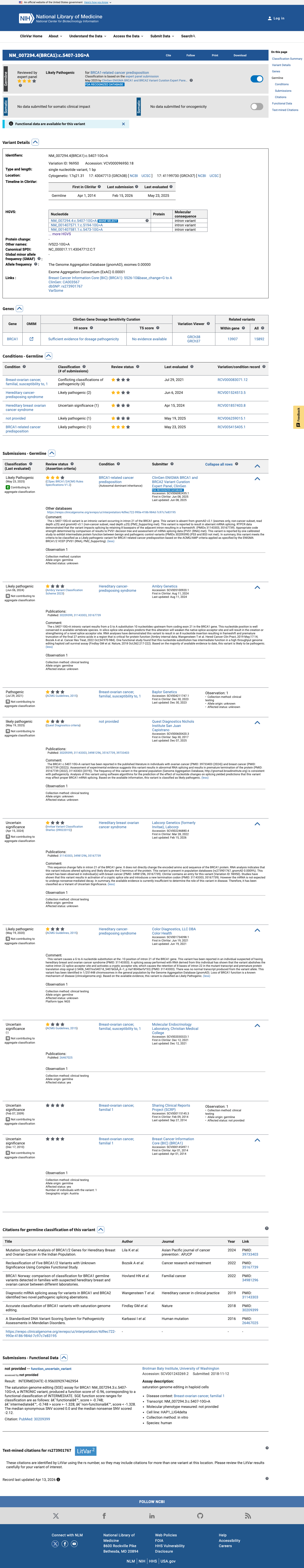
The BRCA1 c.5407-10G>A (p.?) variant has not been observed in somatic cancers in COSMIC and has been reported in ClinVar, where the current aggregate classification is Likely Pathogenic with expert-panel review.
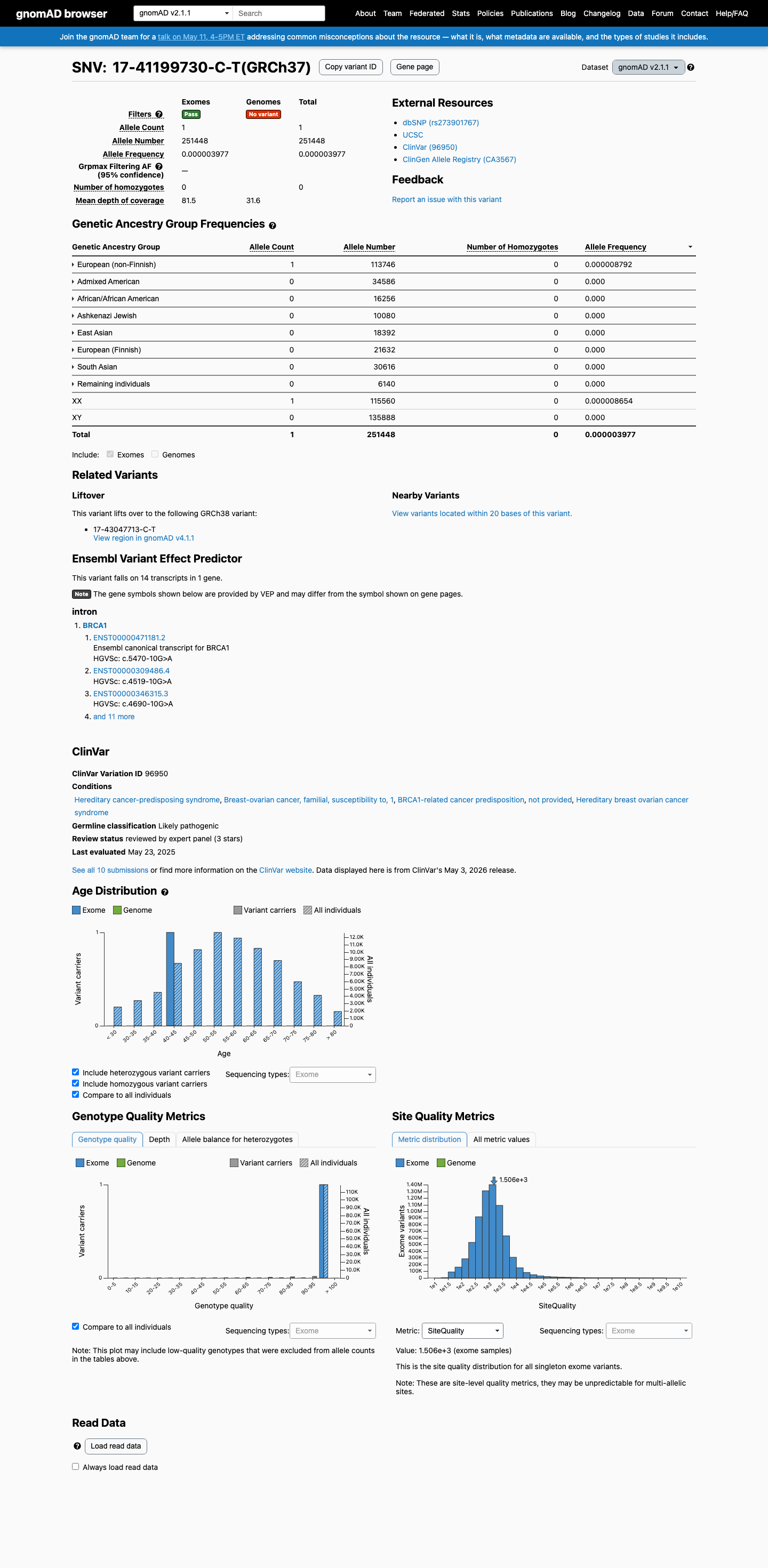
clinvar ↗This variant is absent from gnomAD v4.1 and is present only once in gnomAD v2.1 (1/251448 alleles; AF 3.98e-06), which is too low for benign population criteria but does not satisfy the BRCA1 ENIGMA requirement for complete absence from gnomAD for PM2.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In published BRCA1 functional and RNA studies, this variant showed retention of 8 nucleotides from intron 22 and partial functional impact rather than a clearly damaging or clearly benign result, so the ENIGMA BRCA1 functional tables indicate that PS3 and BS3 are not met.
PMID:30209399 ↗ PMID:31143303 ↗SpliceAI predicts a strong splice effect for this variant with a maximum delta score of 1.00, which exceeds the BRCA1 ENIGMA PP3 threshold of 0.20 for intronic variants outside the native +/-1,2 splice sites and supports PP3 at Supporting strength.
spliceai ↗ cspec ↗