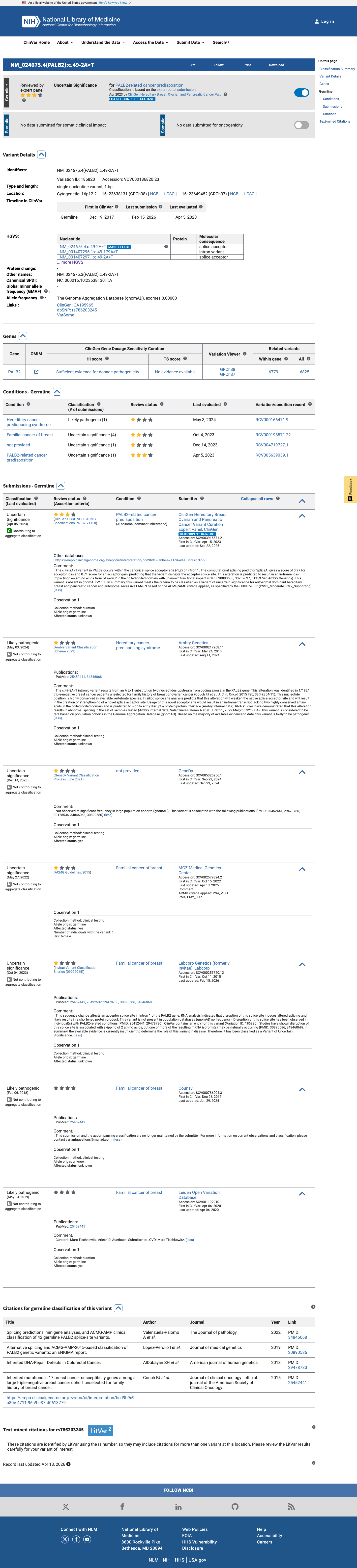
The PALB2 NM_024675.3:c.49-2A>T (NP_078951.2:p.?) variant has been reported in ClinVar, where submissions include likely pathogenic and uncertain significance interpretations, and the ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer expert panel has classified it as uncertain significance.
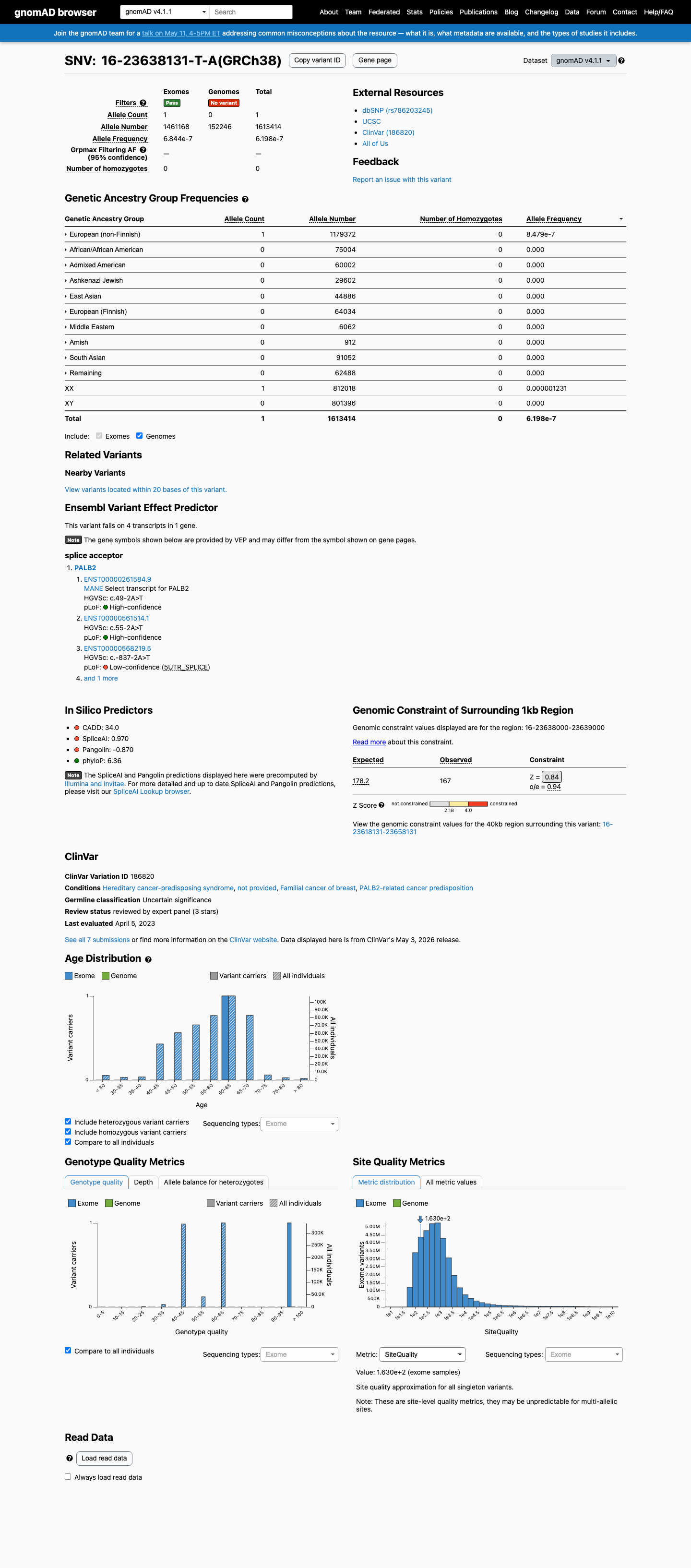
clinvar ↗This variant is absent from gnomAD v2.1 and is present in gnomAD v4.1 at 1/1,613,414 alleles (0.00006%), which is below the PALB2 PM2_Supporting threshold of 0.000333% and well below the BS1 and BA1 population thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗This variant affects a canonical splice acceptor position in a gene for which loss of function is an established disease mechanism, but the exact transcript consequence for this variant remains unresolved in the available evidence.
