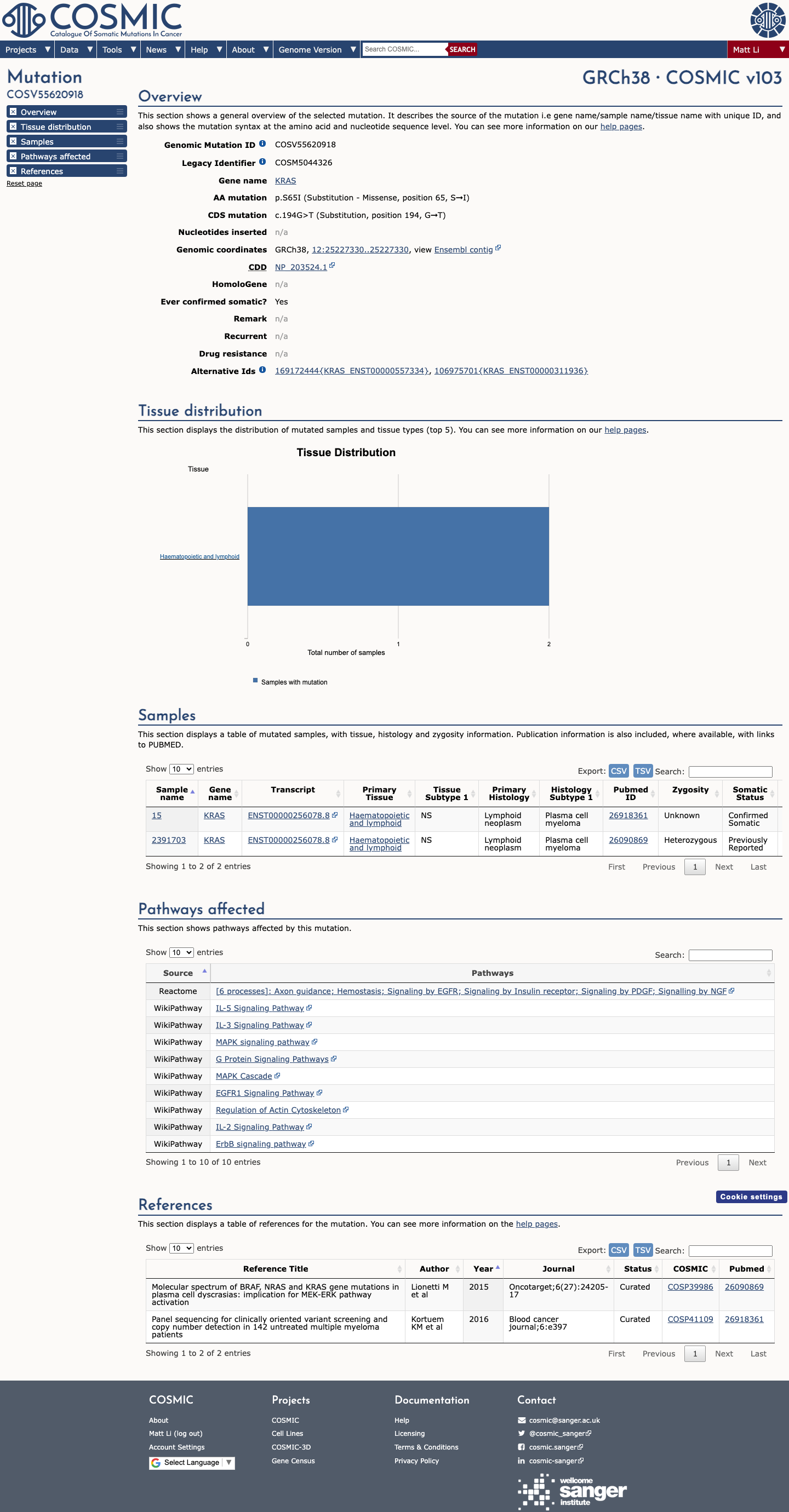
The KRAS c.194G>T (p.Ser65Ile) variant has been reported in ClinVar, including a Likely pathogenic expert-panel classification and additional pathogenic and uncertain-significance submissions.
clinvar ↗This variant is absent from gnomAD v2.1 and absent from gnomAD v4.1, supporting PM2 at supporting strength under the KRAS RASopathy specification.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Available KRAS VCEP functional-study materials list approved assay types for this gene, but no variant-specific approved functional result for p.Ser65Ile was identified in the retrieved evidence, so PS3 was not applied.
