Classification rationale
1
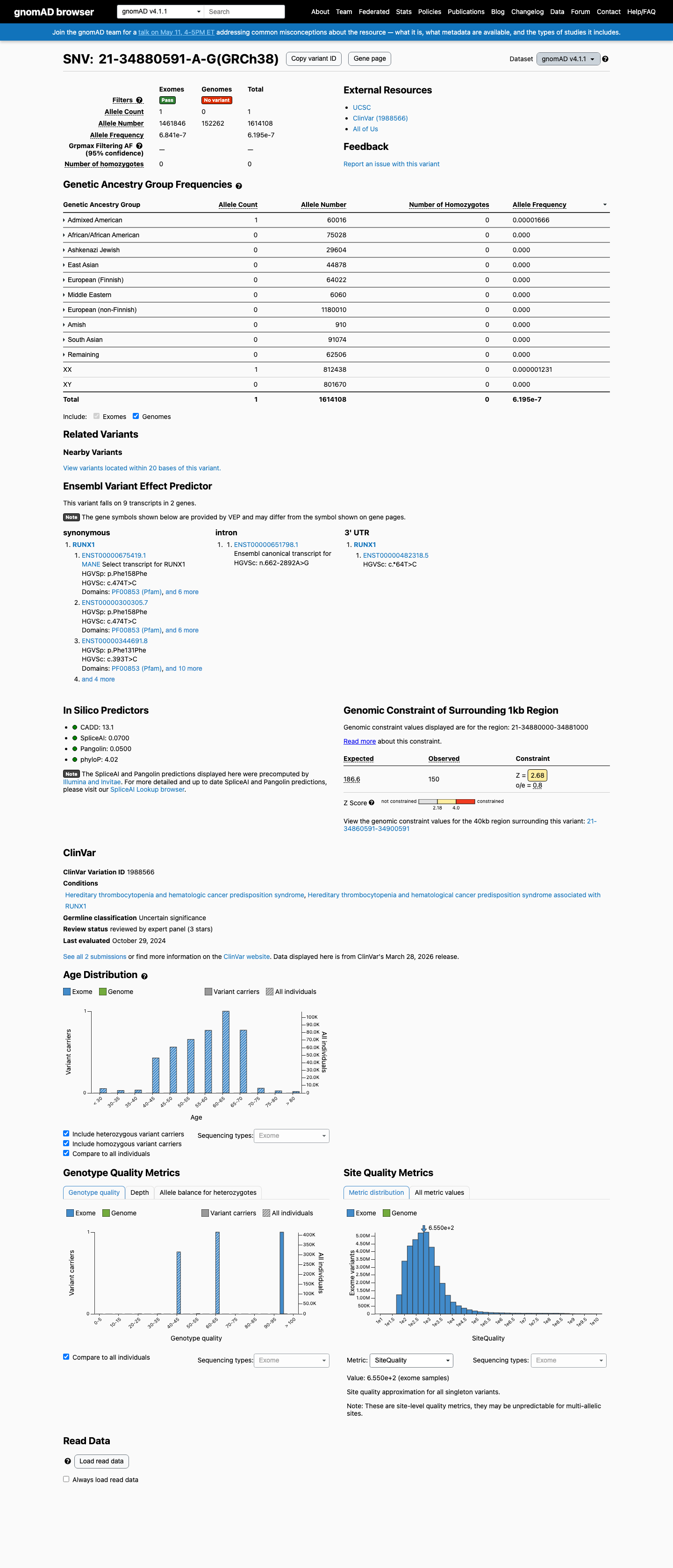
The RUNX1 NM_001001890.2:c.393T>C (p.(Phe131=)) variant has been reported in ClinVar, where the ClinGen Myeloid Malignancy Variant Curation Expert Panel classified the equivalent RUNX1 transcript representation as uncertain significance and one clinical laboratory classified it as likely benign.
clinvar ↗2
This variant is absent from gnomAD v2.1 and is present at very low frequency in gnomAD v4.1 (1/1614108 alleles; AF 6.20e-07), with the highest observed subpopulation frequency of 1.67e-05 remaining below the RUNX1 PM2_Supporting threshold of 0.00005.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
Computational splicing prediction shows no significant splice impact, with a SpliceAI maximum delta score of 0.07, which supports BP4 and BP7 and does not meet the RUNX1 PP3 threshold.
spliceai ↗ cspec ↗