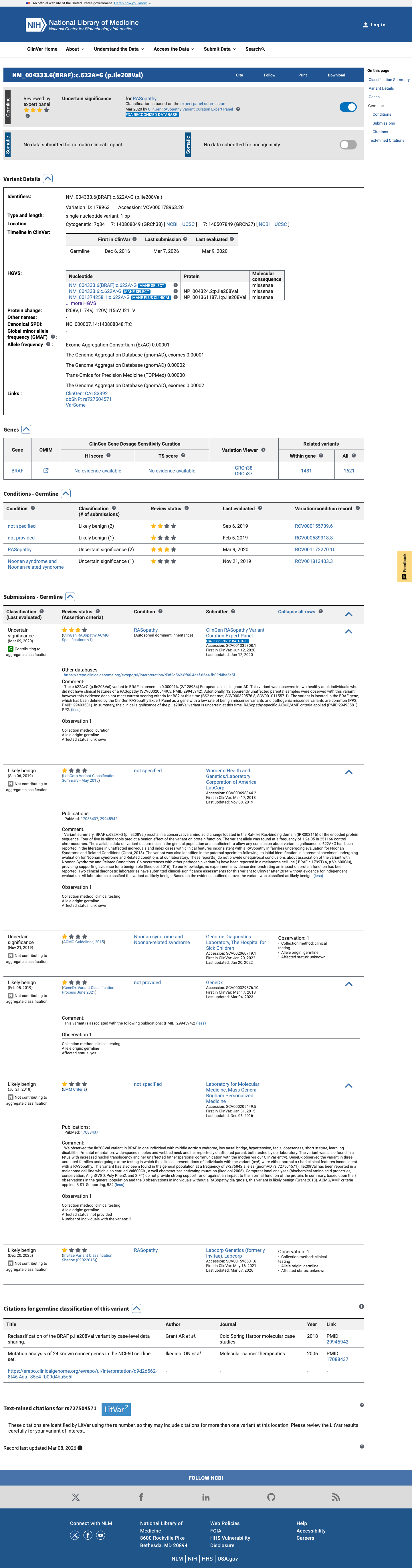
The BRAF c.622A>G (p.Ile208Val) variant has been reported in ClinVar with mostly likely benign laboratory submissions and an expert-panel uncertain significance classification, and published case-level data describe unaffected carriers and multiple individuals without findings consistent with a RASopathy.
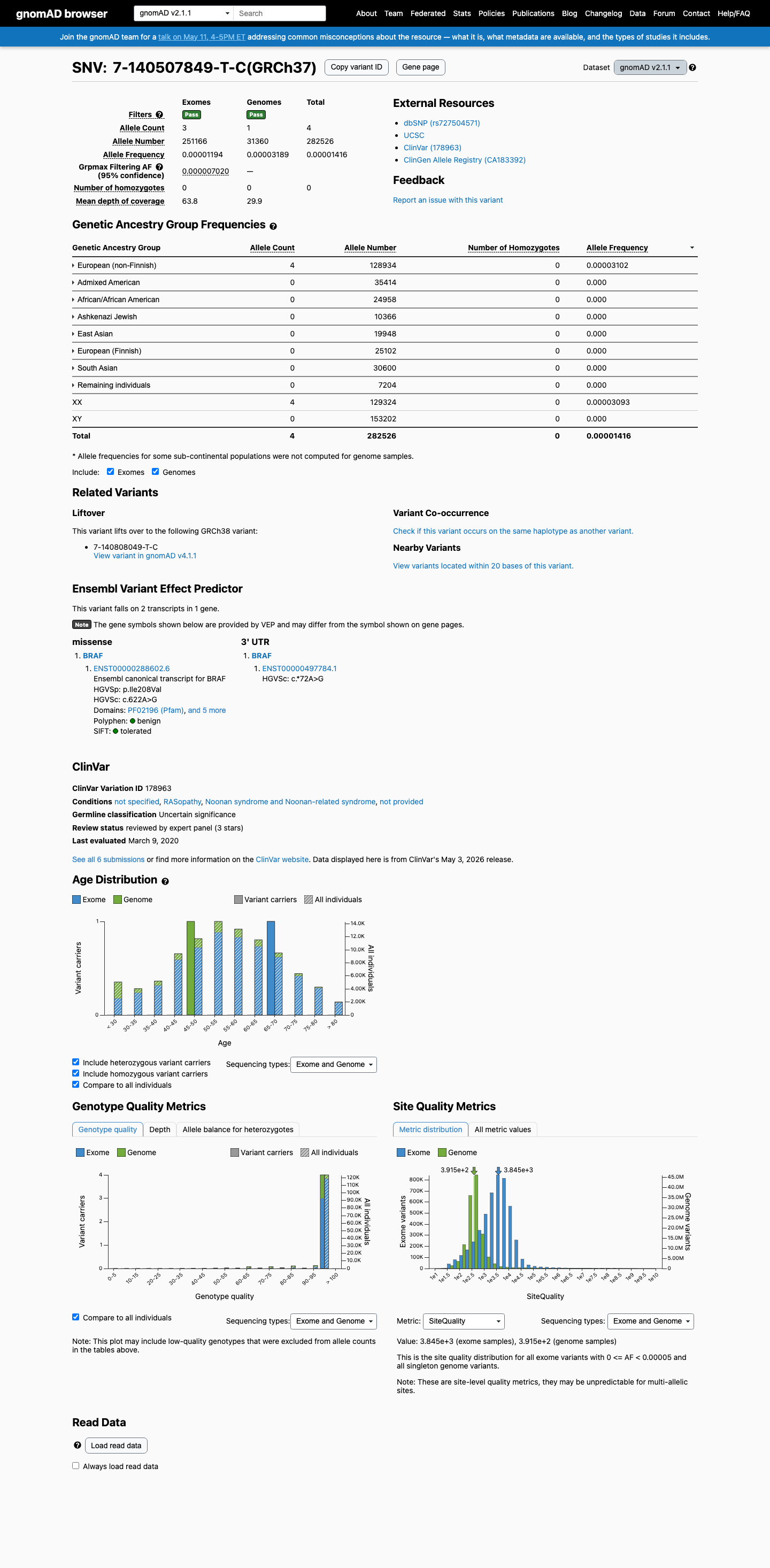
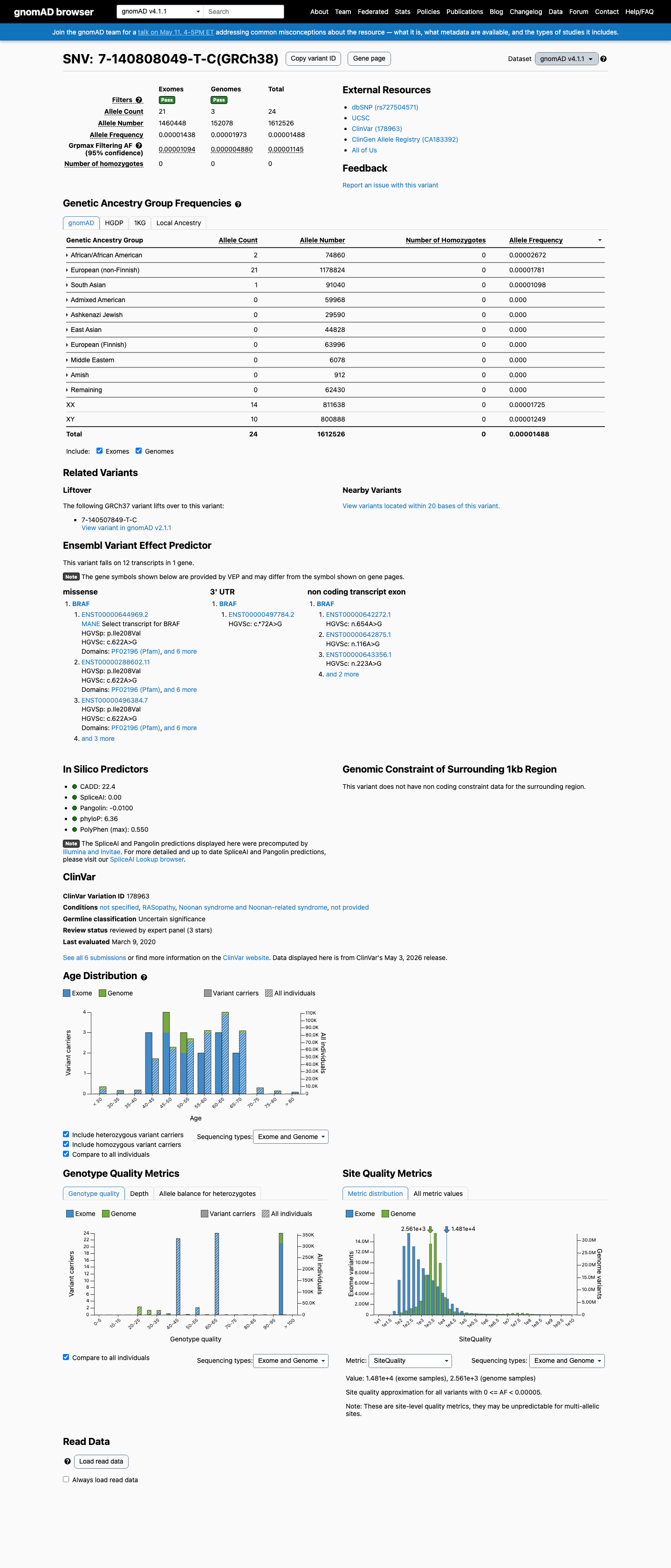
clinvar ↗ PMID:29945942 ↗This variant is present in population databases at low frequency, including 4/282526 alleles in gnomAD v2.1 (0.00142%) and 24/1612526 alleles in gnomAD v4.1 (0.00149%), which is below the BRAF RASopathy BS1 threshold of 0.025% and below the BA1 threshold of 0.05%, but means PM2 is not met because the variant is not absent from controls.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗No approved variant-specific functional study was identified for p.Ile208Val in the reviewed RASopathy functional-study materials.
cspec ↗Computational evidence does not meet the VCEP missense thresholds: REVEL is 0.318, which is below the PP3 cutoff of 0.7 but above the BP4 cutoff of 0.3, while SpliceAI predicts no meaningful splice effect with a maximum delta score of 0.01; the BayesDel score is -0.174758.
spliceai ↗ cspec ↗