The PALB2 c.2014G>C (p.Glu672Gln; p.E672Q) variant has been reported in ClinVar as Benign by the ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel, with additional benign and likely benign clinical laboratory submissions.
clinvar ↗This variant is common in population databases; in gnomAD v4.1 the allele frequency is 2.71302% with a grpmax filtering allele frequency of 5.70171%, and the highest observed population frequency is 15.35088% in the Amish population, which is well above the PALB2 BA1 threshold of 0.1%.
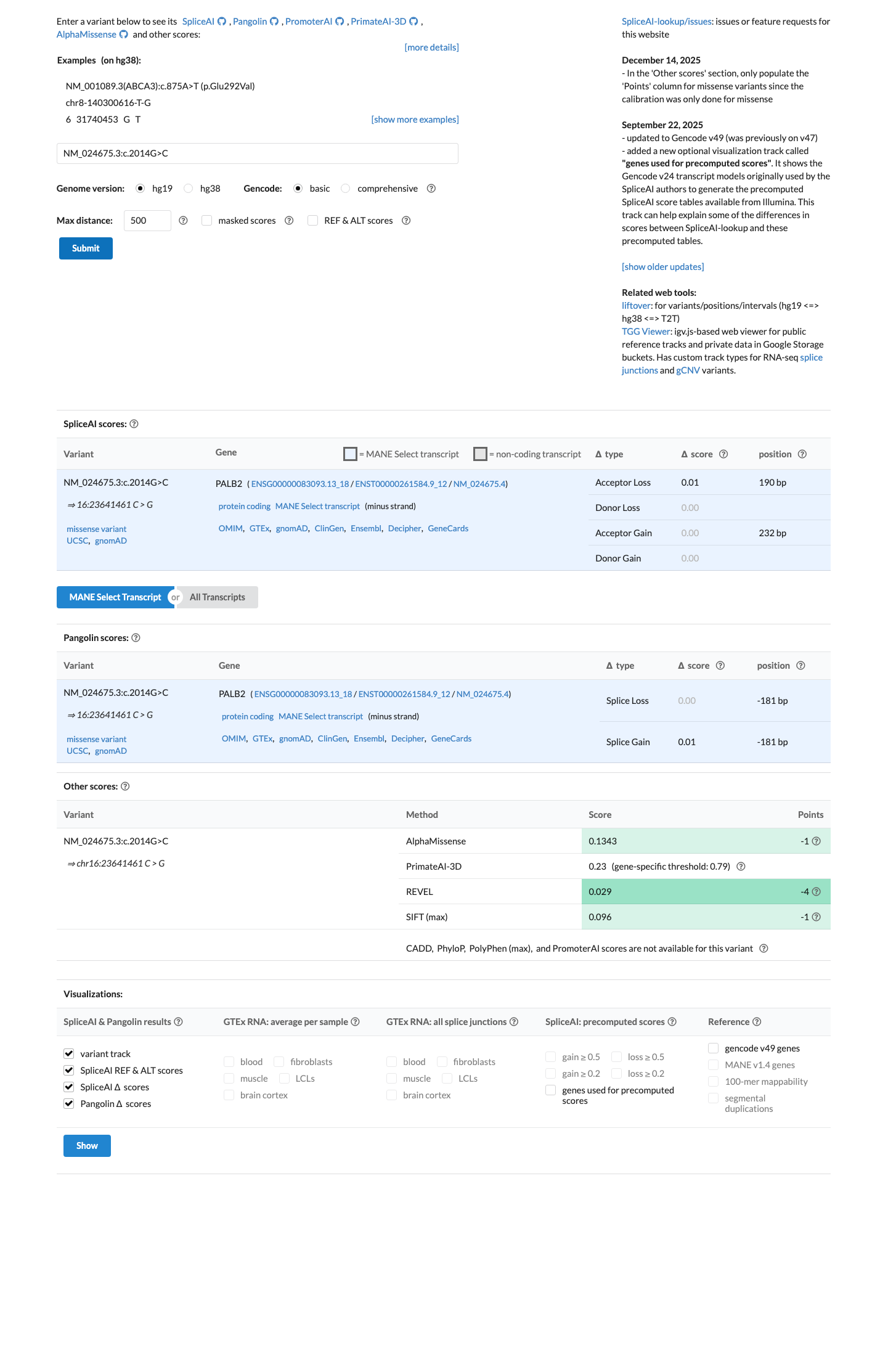
gnomad_v4 ↗ cspec ↗Computational data do not support a deleterious effect: SpliceAI predicts no significant splice impact with a max delta score of 0.01, REVEL is 0.029, and BayesDel is -0.736359; however, the PALB2 VCEP specification does not apply PP3 or BP4 to missense variants.
spliceai ↗ cspec ↗