Classification rationale
1
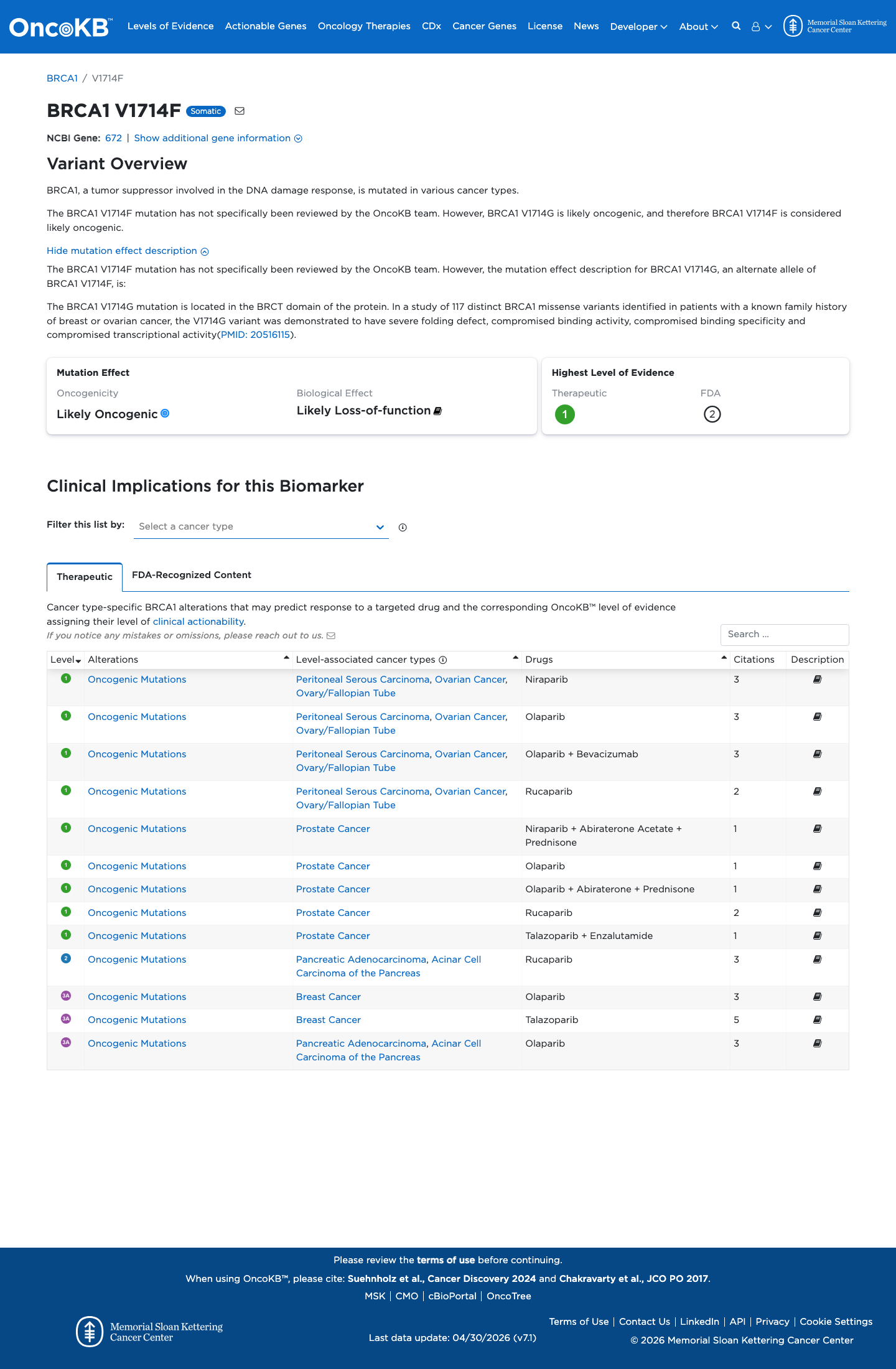
The BRCA1 c.5140G>T (p.Val1714Phe) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar, where the overall classification is Likely Pathogenic with expert panel review.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population databases.
gnomad_v2 ↗ gnomad_v4 ↗3
In a calibrated BRCA1 functional study, this variant showed complete functional impact consistent with loss of function, and ENIGMA assigns PS3_Strong for this variant.
PMID:30209399 ↗4
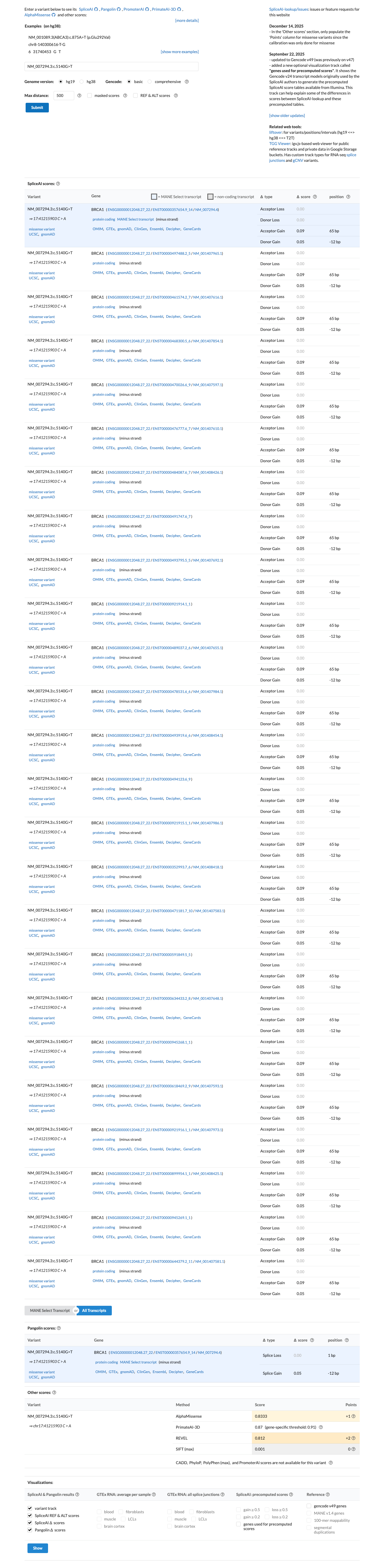
Computational data support a damaging protein effect because the variant is in the BRCT repeats, BayesDel no-AF is 0.482282, and REVEL is 0.812, while SpliceAI predicts no significant splice effect with a max delta score of 0.09.
spliceai ↗ cspec ↗