The PALB2 c.2787_2788dup (p.Asn930IlefsTer6; p.N930Ifs*6) variant has been reported in ClinVar as pathogenic, including an expert panel submission.
clinvar ↗This variant is absent from gnomAD v4.1 and gnomAD v2.1, and its observed population frequency is below the PALB2 PM2_Supporting threshold of 0.000333%.
gnomad_v4 ↗ gnomad_v2 ↗ cspec ↗This frameshift duplication is predicted to create p.(Asn930IlefsTer6), introducing a premature termination codon consistent with loss of function in a gene where loss of function is an established disease mechanism, and it truncates the protein upstream of p.Tyr1183, supporting PVS1 and PM5_Supporting under the PALB2 specification.
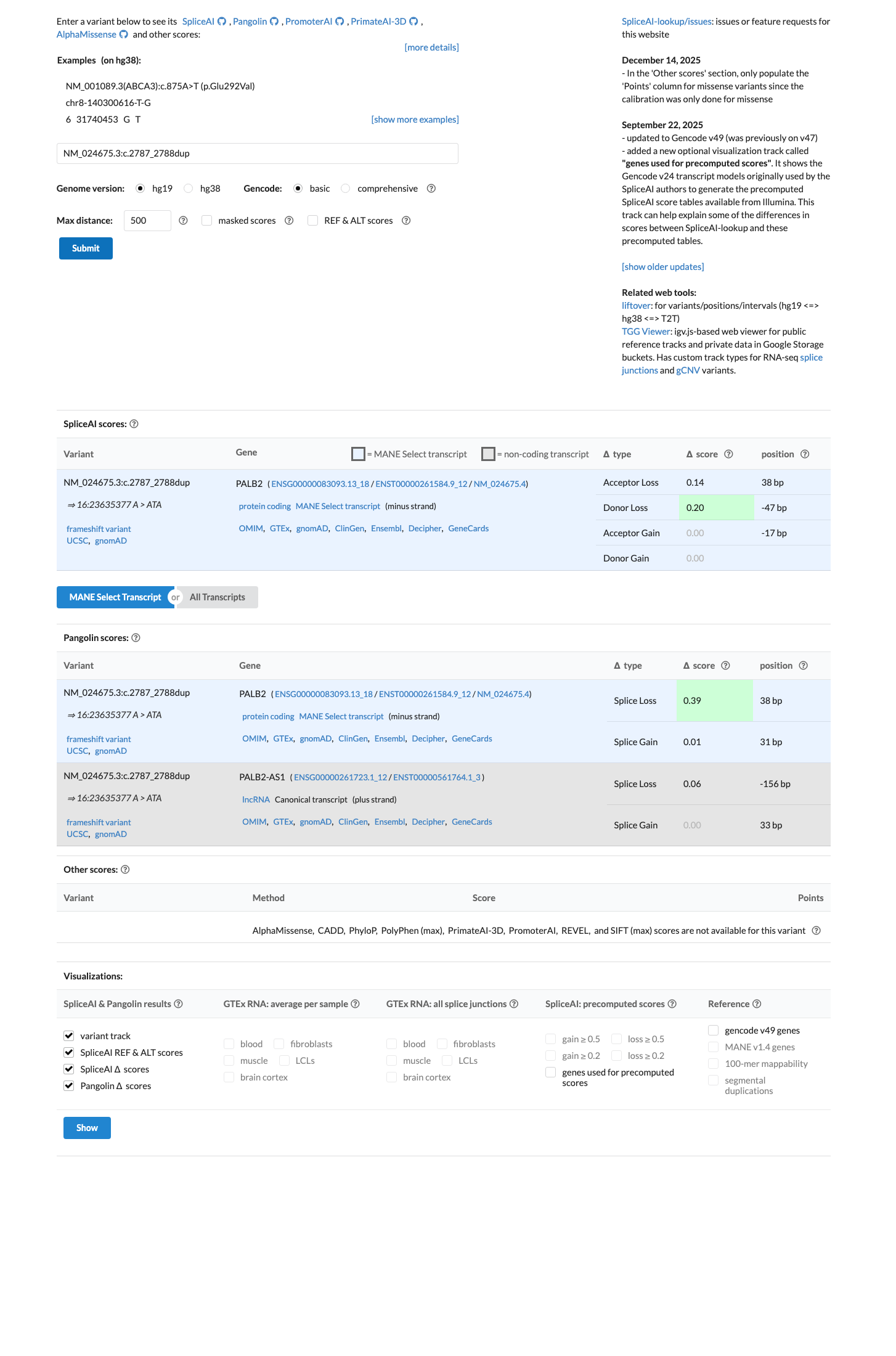
cspec ↗SpliceAI shows a maximum delta score of 0.20, which suggests possible splice impact, but this was not applied as a separate PP3 or BP4 pathogenic criterion for this frameshift variant under the PALB2 rule set.
spliceai ↗ cspec ↗