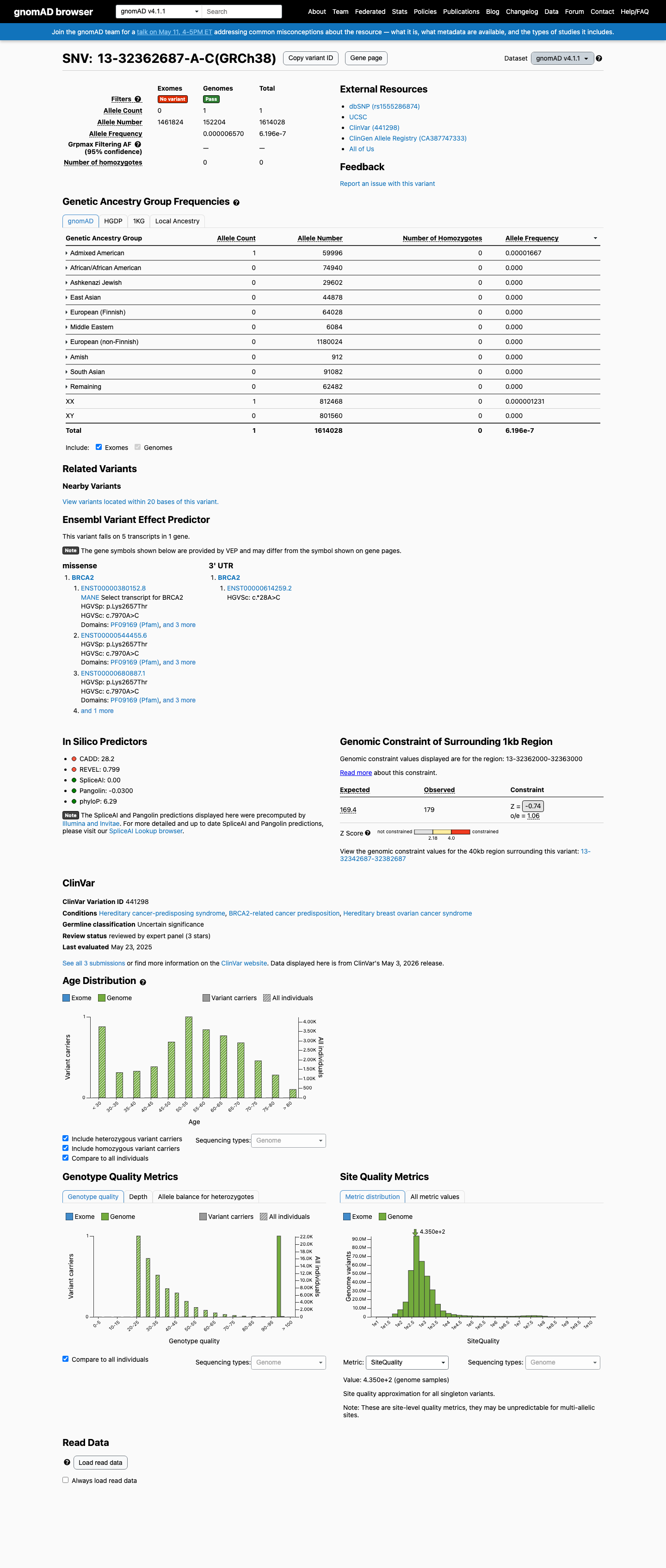
The BRCA2 c.7970A>C (p.Lys2657Thr, K2657T) variant has been reported in ClinVar, where the ClinGen ENIGMA expert panel classified it as uncertain significance and other submitters reported likely pathogenic and uncertain significance assertions.
clinvar ↗This variant is absent from gnomAD v2.1 and is present once in gnomAD v4.1 (1/1614028 alleles overall; highest observed frequency 1/59996 in Admixed American individuals), supporting rarity but not complete absence from population databases.
gnomad_v2 ↗ gnomad_v4 ↗In a calibrated BRCA2 functional study summarized by the ENIGMA BRCA1/2 specification, p.Lys2657Thr showed protein function similar to pathogenic control variants, supporting PS3 at strong strength.
cspec ↗ PMID:33609447 ↗Computational evidence predicts no significant splice impact (SpliceAI max delta score 0.01), while the missense prediction profile is mixed under ENIGMA thresholds (REVEL 0.799; BayesDel 0.270216), so PP3 and BP4 are not independently met.
cspec ↗ spliceai ↗A BRCA2 clinical-history likelihood ratio of 1.188967663938105 from 1 proband falls in the ENIGMA neutral zone (>0.48 and <2.08), so neither PP4 nor BP5 is supported by the reviewed multifactorial clinical-history data.
cspec ↗ PMID:31853058 ↗