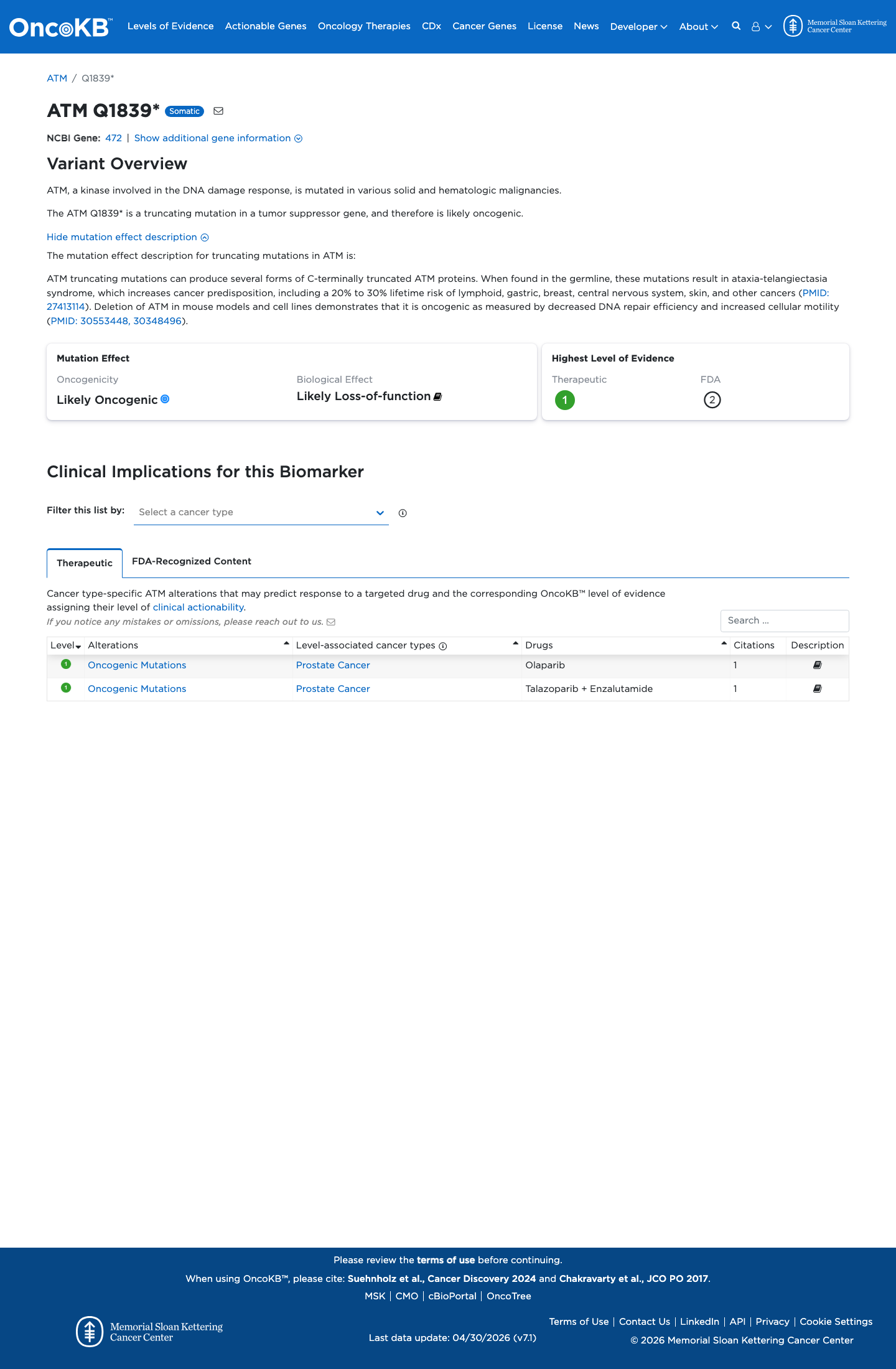
The ATM c.5515C>T (p.Gln1839Ter; p.Q1839*) variant has been reported in ClinVar as pathogenic, including expert panel review.
