Classification rationale
1
The POLE c.745C>A (p.Arg249=) variant has not been observed in COSMIC and has been reported in ClinVar as likely benign by two clinical laboratory submissions.
clinvar ↗2
3
This variant is not listed among the recurrent or hotspot POLE exonuclease-domain substitutions in the León-Castillo POLE review materials.
4
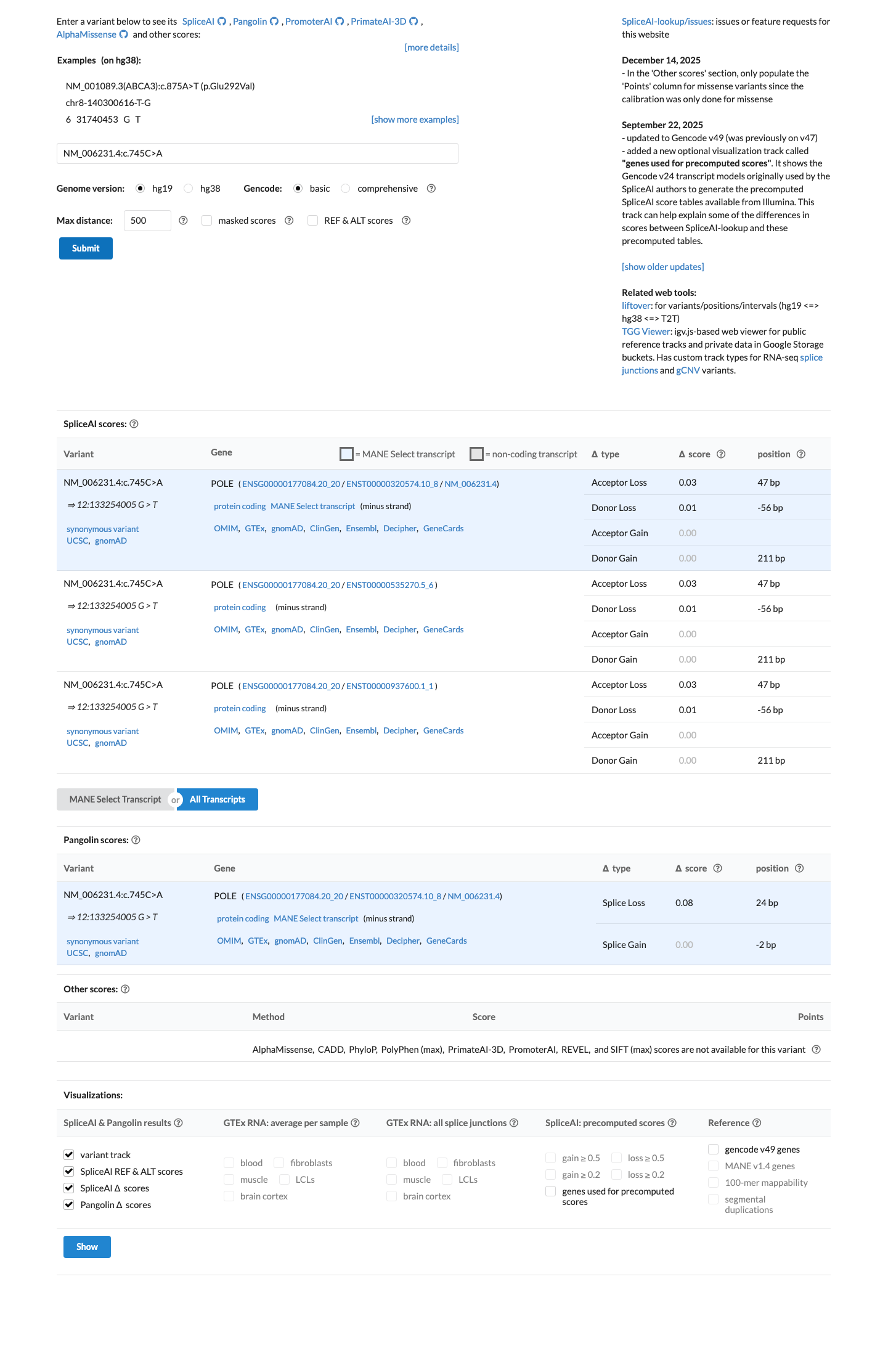
This is a synonymous change outside the canonical splice dinucleotides, and SpliceAI predicts no significant splice impact with a maximum delta score of 0.03.
spliceai ↗