Classification rationale
1
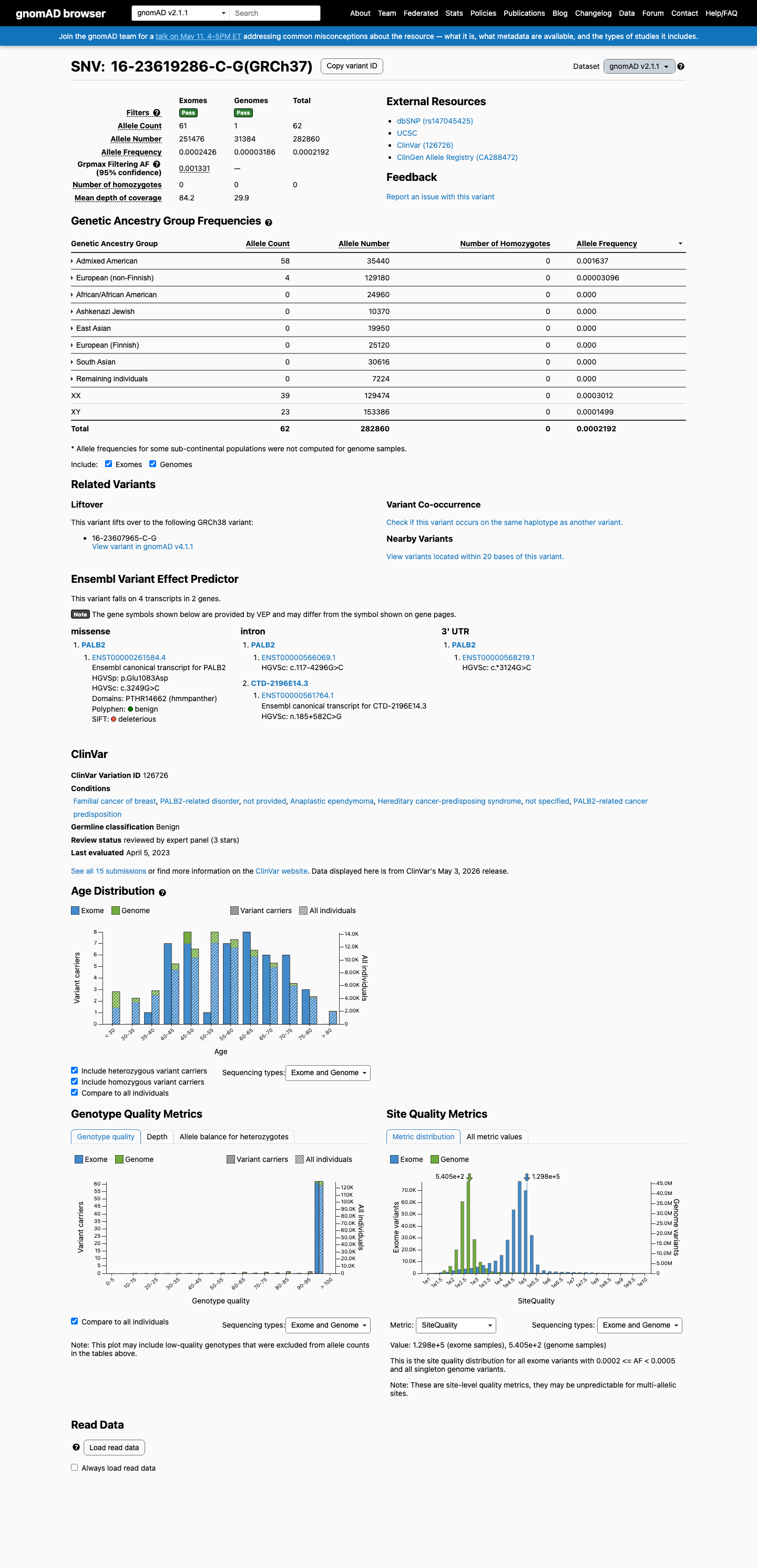
The PALB2 c.3249G>C (p.Glu1083Asp, p.E1083D) variant has been reported in ClinVar and is classified there as benign by the ClinGen Hereditary Breast, Ovarian and Pancreatic Cancer Variant Curation Expert Panel.
