Classification rationale
1
The BRIP1 NM_032043.3:c.1474-3T>C (NP_114432.2:p.?) variant has been reported in ClinVar with predominantly benign and likely benign submissions, although uncertain significance submissions are also present.
clinvar ↗2
This variant is present in population databases, including gnomAD v4.1 at an overall allele frequency of 0.01185% (191/1611312) and a highest observed South Asian frequency of 0.20440% (186/90996), with similar South Asian enrichment in gnomAD v2.1 at 0.16343% (50/30594); these values are above a 0.1% PM2 rarity threshold but below BA1 and BS1 thresholds.
gnomad_v2 ↗ gnomad_v4 ↗3
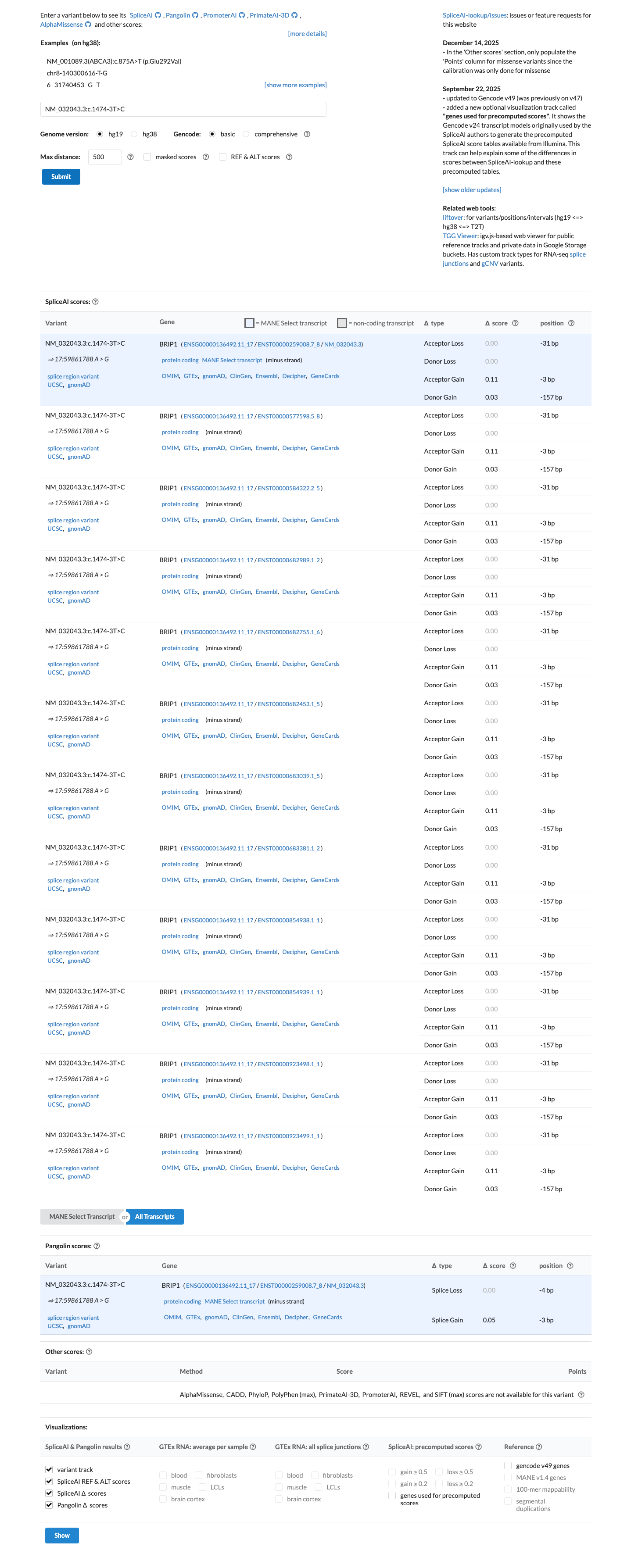
In silico splice prediction does not support a significant splice effect, with a SpliceAI maximum delta score of 0.11, supporting BP4 and arguing against PP3.
spliceai ↗