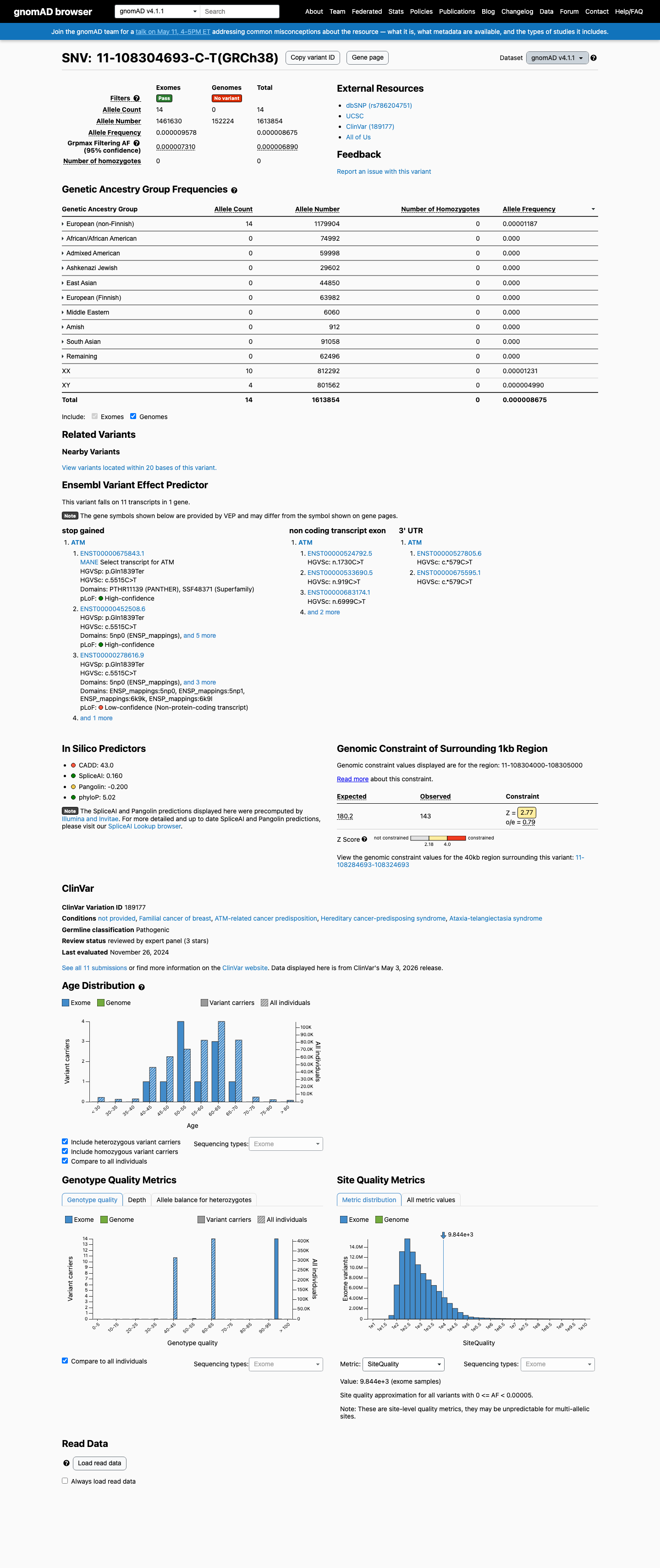
clinvar ↗This variant is absent from gnomAD v2.1 and is present in gnomAD v4.1 at a very low overall allele frequency of 0.00087% (14/1613854 alleles) with no homozygotes, which is below the ATM PM2_Supporting threshold of 0.001%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗This nonsense variant introduces a premature stop codon at residue 1839, and the ATM VCEP PVS1 framework supports loss of function as a disease mechanism for ATM.
