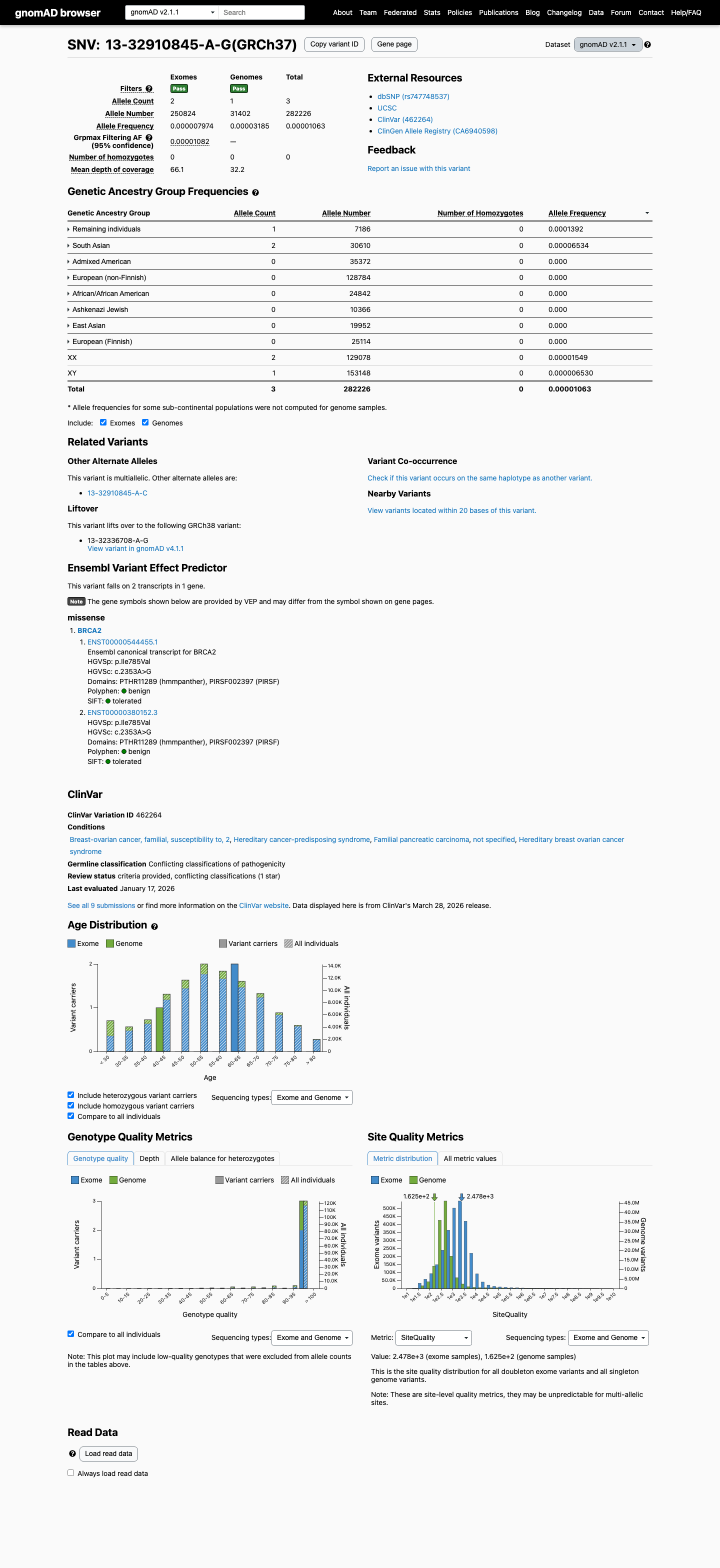
clinvar ↗This variant is present in gnomAD v2.1 at AF 1.06298e-05 (3/282226 alleles) with grpmax FAF 1.082e-05, which is below ENIGMA BA1 and BS1 thresholds and also means the variant is not absent from controls for PM2.
gnomad_v2 ↗ cspec ↗In a published mouse embryonic stem cell-based assay, I785V was reported as functionally indistinguishable from wild-type BRCA2; however, this variant was not identified with a preassigned PS3 or BS3 code in the ENIGMA curated functional table, so the functional evidence remains for manual review.
