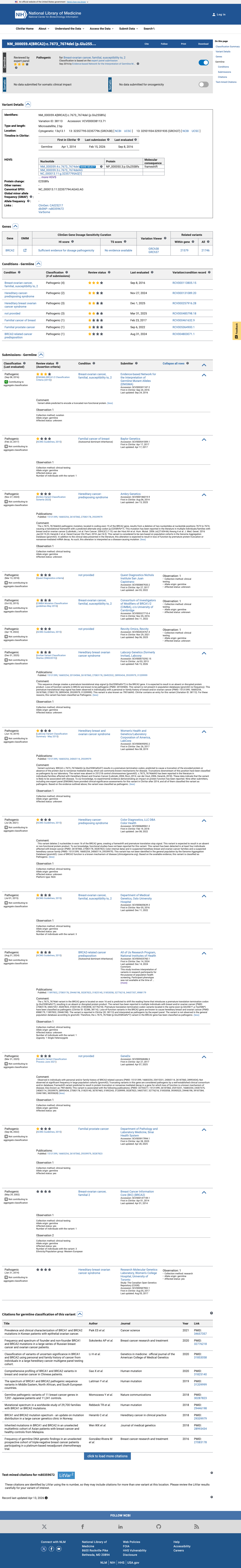
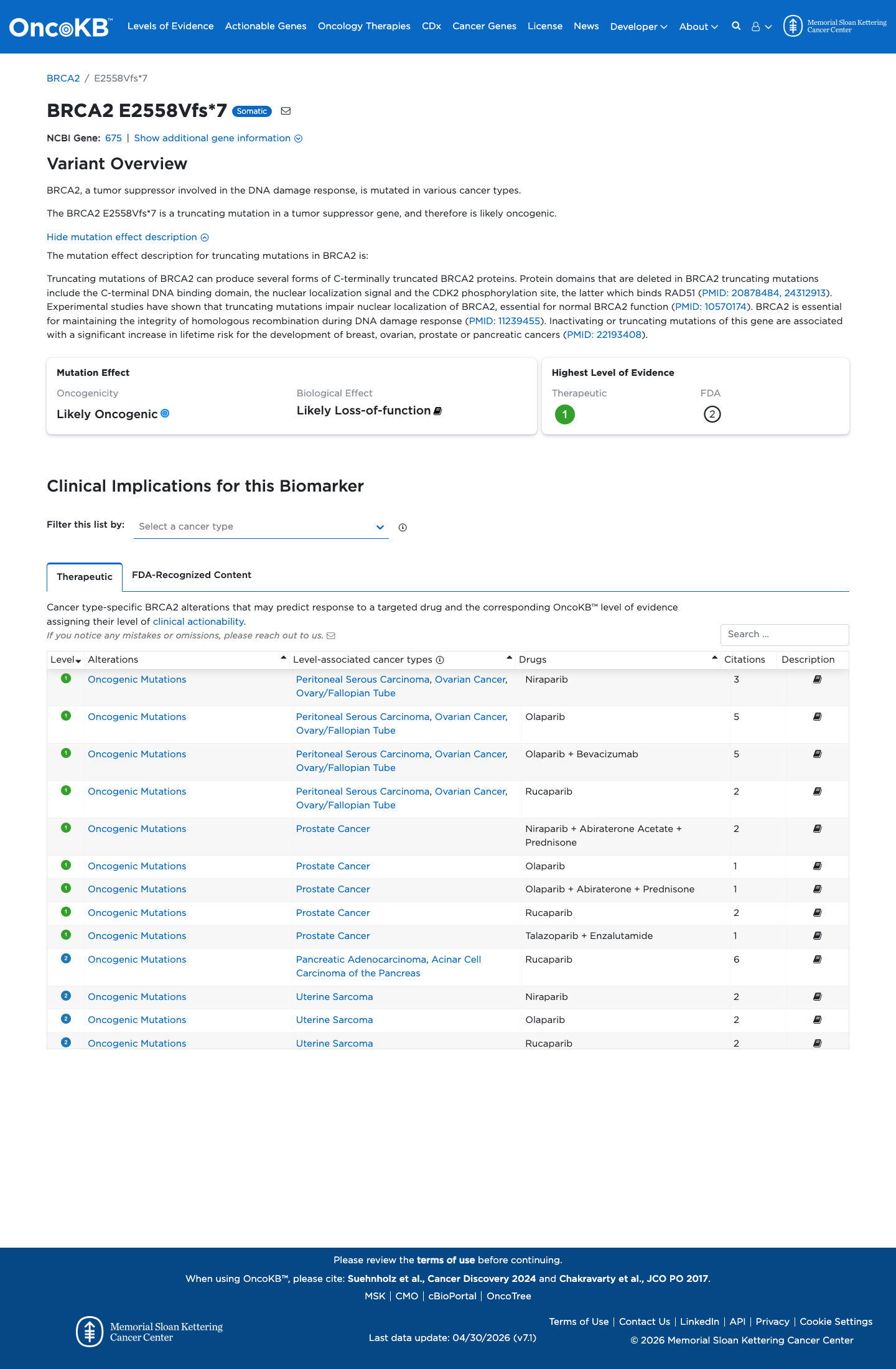
The BRCA2 c.7673_7674delAG (p.Glu2558ValfsTer7; p.E2558Vfs*7) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as Pathogenic, including ENIGMA expert panel review.
clinvar ↗This variant is absent from gnomAD v2.1 and present once in gnomAD v4.1 (1/1,613,678 alleles; total AF 6.197e-07; highest observed population AF 1.098e-05 in South Asian individuals), supporting extremely low population frequency but not complete absence from population databases.
gnomad_v2 ↗ gnomad_v4 ↗Under the ENIGMA BRCA2 loss-of-function framework, this exon 16 frameshift is eligible for PVS1, and the exon-level truncating-variant table assigns additional PM5_PTC evidence at Strong strength for this exon.
cspec ↗SpliceAI predicts possible splice impact with a maximum delta score of 0.24, while REVEL and BayesDel are not applicable to this deletion; this computational result was reviewed but not used as separate PP3 evidence because the variant was adjudicated through the loss-of-function framework.
spliceai ↗ cspec ↗