Classification rationale
1
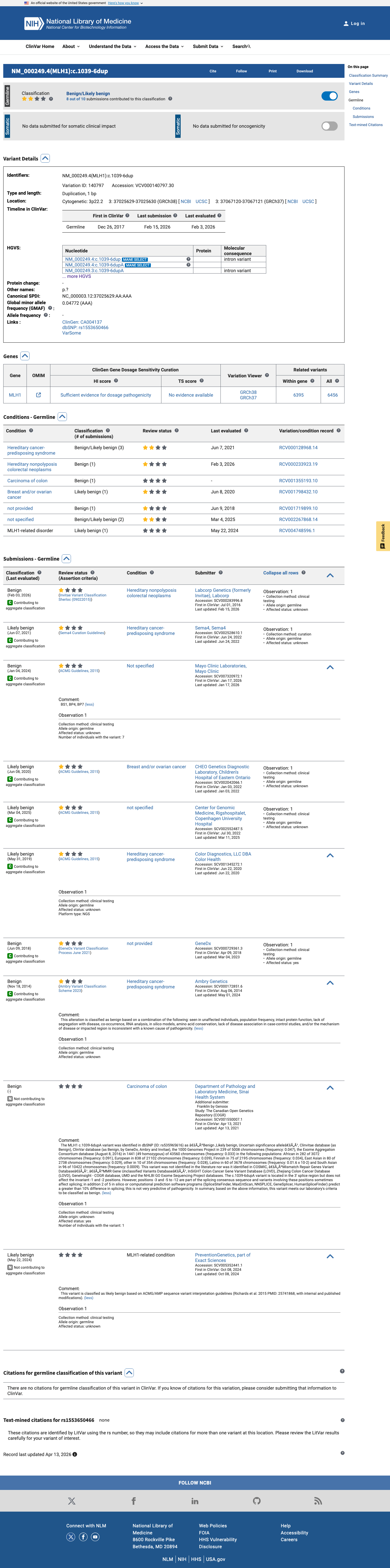
The MLH1 c.1039-6dup (p.?) variant has been reported in ClinVar as benign by 5 clinical laboratories and likely benign by 4 clinical laboratories.
clinvar ↗2
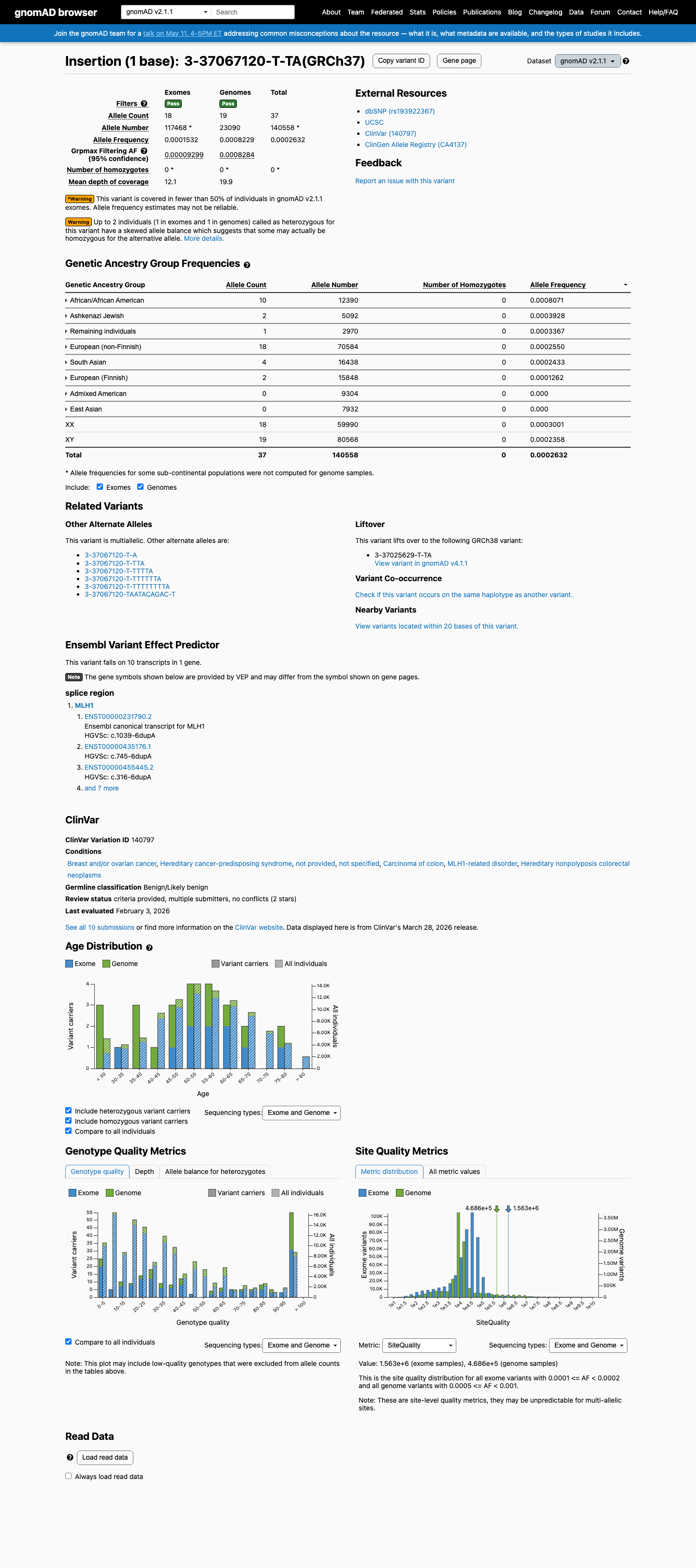
This variant is present in population databases; in gnomAD v4.1 the overall allele frequency is 0.000530357 and the grpmax filtering allele frequency is 0.00117106, which is above the MLH1 VCEP BA1 threshold of 0.001.
gnomad_v4 ↗ cspec ↗3
SpliceAI predicts no significant splice effect, with a maximum delta score of 0.01, which is below the BP4 no-impact threshold of 0.1 and below the PP3 splice-defect threshold of 0.2 in the MLH1 VCEP rules.
spliceai ↗ cspec ↗