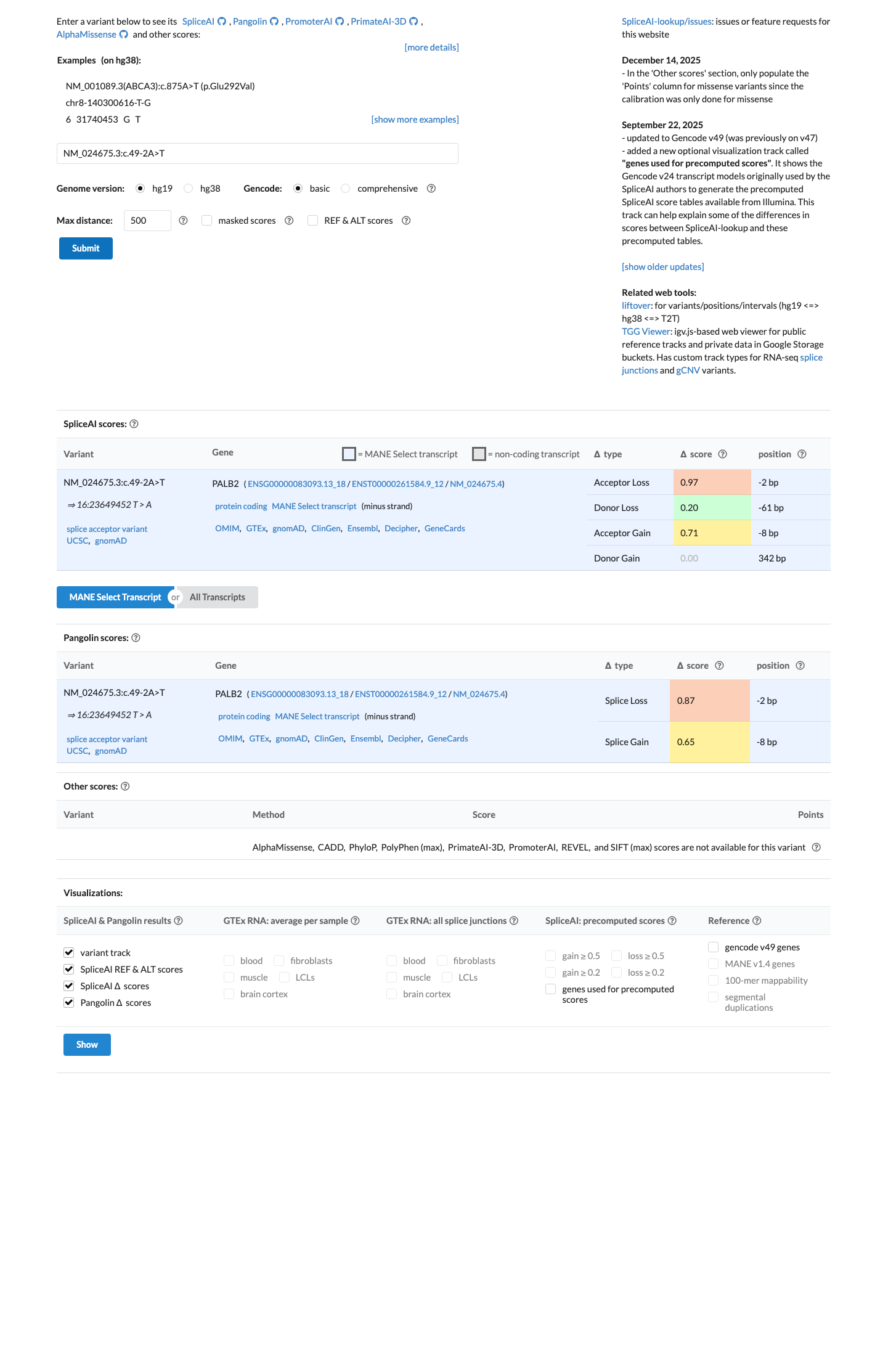
cspec ↗Available computational evidence is mixed: SpliceAI shows a max delta score of 0.00, whereas BayesDel is 0.63, and these data do not independently resolve the splice consequence for this canonical splice-site variant.
spliceai ↗