Classification rationale
1
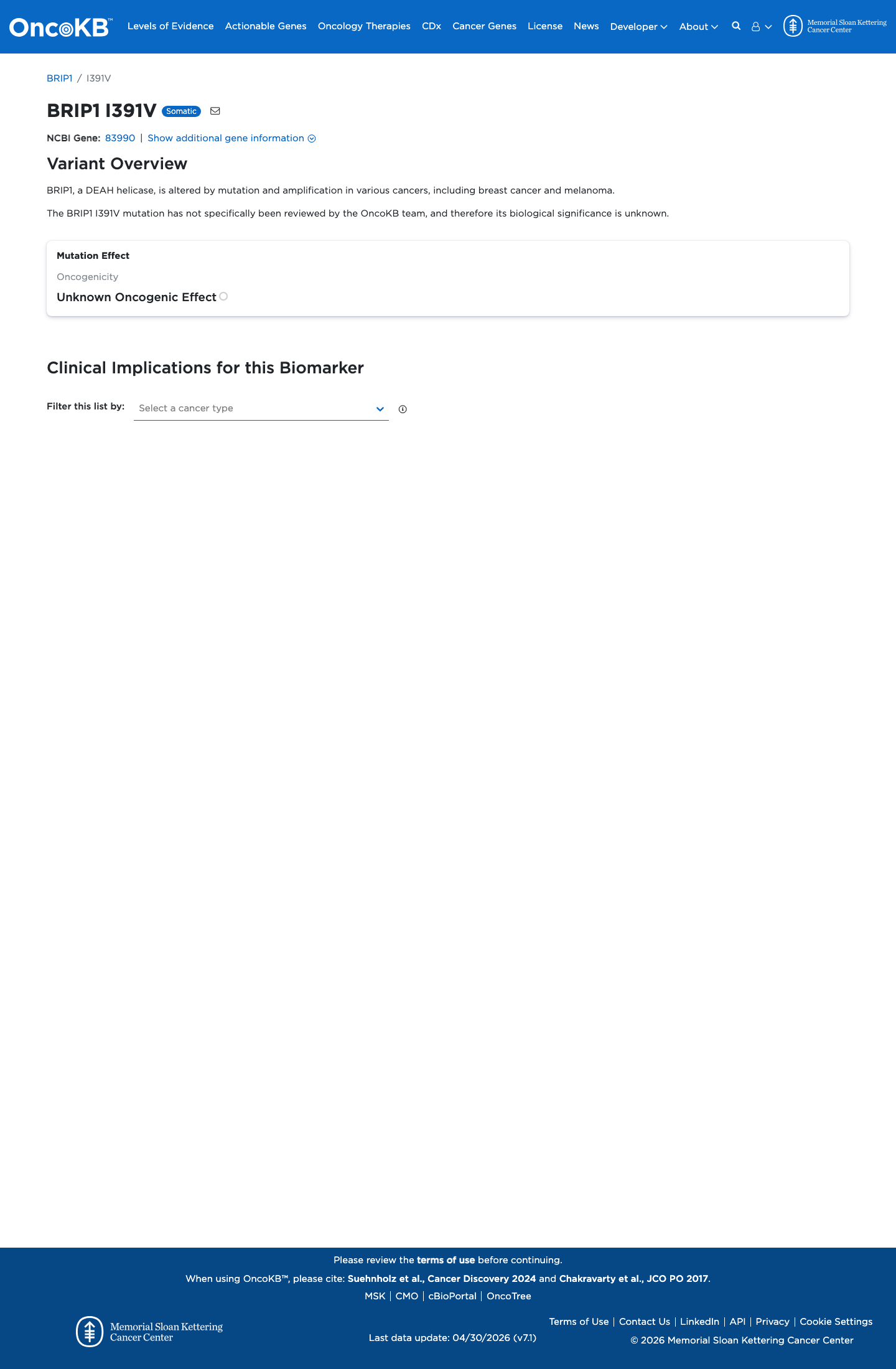
The BRIP1 c.1171A>G (p.Ile391Val) variant has been reported in ClinVar and is currently classified there as a variant of uncertain significance with single-submitter review status.