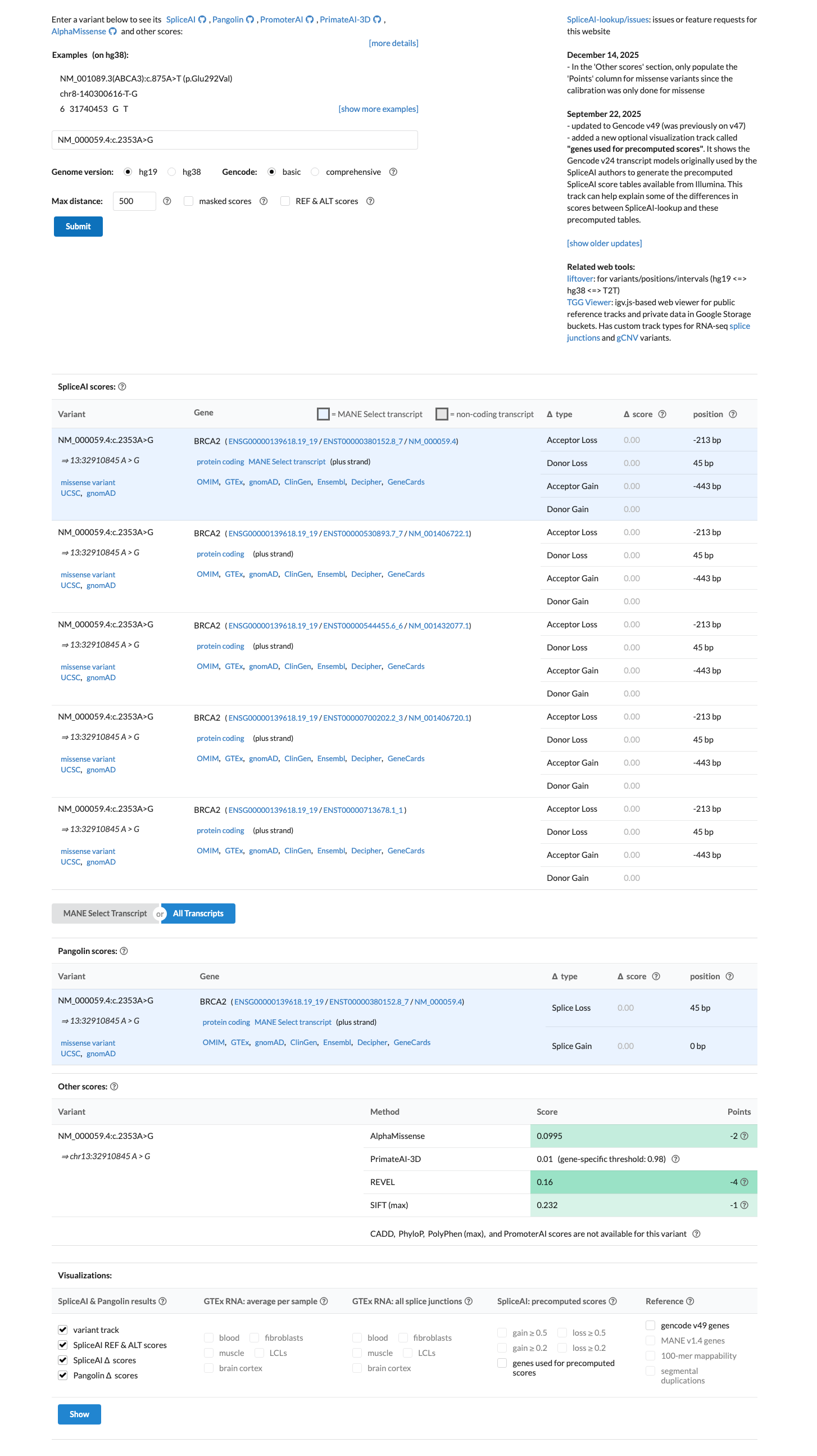
PMID:32393398 ↗ cspec ↗This missense change lies outside the BRCA2 PALB2-binding and DNA-binding domains used for ENIGMA missense pathogenicity assessment, SpliceAI predicts no splice impact (max delta 0.00), BayesDel is -0.394429, and REVEL is 0.16, supporting BP1_Strong and arguing against PP3.
cspec ↗ spliceai ↗