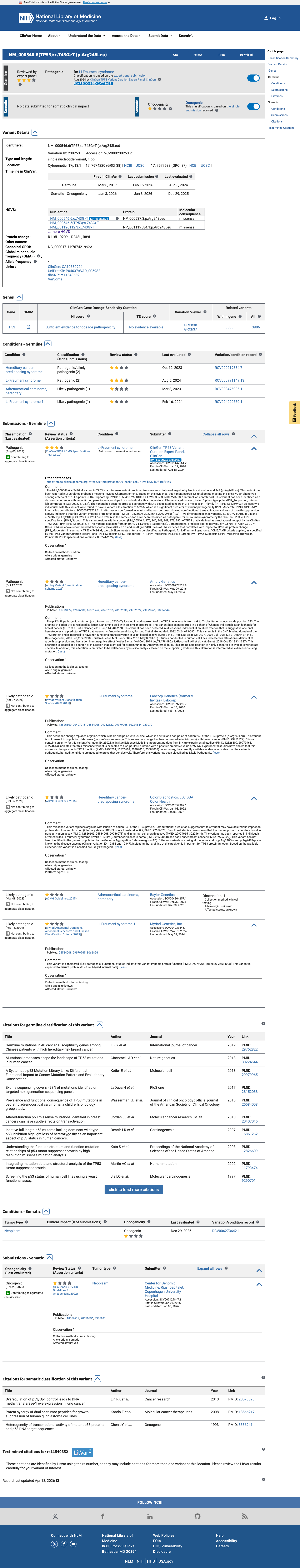
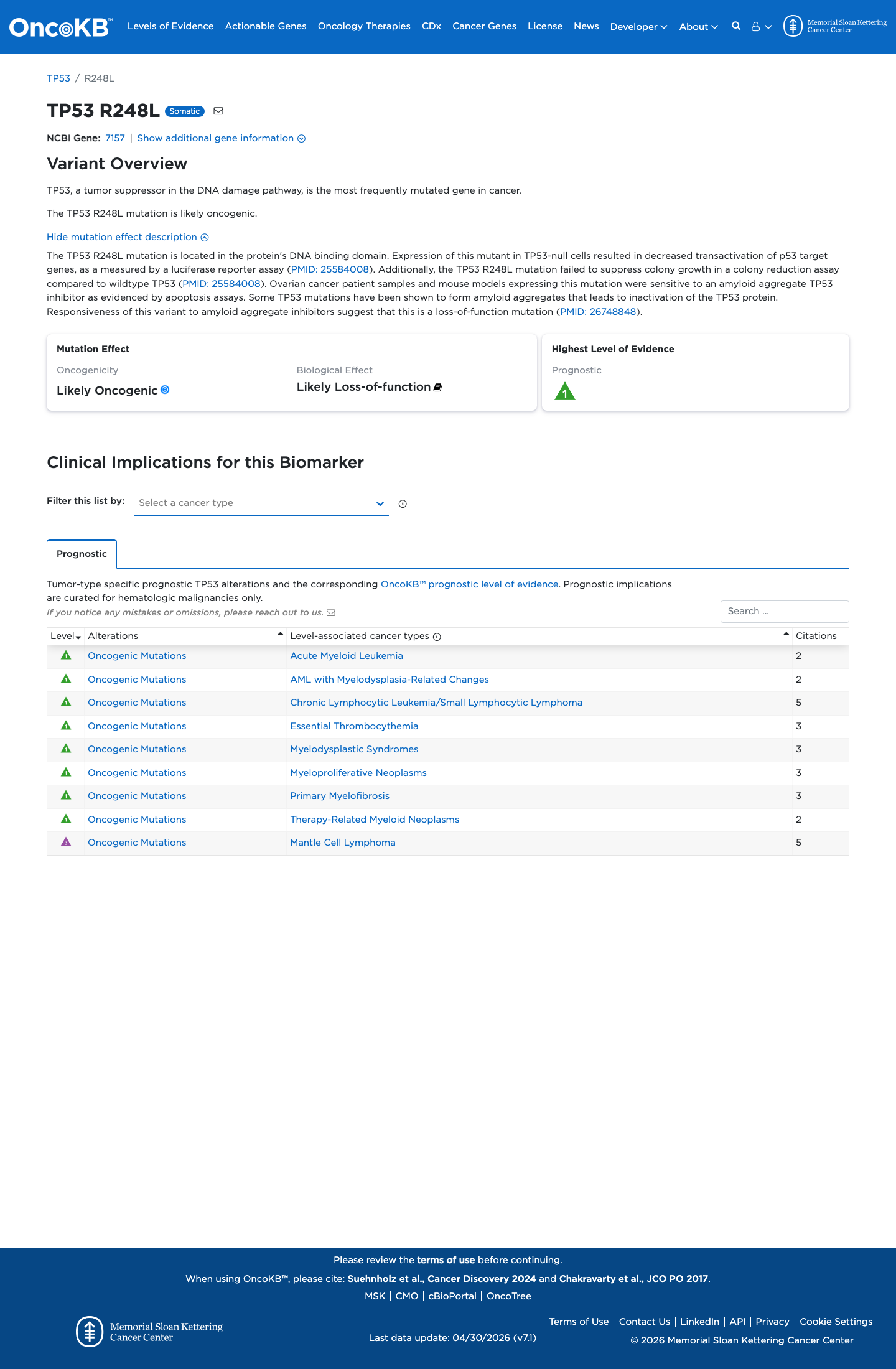
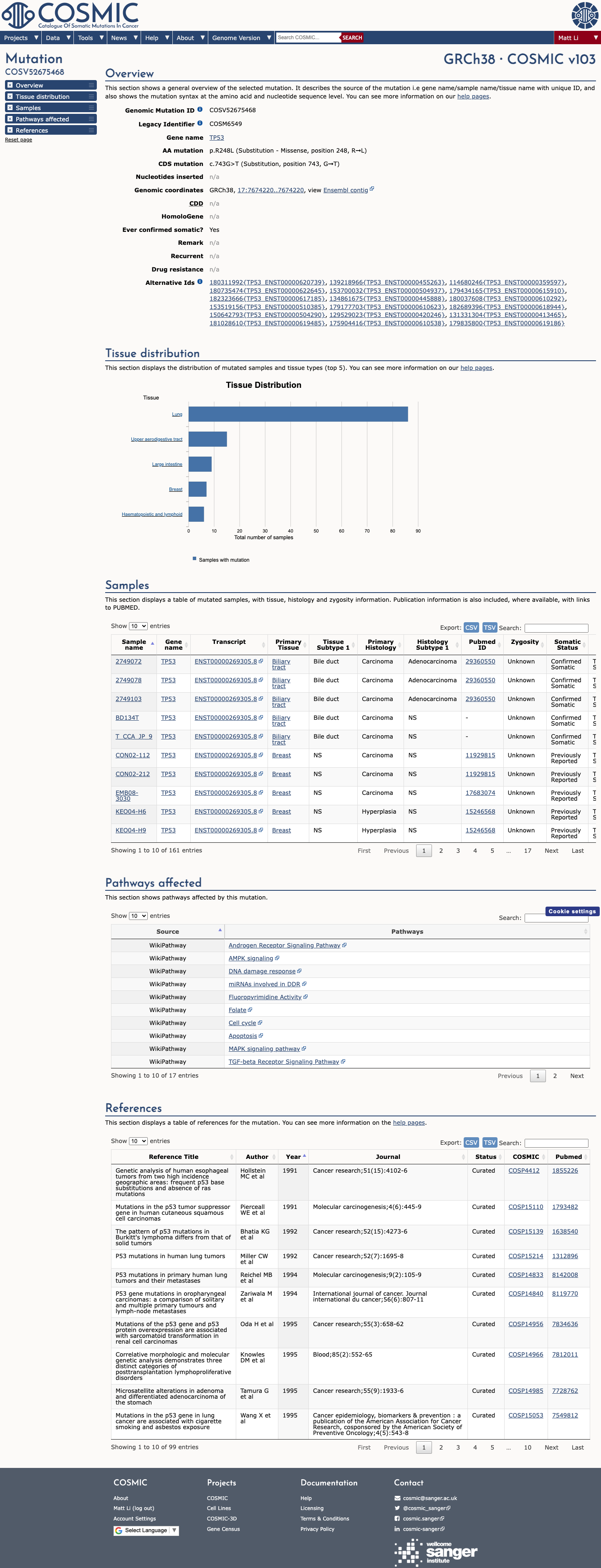
The TP53 c.743G>T (p.Arg248Leu) variant has been observed in somatic cancers in COSMIC 161 times and has been reported in ClinVar with an expert panel pathogenic classification.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity under the TP53 VCEP PM2_Supporting threshold.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In the TP53 VCEP functional worksheet, p.Arg248Leu is summarized as non-functional with loss-of-function results across eligible assays and is pre-assigned PS3, supporting a damaging effect on p53 function.
