Classification rationale
1
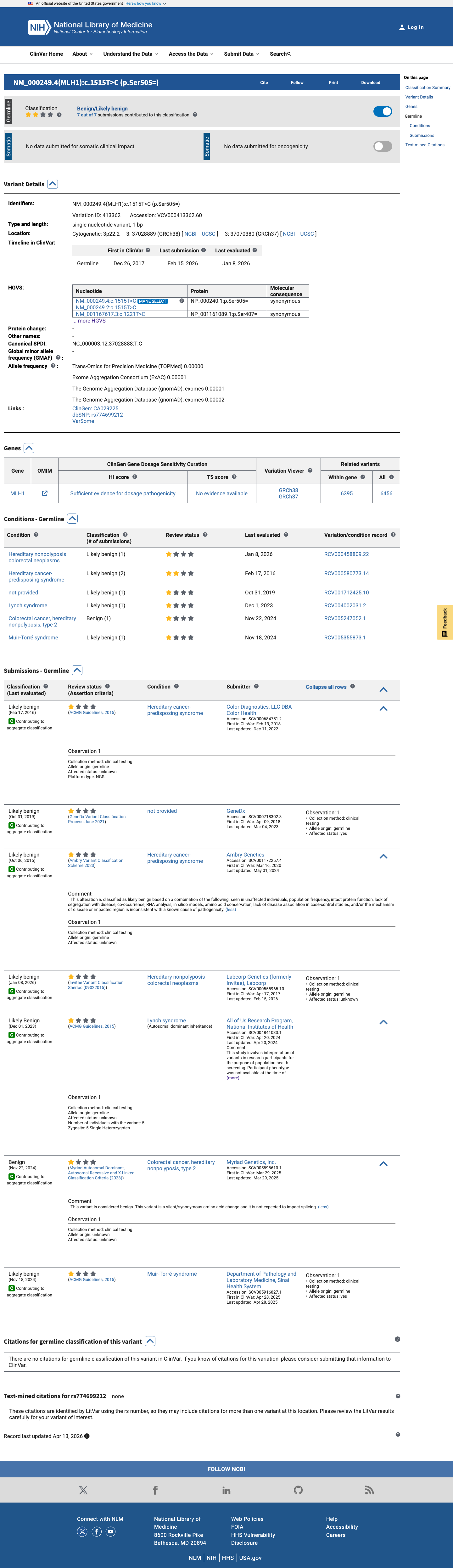
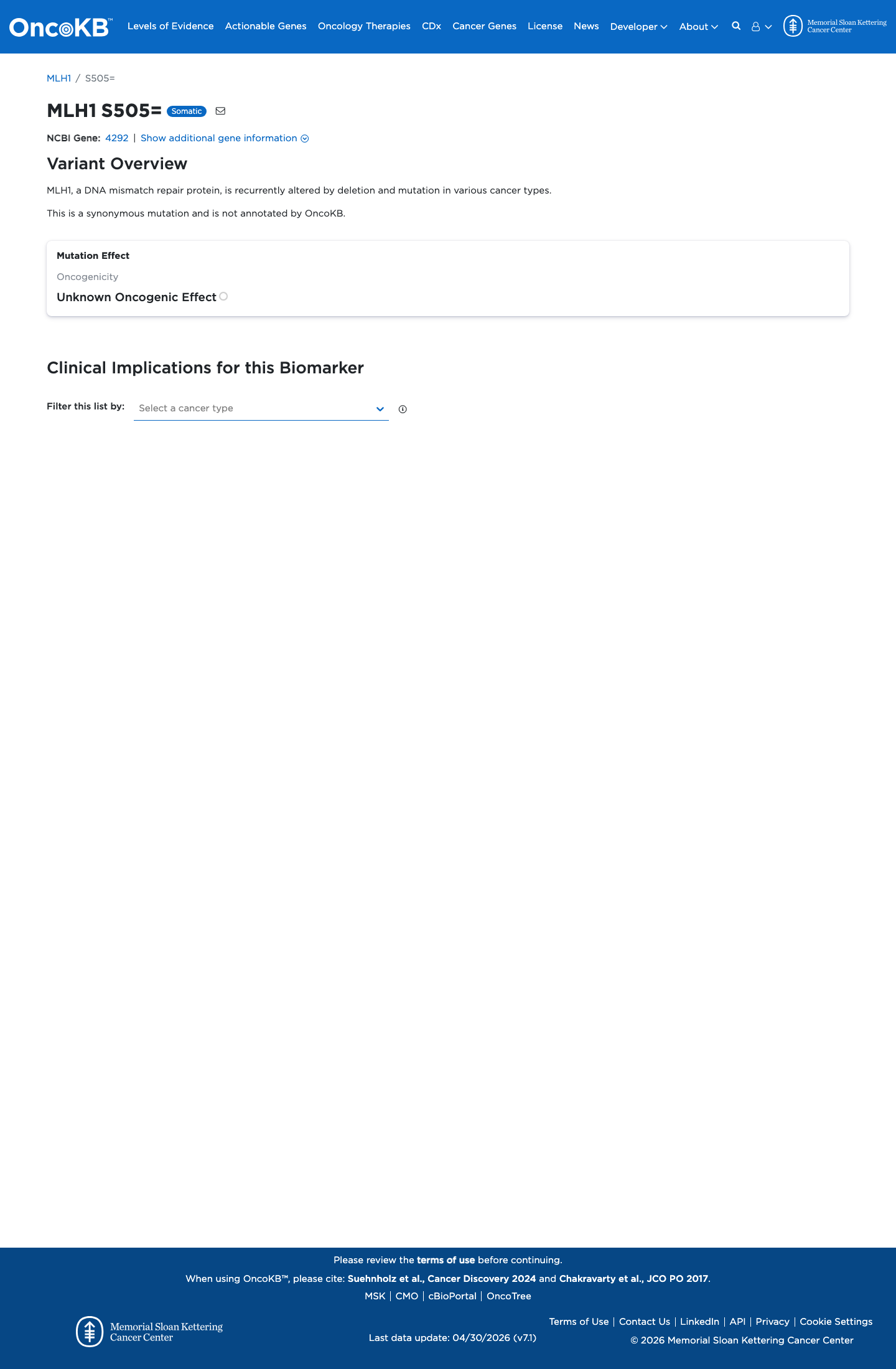
The MLH1 c.1515T>C (p.Ser505=) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar predominantly as likely benign or benign.
clinvar ↗2
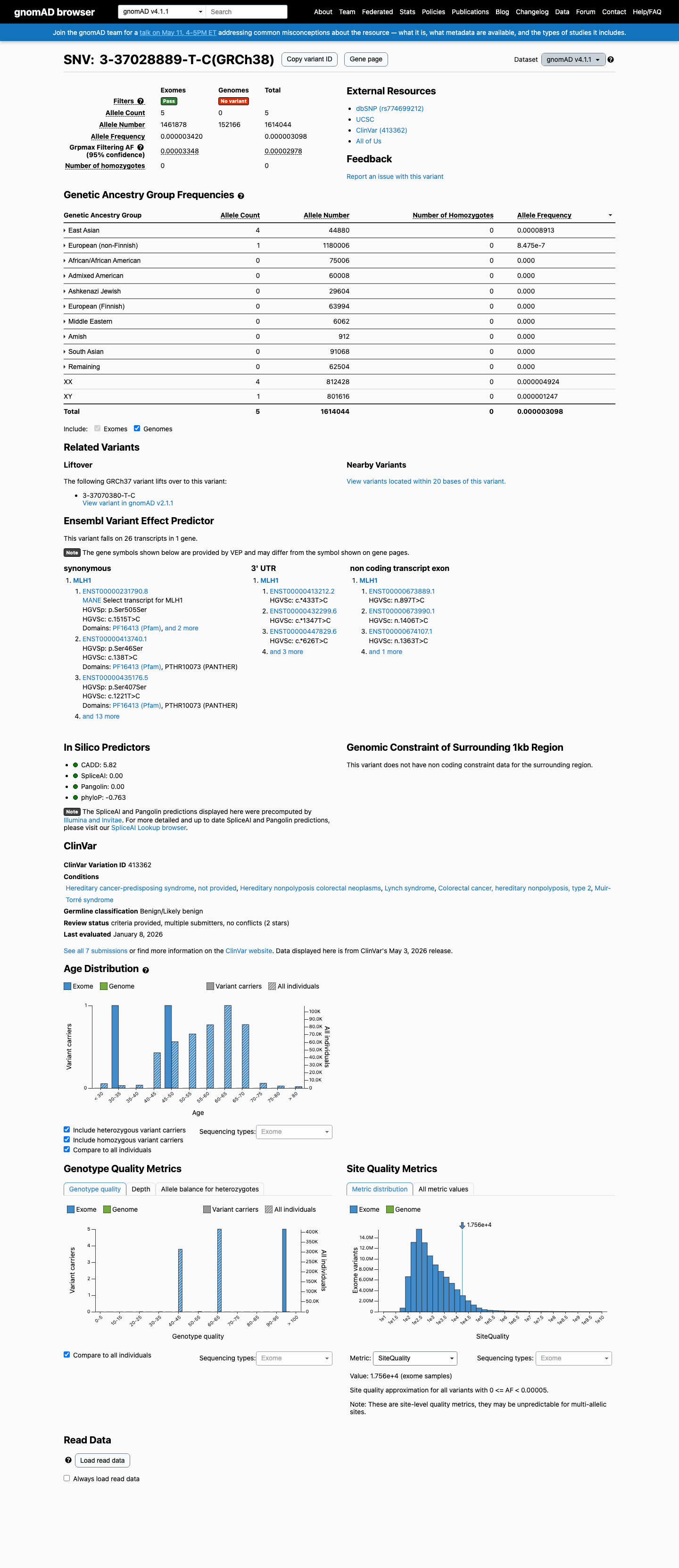
