Classification rationale
1
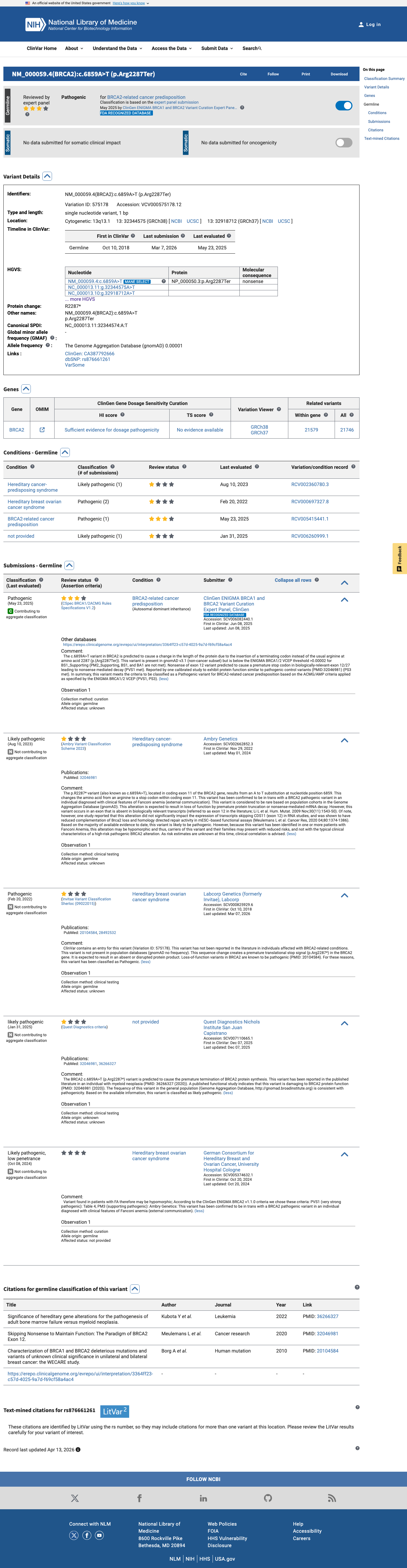
The BRCA2 c.6859A>T (p.Arg2287Ter; p.R2287*) variant has not been observed in COSMIC and has been reported in ClinVar as Pathogenic with expert panel review.
clinvar ↗2
This variant is absent from gnomAD v2.1 and is present once in gnomAD v4.1 (1/1,556,108 alleles; AF 6.43e-07; no homozygotes), consistent with extreme rarity.
gnomad_v2 ↗ gnomad_v4 ↗3
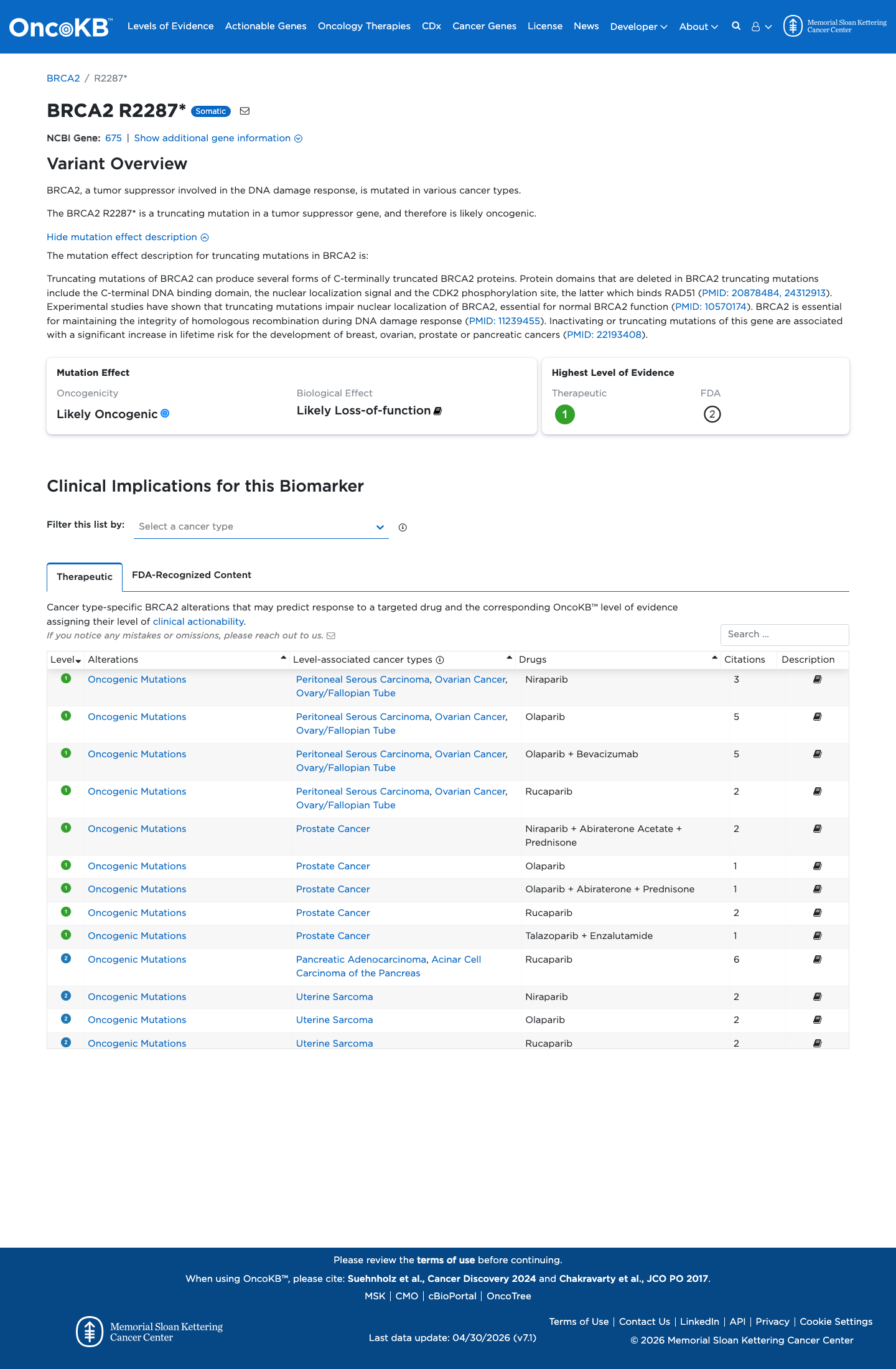
Under the ENIGMA BRCA2 specification, this exon 12 premature termination variant is a null variant in a gene where loss of function is an established disease mechanism, and the exon-level table assigns PVS1 to protein-truncating variants in exon 12.
cspec ↗4
SpliceAI predicts no significant splice impact, with a maximum delta score of 0.10, which does not support PP3 for splice disruption.
spliceai ↗ cspec ↗