clinvar ↗2
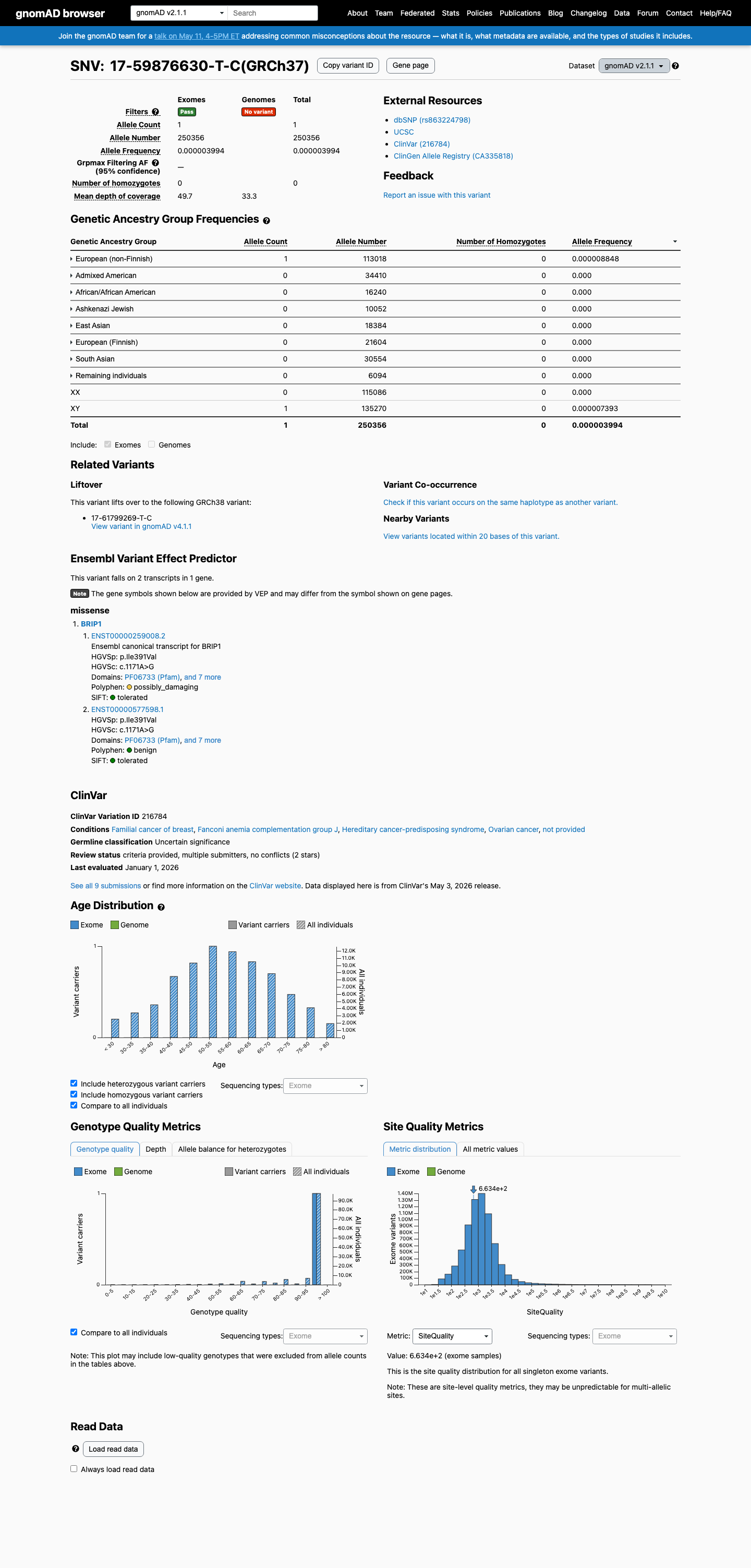
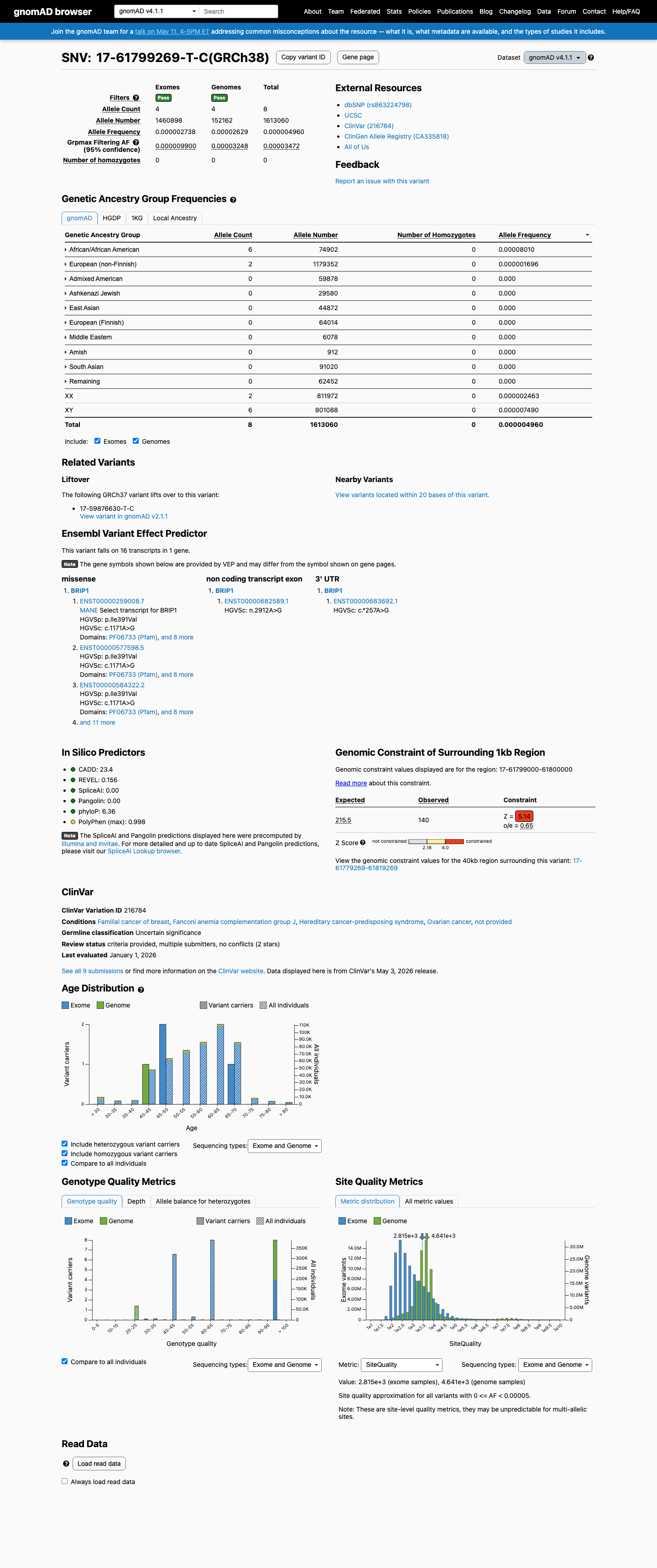
This variant is very rare in population databases, with gnomAD v2.1 AF 3.99431e-06 (1/250356) and gnomAD v4.1 AF 4.95952e-06 (8/1613060), which are below the 0.1% PM2 threshold.
