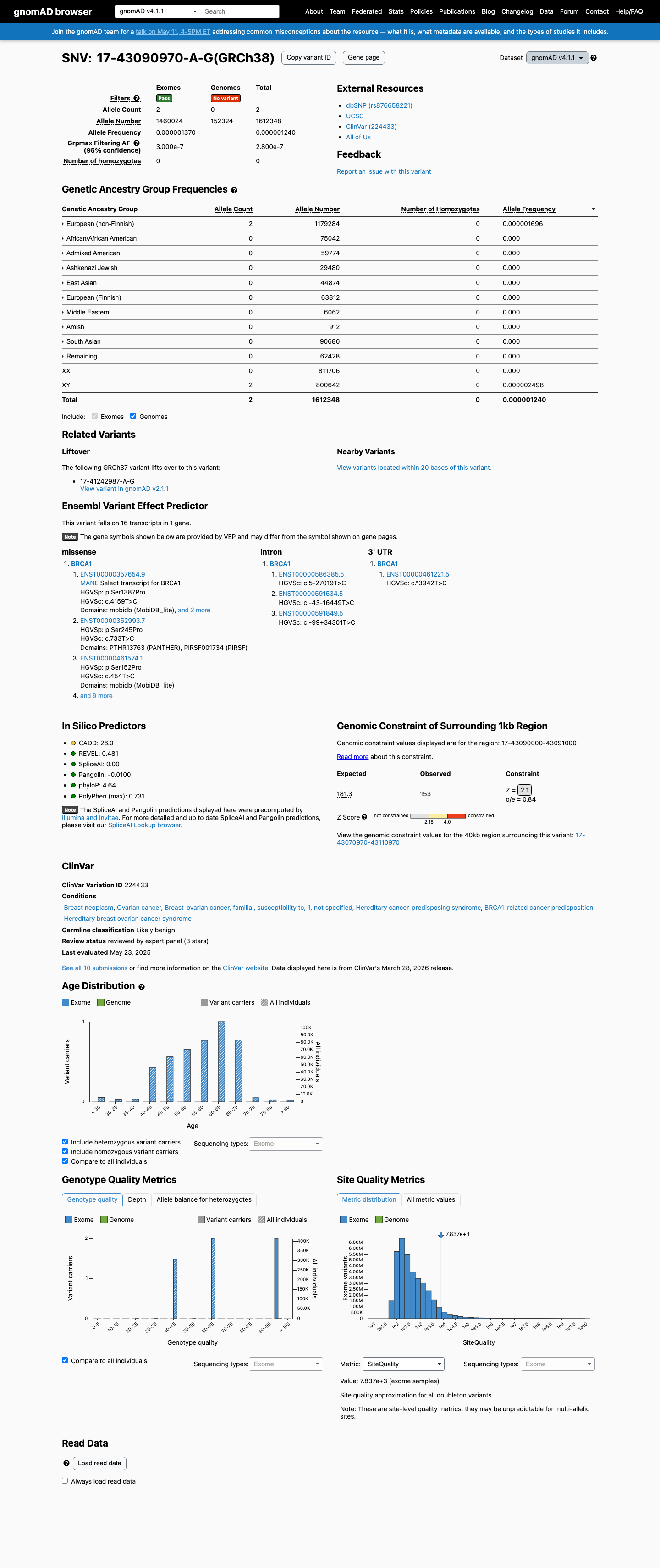
The BRCA1 c.4159T>C (p.Ser1387Pro) variant has been reported in ClinVar, where the ClinGen ENIGMA BRCA1/2 expert panel classifies it as Likely Benign despite conflicting individual submissions.
clinvar ↗This variant is present at very low frequency in population databases, including gnomAD v2.1 at 1/248290 alleles (AF 0.00040%) and gnomAD v4.1 at 2/1612348 alleles (AF 0.00012%; grpmax FAF 2.8e-07), which is too low for BA1 or BS1 and means PM2 is not met because the variant is not absent from controls.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗No exact curated functional assay result for p.(Ser1387Pro) was identified in the ENIGMA BRCA1 functional tables, so PS3 and BS3 were not applied from the available functional evidence.
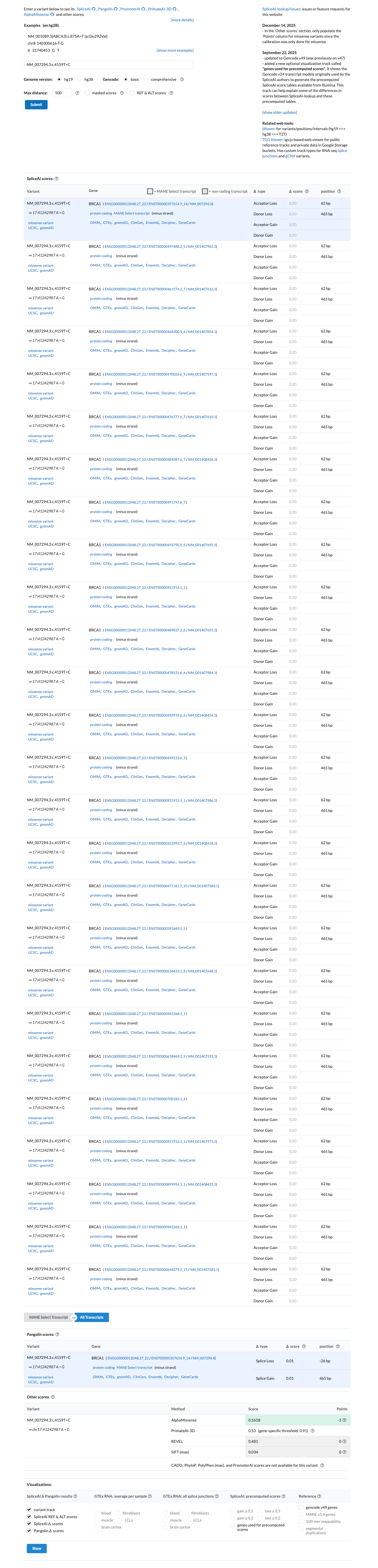
In silico data support a benign interpretation under the ENIGMA BRCA1 framework because p.(Ser1387Pro) lies outside the BRCA1 clinically important domains, SpliceAI predicts no splice impact (max delta 0.00), and the missense change therefore meets BP1_Strong; BayesDel is 0.148379 and PP3 is not met.
cspec ↗ spliceai ↗