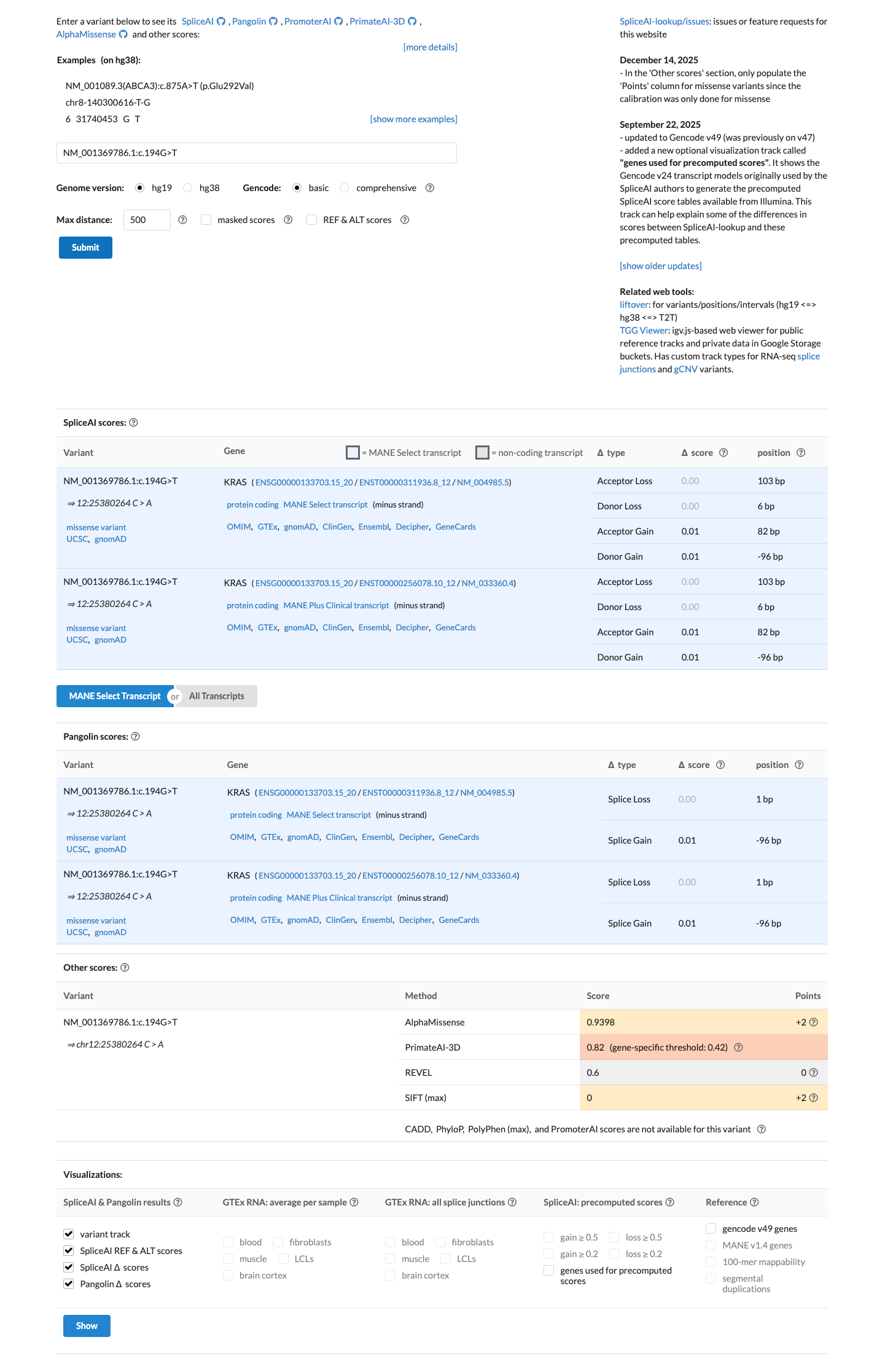
oncokb ↗Computational data do not meet the KRAS RASopathy missense thresholds for either PP3 or BP4: REVEL is 0.60, which is below the PP3 threshold of at least 0.7 and above the BP4 threshold of 0.3 or lower, BayesDel is 0.0836572, and SpliceAI predicts no meaningful splice effect with a maximum delta score of 0.01.
cspec ↗ spliceai ↗