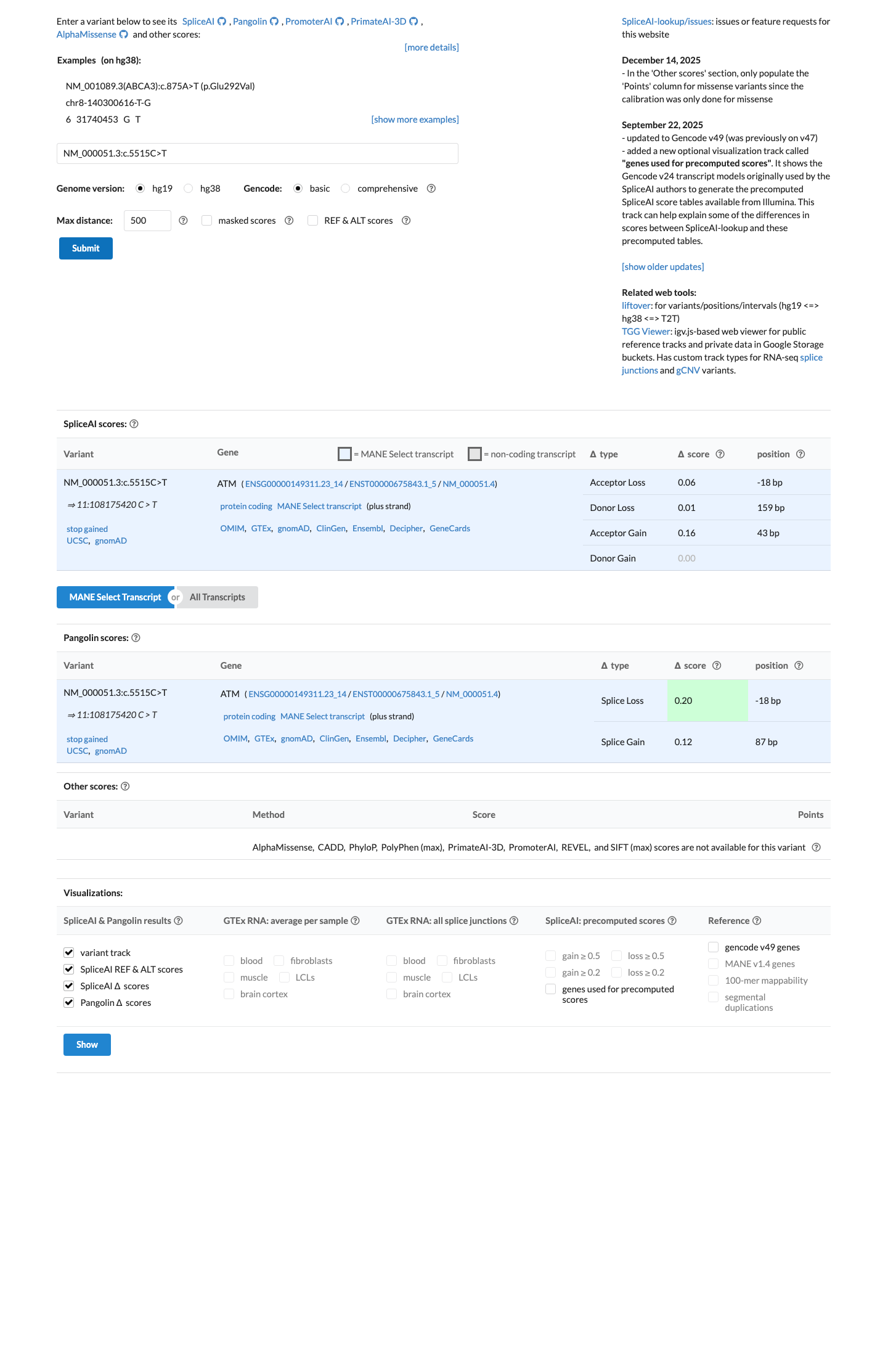
cspec ↗SpliceAI predicts no strong splice effect for this variant (max delta score 0.16), which is below the ATM PP3 splicing threshold of 0.2 and above the BP4 threshold of 0.1, so computational evidence does not independently change the interpretation.
spliceai ↗ cspec ↗