Classification rationale
1
The PALB2 c.154G>A (p.Val52Ile) variant has been reported in ClinVar with conflicting single-submitter interpretations of uncertain significance and likely benign.
clinvar ↗2
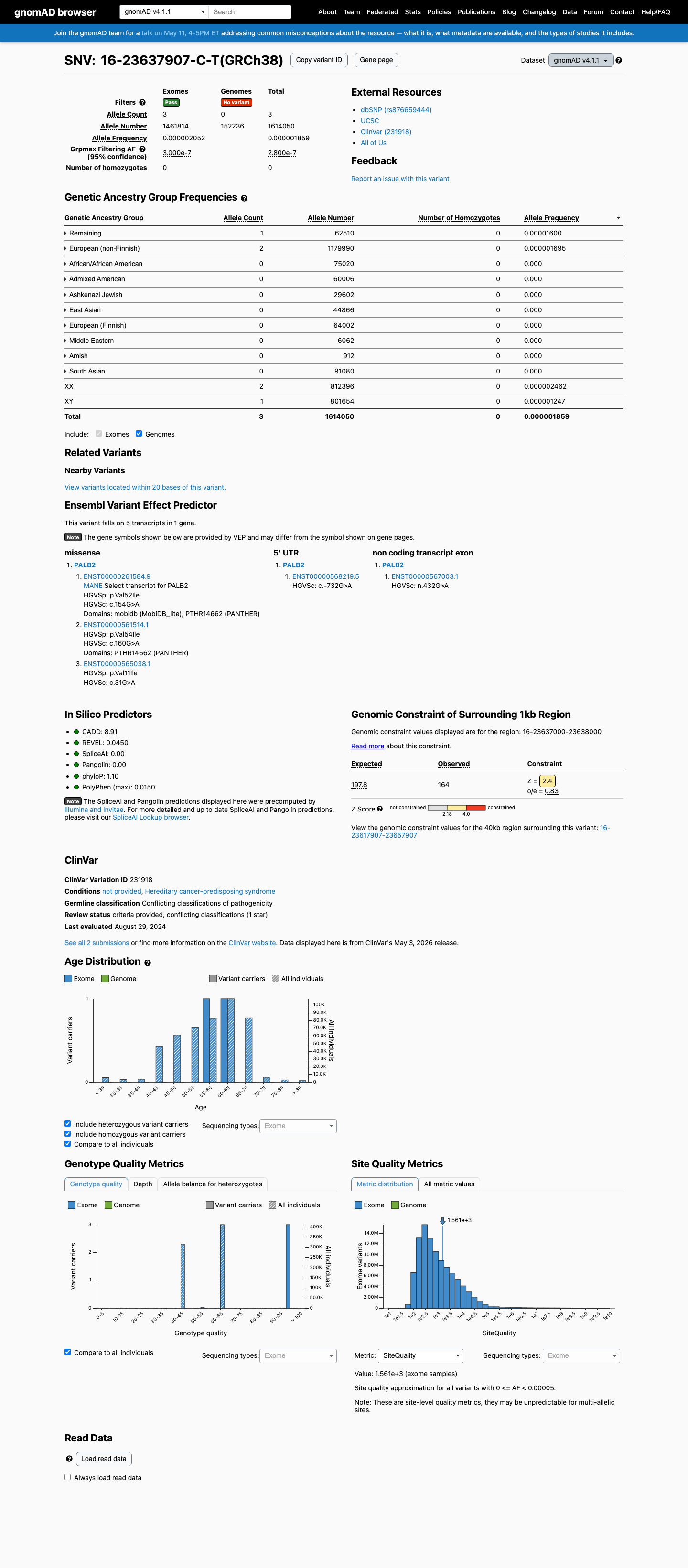
This variant is absent from gnomAD v2.1 and is present at very low overall frequency in gnomAD v4.1 (3/1,614,050 alleles; 0.00019%), but the highest observed subpopulation frequency is 0.00160% (1/62,510 alleles), which is above the PALB2 PM2 threshold exception and remains below the BS1 and BA1 population thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
SpliceAI predicts no significant splice impact (max delta score 0.01), and REVEL (0.045) and BayesDel (-0.518425) are low; however, the PALB2 VCEP does not use PP3 or BP4 for missense variants, while BP1 is applicable to all PALB2 missense variants.
spliceai ↗ cspec ↗