Classification rationale
1
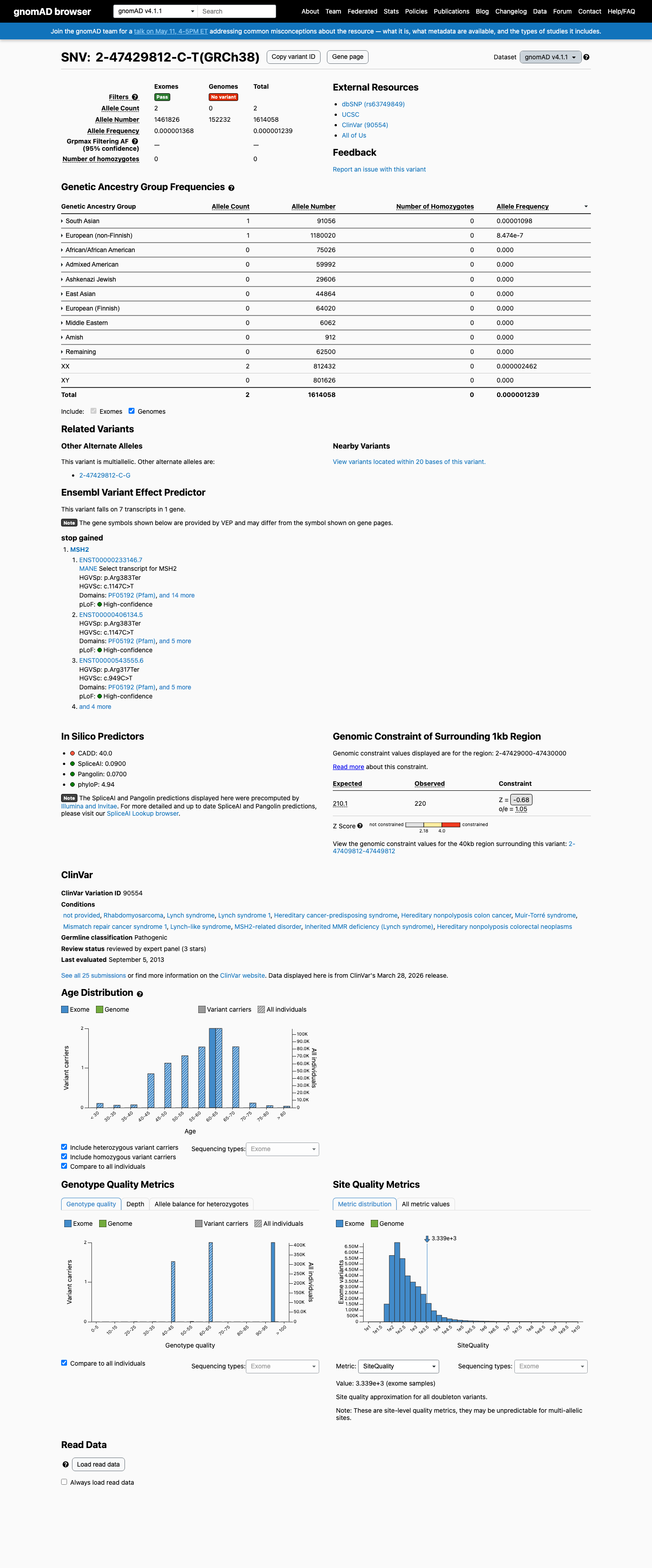
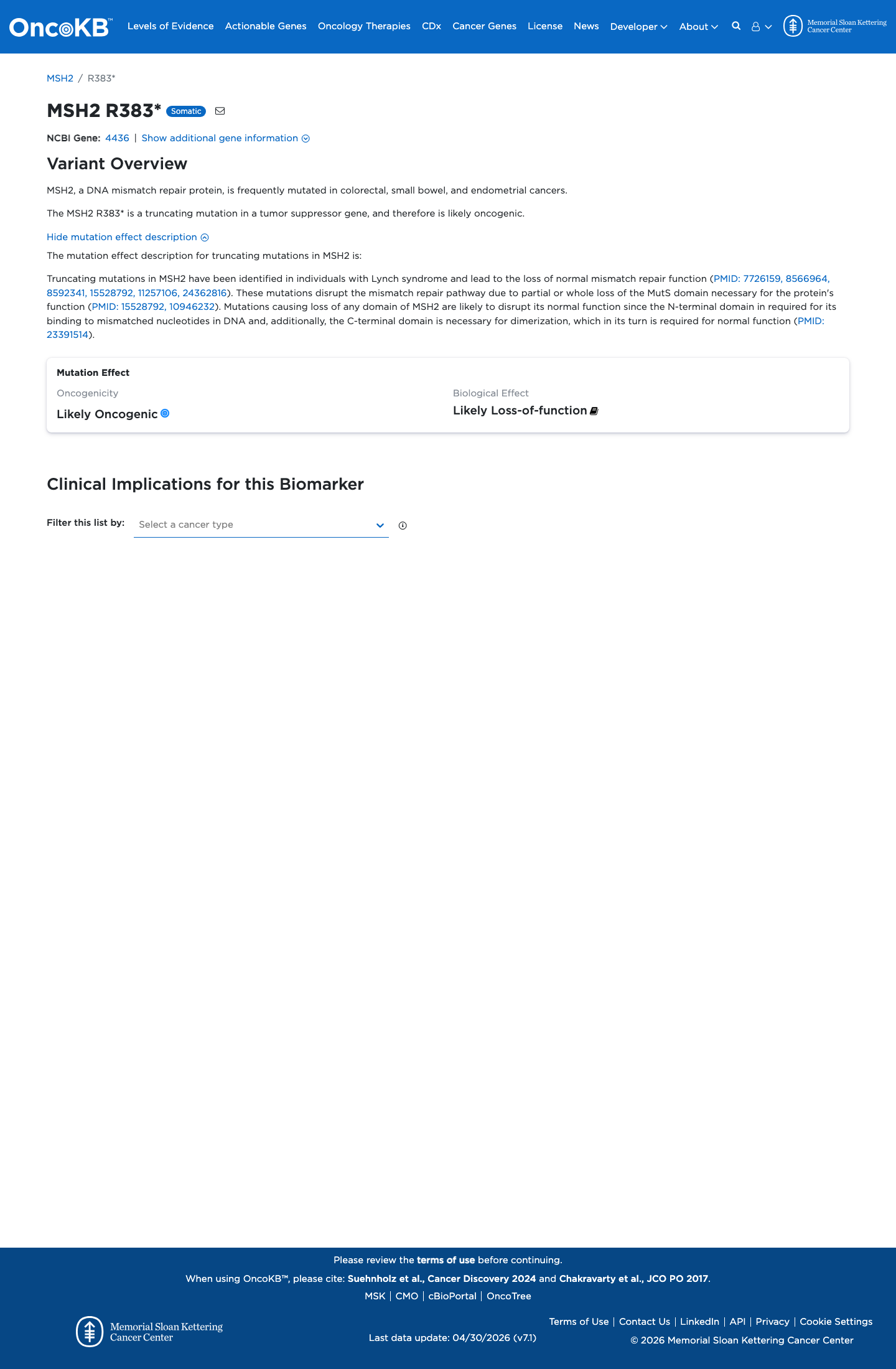
The MSH2 NM_000251.3:c.1147C>T (p.Arg383Ter; p.R383*) variant has been reported in ClinVar and includes an expert panel Pathogenic classification.
clinvar ↗2
This variant is absent from gnomAD v2.1 and is present in gnomAD v4.1 at 2/1614058 alleles (AF 1.23911e-06; 0.00012%), which is below the MSH2 PM2 threshold of 0.00002.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
This nonsense variant introduces a premature stop at codon 383, which is well upstream of the MSH2 codon 891 cutoff for very strong PVS1 evidence and is consistent with loss of function as an established disease mechanism for MSH2-related disease.
cspec ↗