clinvar ↗2
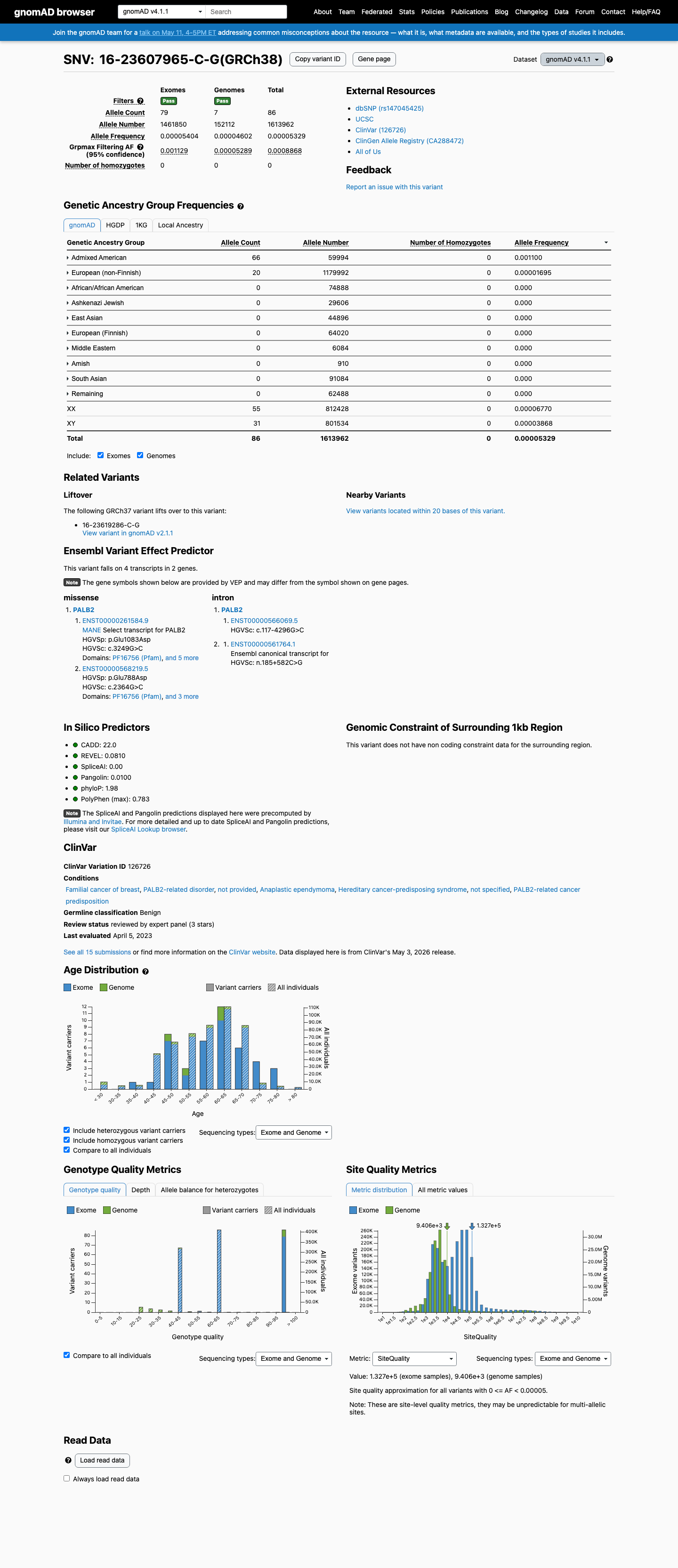
In gnomAD v4, the variant has a total allele frequency of 0.00533% and a grpmax filtering allele frequency of 0.088684%, which is above the PALB2 BS1 threshold of 0.01% but below the BA1 threshold of 0.1%.
gnomad_v4 ↗ cspec ↗3
Computational data predict no significant splice effect, with a SpliceAI max delta score of 0.00; REVEL is 0.081 and BayesDel is -0.49371, although PALB2 VCEP guidance does not apply missense PP3 or BP4 for this gene.
spliceai ↗ cspec ↗