The BRCA2 c.794-2A>G (p.?) variant has been reported in ClinVar, where the ClinGen ENIGMA BRCA1 and BRCA2 Variant Curation Expert Panel classifies it as uncertain significance, while other submitters have reported likely pathogenic or pathogenic interpretations.
clinvar ↗This variant is absent from gnomAD v2.1 and present once in gnomAD v4.1 at an overall allele frequency of 6.260650932398744e-07 (1/1597278 alleles; 0 homozygotes), indicating that it is very rare in population databases.
gnomad_v2 ↗ gnomad_v4 ↗In the ENIGMA BRCA2 exon-level splice/PVS1 table, this exact splice acceptor variant is associated with observed in-frame exon 10 skipping and assigned PVS1_N/A (RNA), so current VCEP evidence does not support treating it as a qualifying loss-of-function event under PVS1.
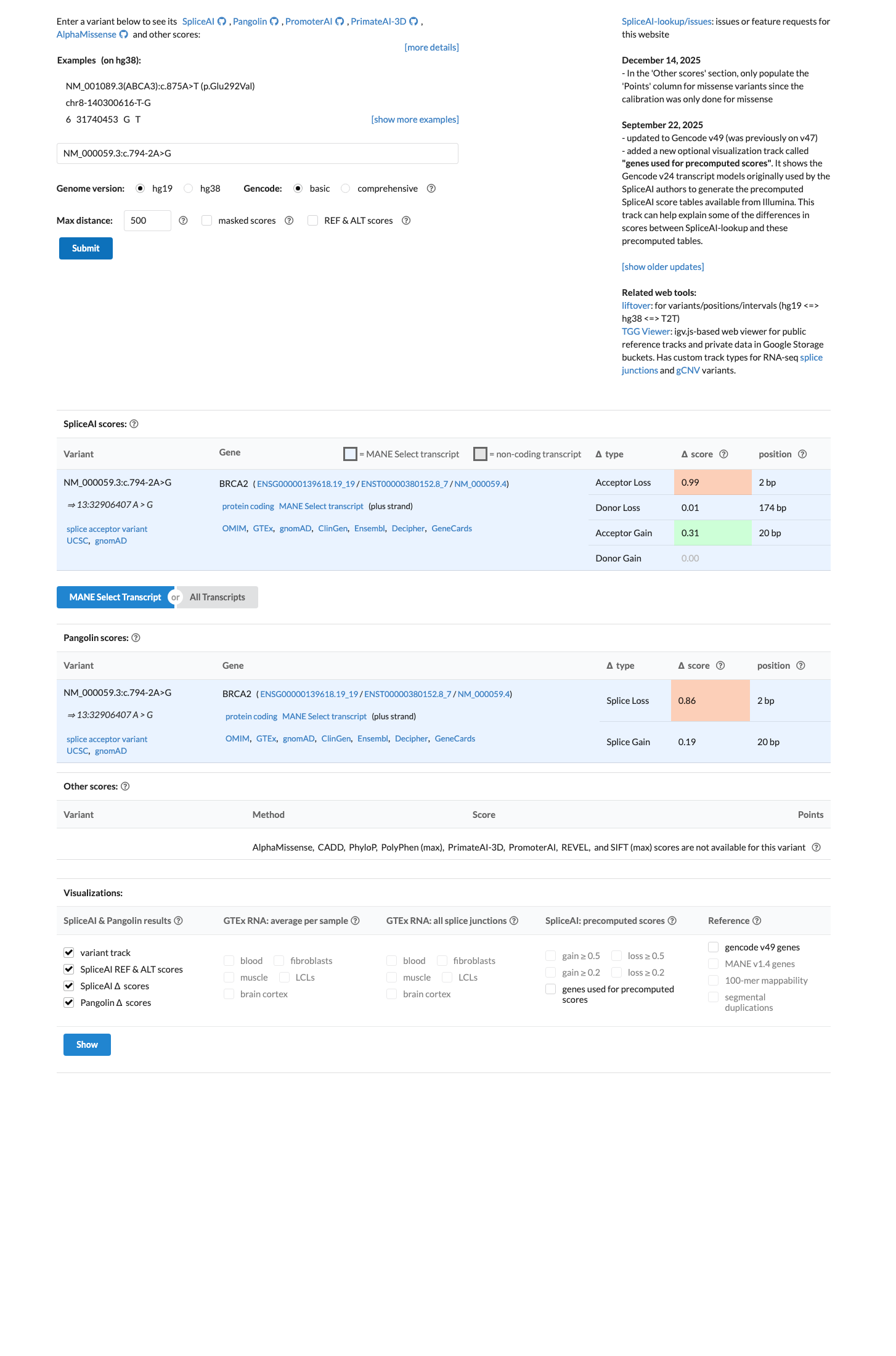
cspec ↗SpliceAI predicts strong splice impact for this variant with a maximum delta score of 0.99 (DS_AL 0.99, DS_AG 0.31), but the BRCA2 ENIGMA framework does not apply PP3 to canonical ±1,2 splice-site variants and this splice prediction should not be double-counted against the variant-specific RNA/PVS1 assessment.
spliceai ↗ cspec ↗