PMID:12826609 ↗In the TP53 VCEP bioinformatic worksheet, this missense change is pre-assigned PP3_moderate with Align-GVGD class C65 and BayesDel 0.570318, while SpliceAI predicts no meaningful splice impact with a maximum delta score of 0.01; REVEL is also high at 0.96.
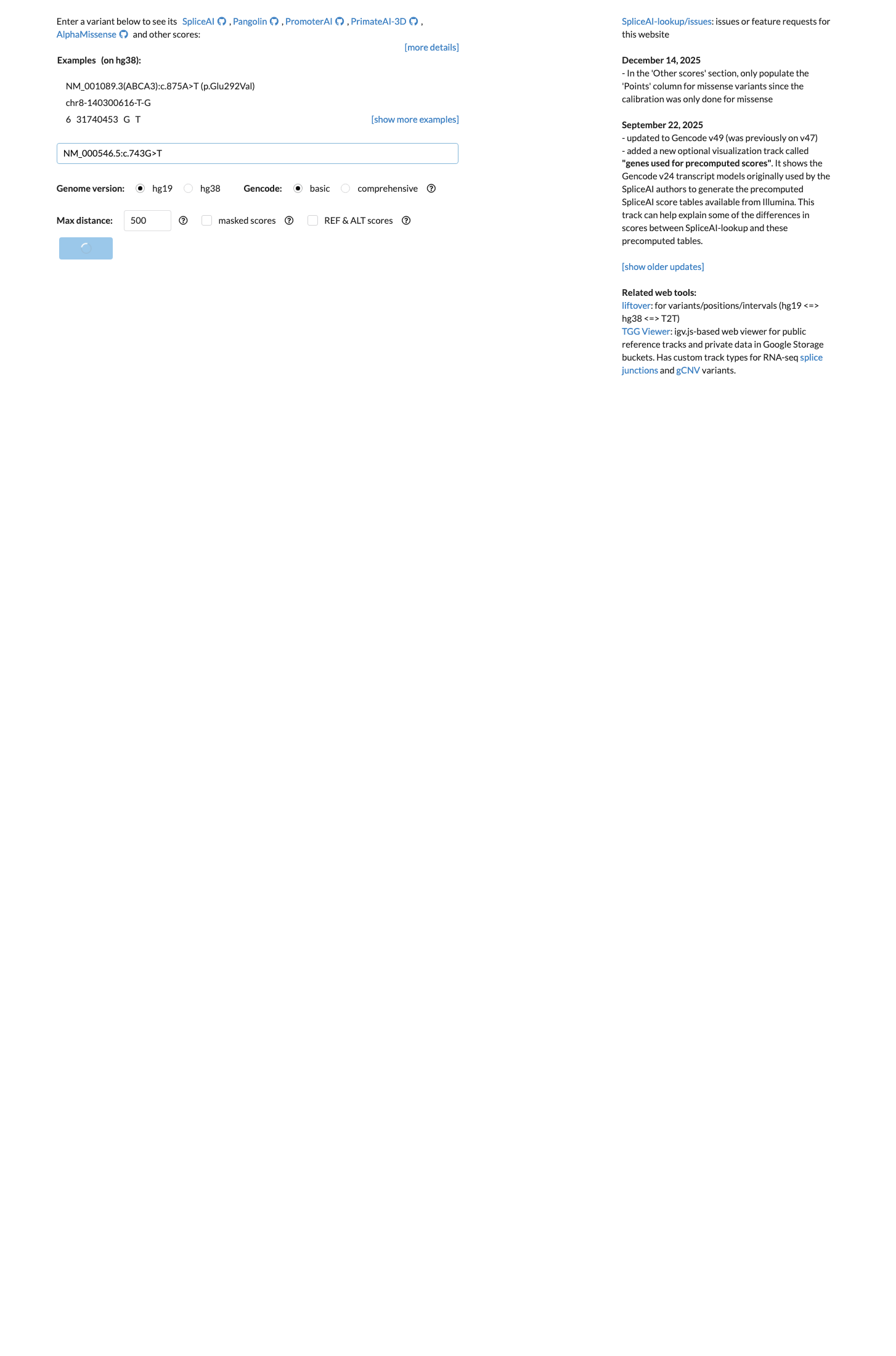
spliceai ↗