Classification rationale
1
The MSH2 NM_000251.3:c.1661+1G>A (NP_000242.1:p.?) variant has been reported in ClinVar, including an InSiGHT expert panel classification of likely pathogenic and additional pathogenic/likely pathogenic clinical laboratory submissions.
clinvar ↗2
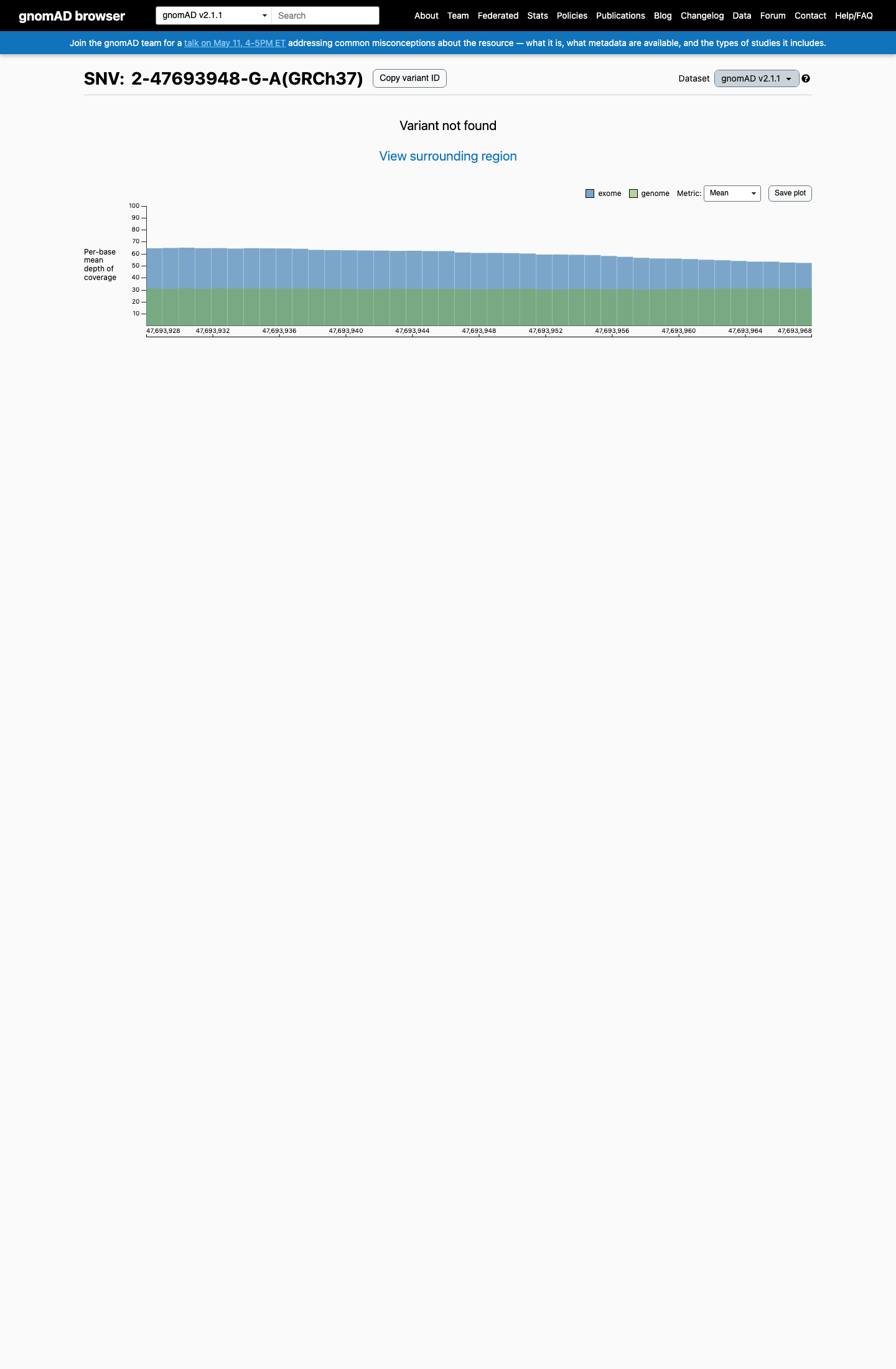
This variant is absent from gnomAD v2.1 and is present only once in gnomAD v4.1 (1/1,612,780 alleles; AF 6.20e-07), which is below the MSH2 VCEP PM2 threshold of 0.00002.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
SpliceAI predicts a strong splice effect for this canonical donor variant (max delta score 0.97), supporting predicted disruption of normal splicing; however, this computational evidence was not counted separately because the same splice effect is captured under PVS1 for a canonical +1 splice-site variant.
spliceai ↗ cspec ↗