gnomad_v2 ↗ gnomad_v4 ↗3
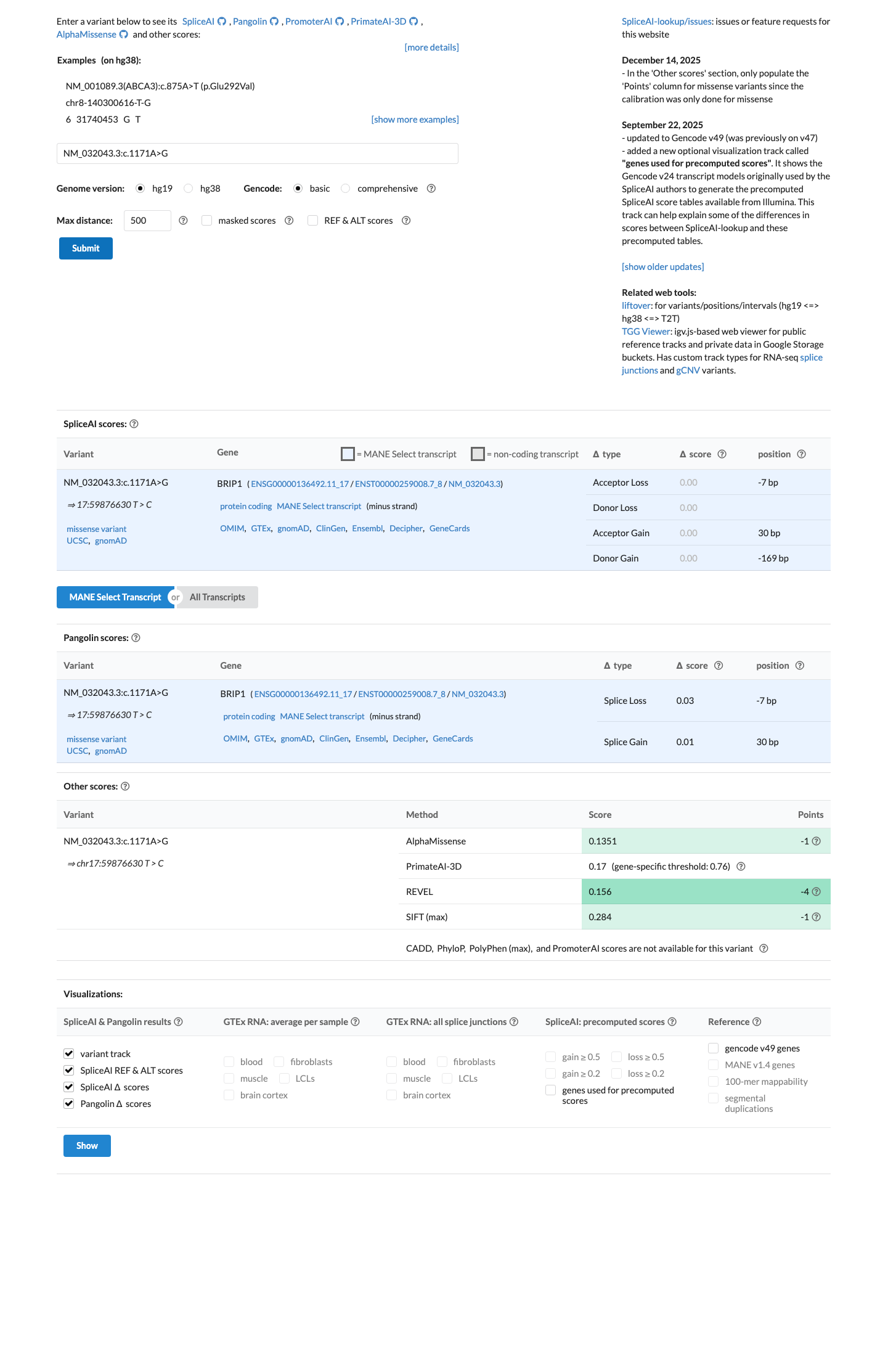
Computational data do not support a damaging effect: SpliceAI predicts no significant splice impact with a maximum delta score of 0.00, REVEL is 0.156, and BayesDel is -0.370746.
spliceai ↗