Classification rationale
1
The LPL c.1385T>C (p.Phe462Ser; p.F462S) variant has been reported in ClinVar as uncertain significance by two clinical laboratories.
clinvar ↗2
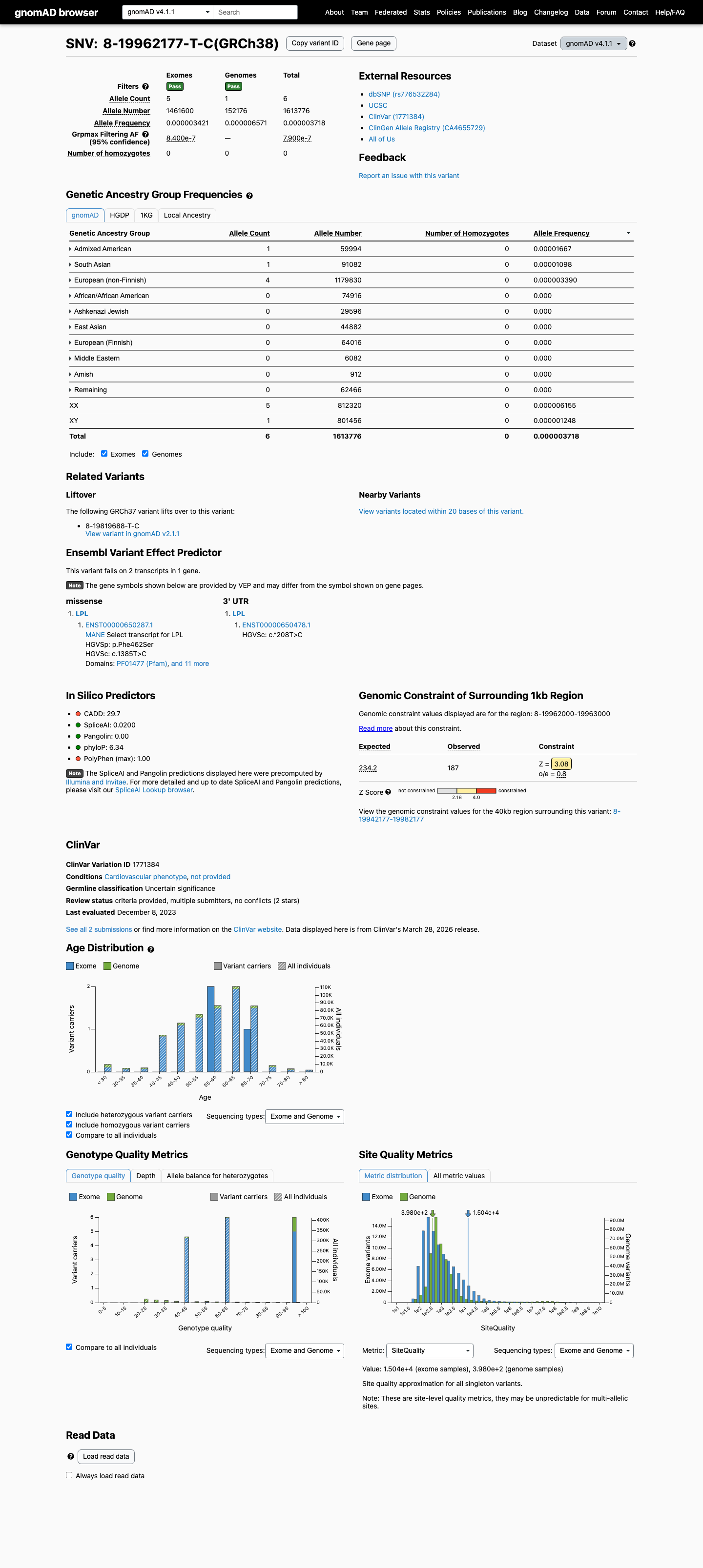
This variant is rare in population databases, with allele frequencies of 0.00080% in gnomAD v2.1 (2/251272) and 0.00037% in gnomAD v4.1 (6/1613776), which are both below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
Computational evidence supports a deleterious effect, with REVEL 0.735 and BayesDel 0.304545, while SpliceAI predicts no significant splice impact with a maximum delta score of 0.04.
spliceai ↗