Classification rationale
1
2
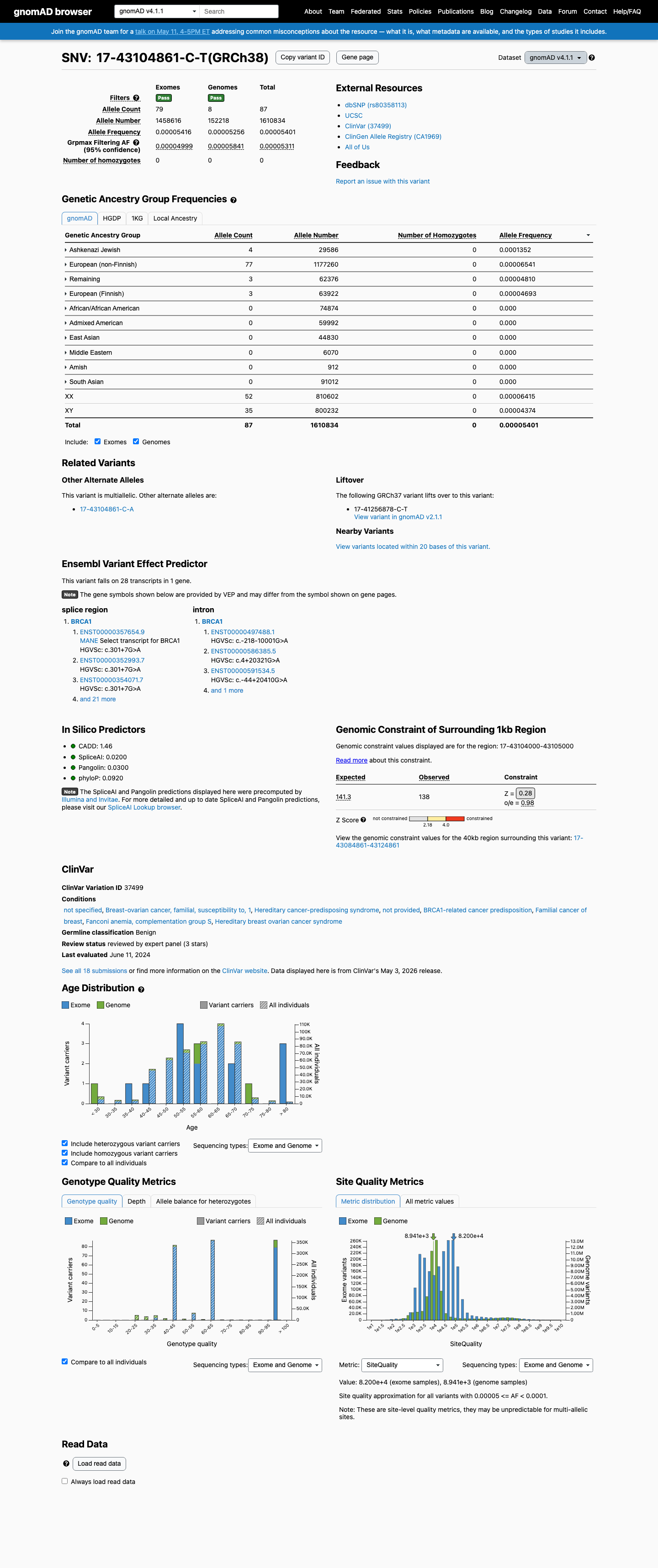
This variant is absent from gnomAD v2.1 but is present in gnomAD v4.1 at AF 5.40093e-05 (87/1610834 alleles; 0 homozygotes), so it is not absent from population controls.
gnomad_v2 ↗ gnomad_v4 ↗3
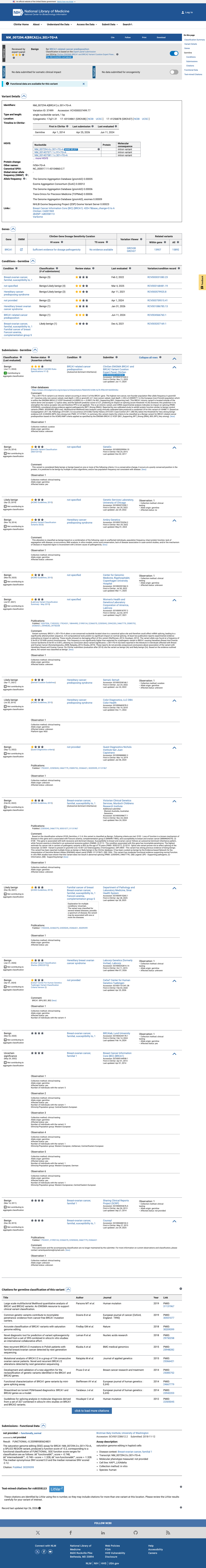
Functional evidence supports a benign effect: ENIGMA BRCA1 Table 9 assigns BS3_Strong for this variant, and RNA studies reported no aberrant splicing.
