Classification rationale
1
The MSH6 c.2091T>C (p.Asp697=) variant has not been observed in somatic cancers in COSMIC.
2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, and its gnomAD v4.1 frequency is therefore below the MSH6 PM2_Supporting threshold of less than 0.00002.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
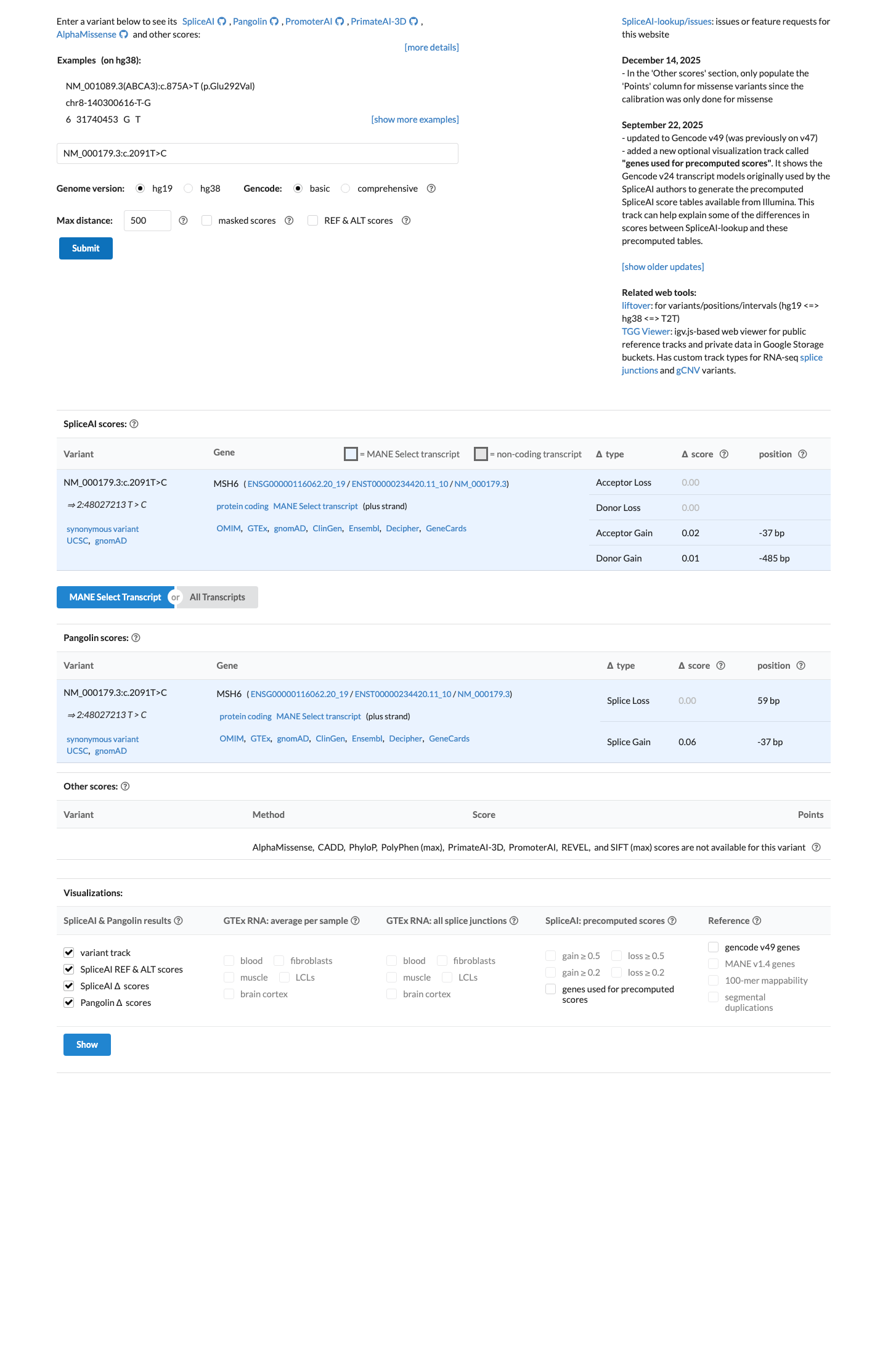
This synonymous exon 4 variant is far from the splice junction boundaries used for MSH6 BP7, and SpliceAI predicts no meaningful splice effect with a maximum delta score of 0.02, supporting BP7 and BP4 while arguing against PP3 or PVS1 based on the currently available evidence.
spliceai ↗ cspec ↗