The PALB2 c.3512del (p.(Leu1171CysfsTer20), p.(L1171Cfs*20)) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar as uncertain significance by the ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, which is below the PALB2 PM2_Supporting threshold of 0.000333% and does not meet the BA1 (>0.1%) or BS1 (>0.01%) population thresholds.
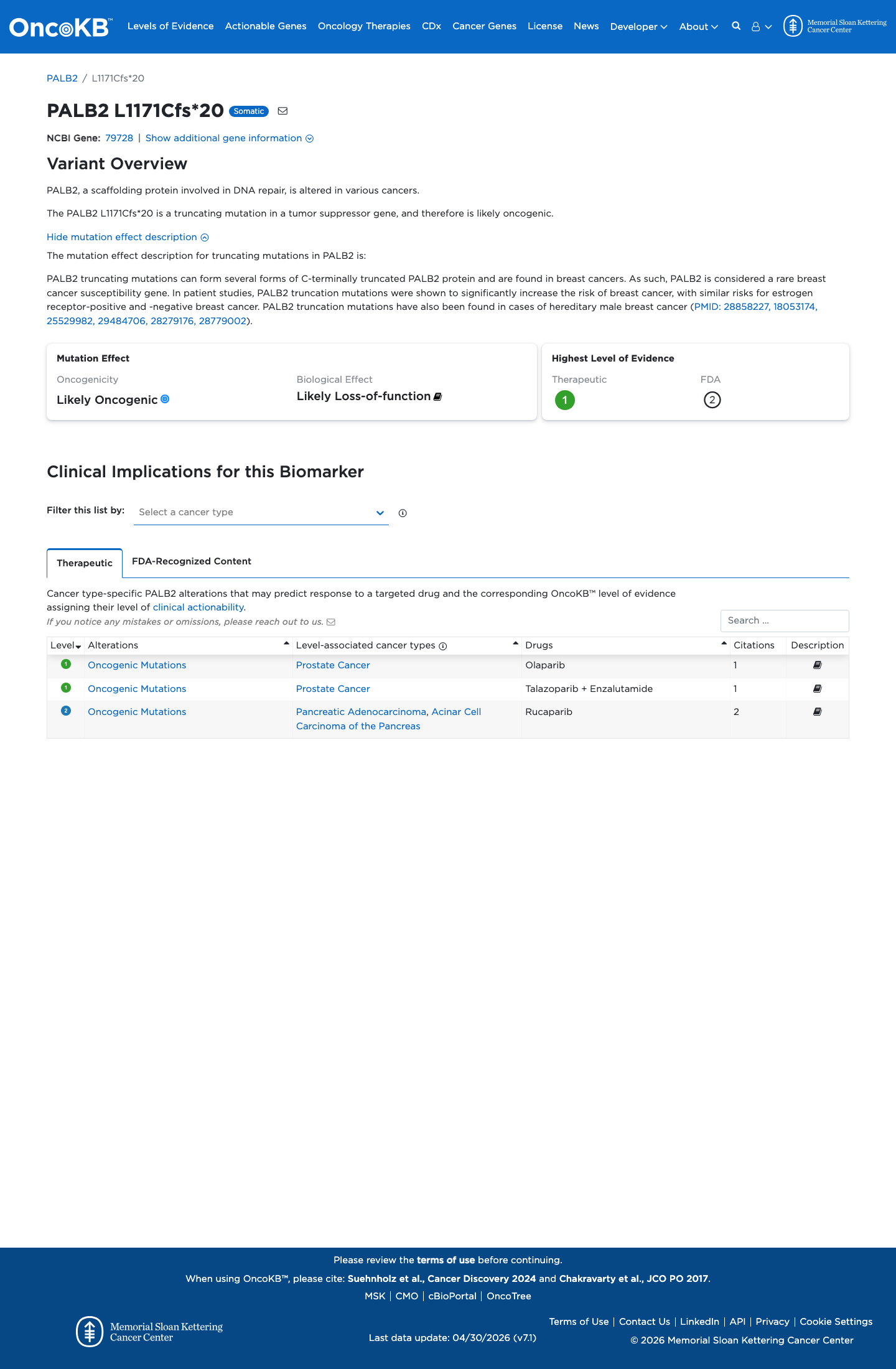
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗PALB2 loss of function is an established disease mechanism, and the PALB2 specification states that variants predicted to escape nonsense-mediated decay but disrupting the indispensable C-terminal WD40 domain can still receive full PVS1; this terminal frameshift alters that WD40 tail.
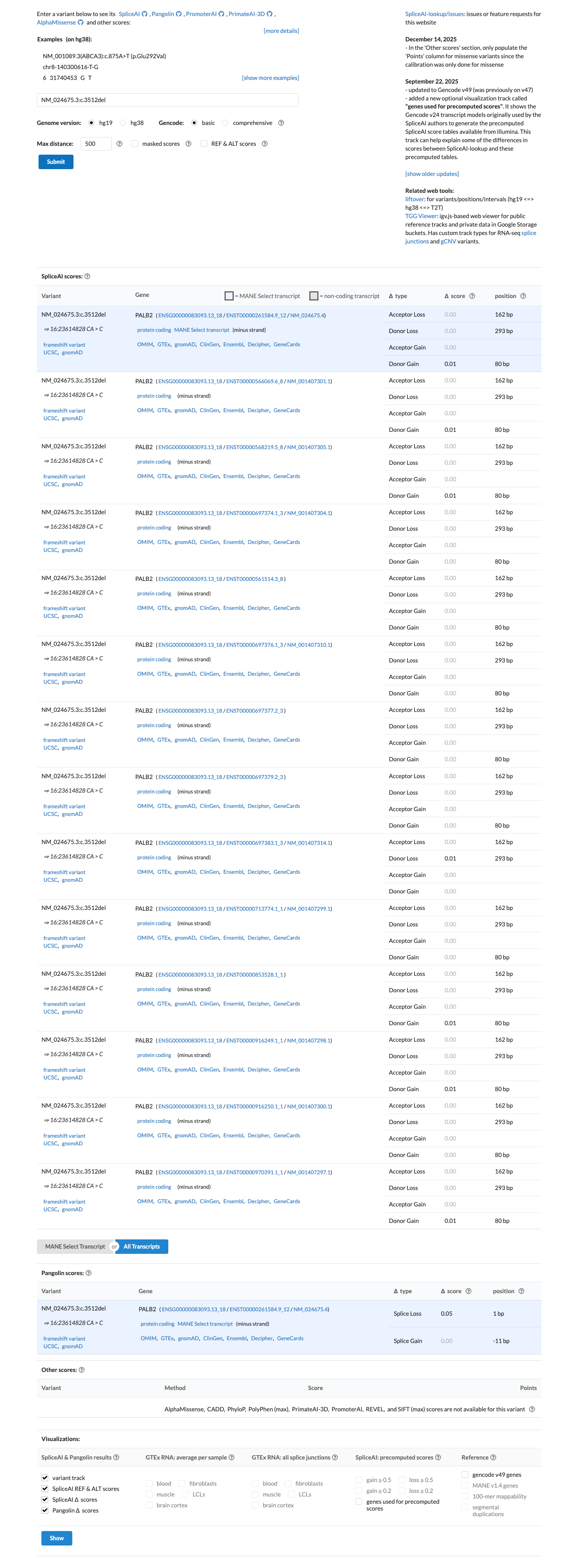
cspec ↗SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01, which is below the PALB2 PP3 threshold of 0.2 and within the BP4 splice threshold of 0.1, although that splice prediction does not remove concern for the protein-disrupting frameshift effect.
spliceai ↗ cspec ↗