PMID:22505045 ↗ PMID:24667779 ↗4
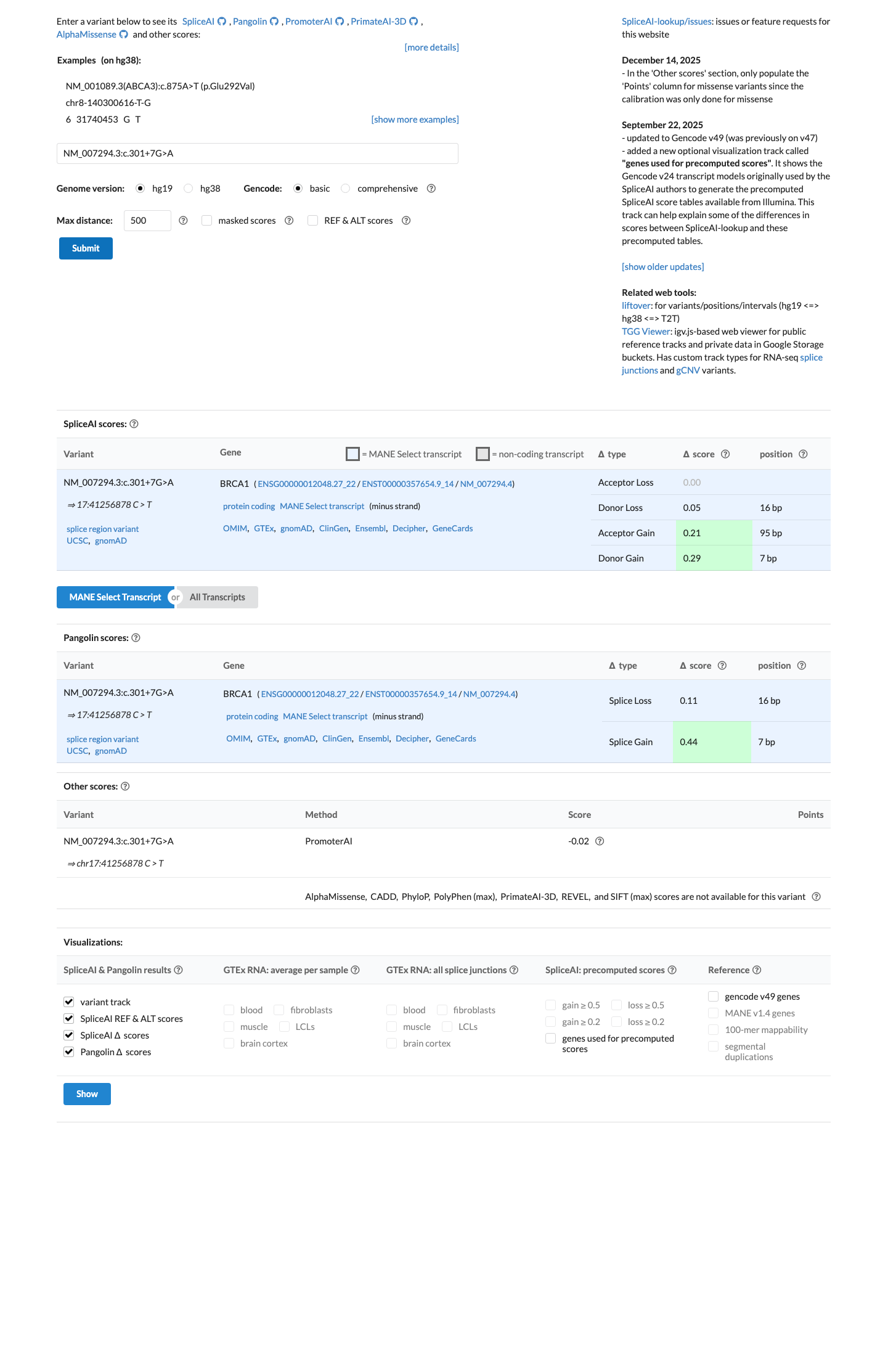
Computational splicing evidence predicts possible splice impact, with a SpliceAI maximum delta score of 0.29, which meets the ENIGMA BRCA1 PP3 threshold and does not meet the BP4 threshold.
spliceai ↗ cspec ↗