This variant is present in gnomAD, including 5/1614044 alleles overall in v4.1 (AF 0.00000310) and 4/44880 East Asian alleles (AF 0.0000891), with a grpmax filtering allele frequency of 0.0000298 that is below the BA1 and BS1 thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
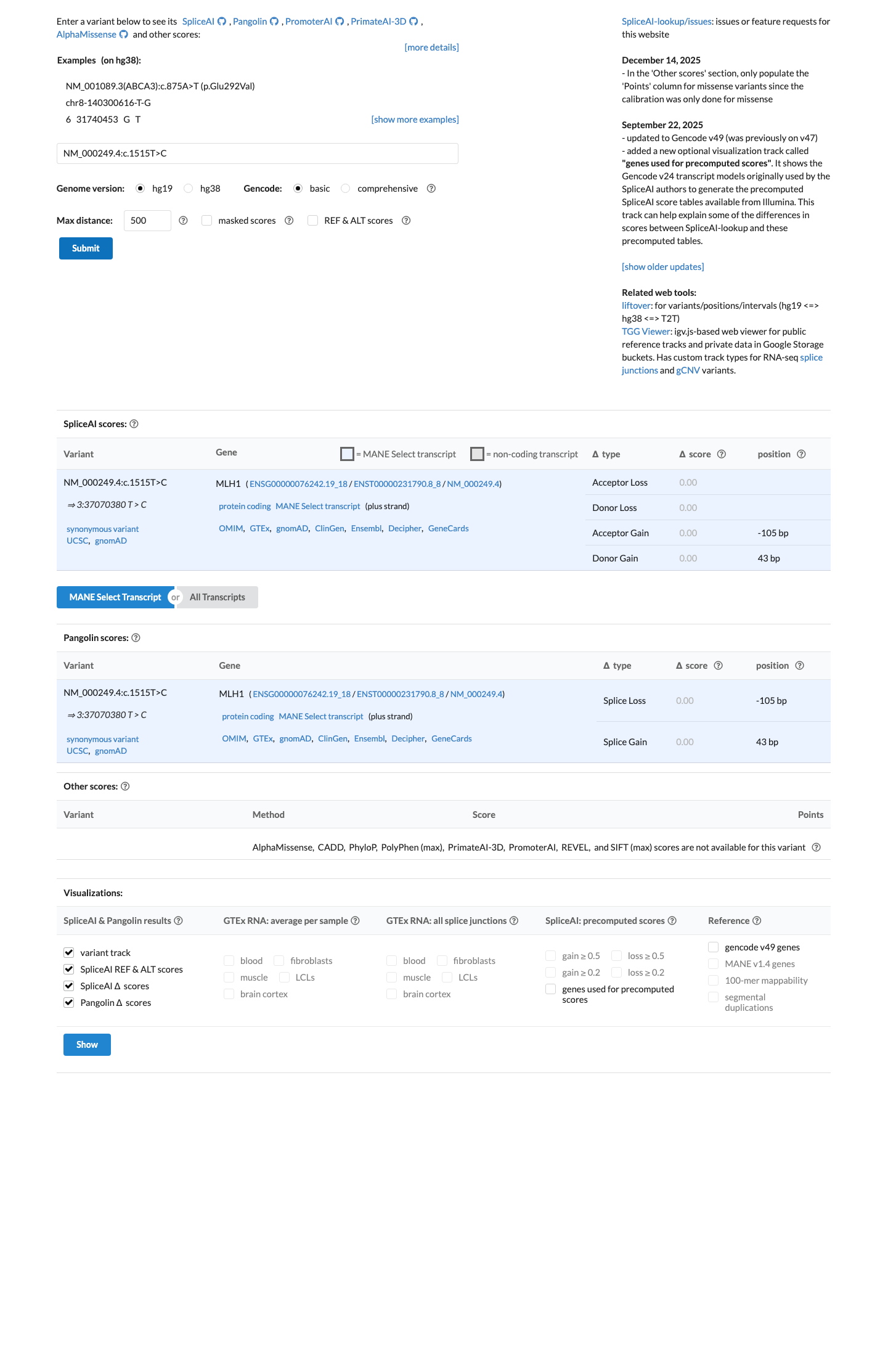
In silico splicing analysis predicts no significant splice impact with a SpliceAI max delta score of 0.00, supporting BP4 for a synonymous variant; no MLH1 HCI prior entry for c.1515T>C was identified, and the available REVEL score was 0.343.
spliceai ↗ cspec ↗