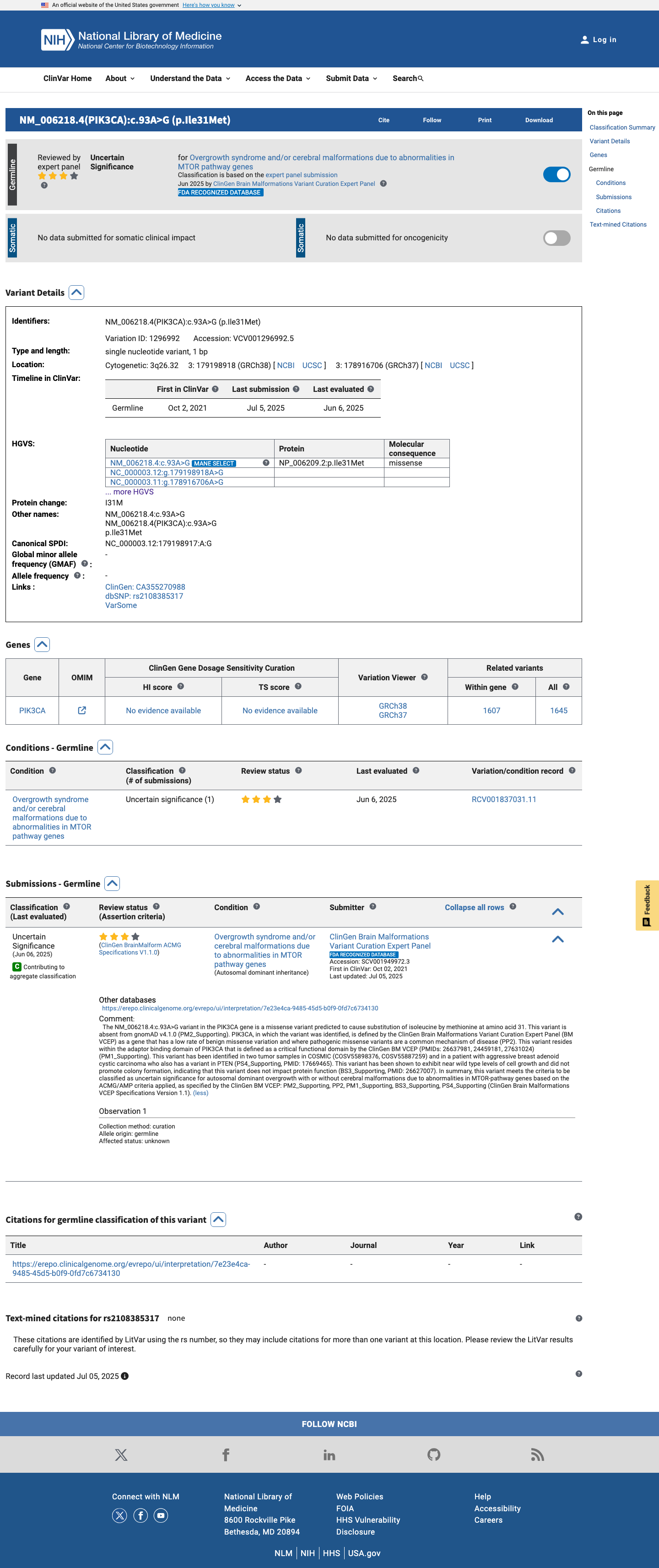
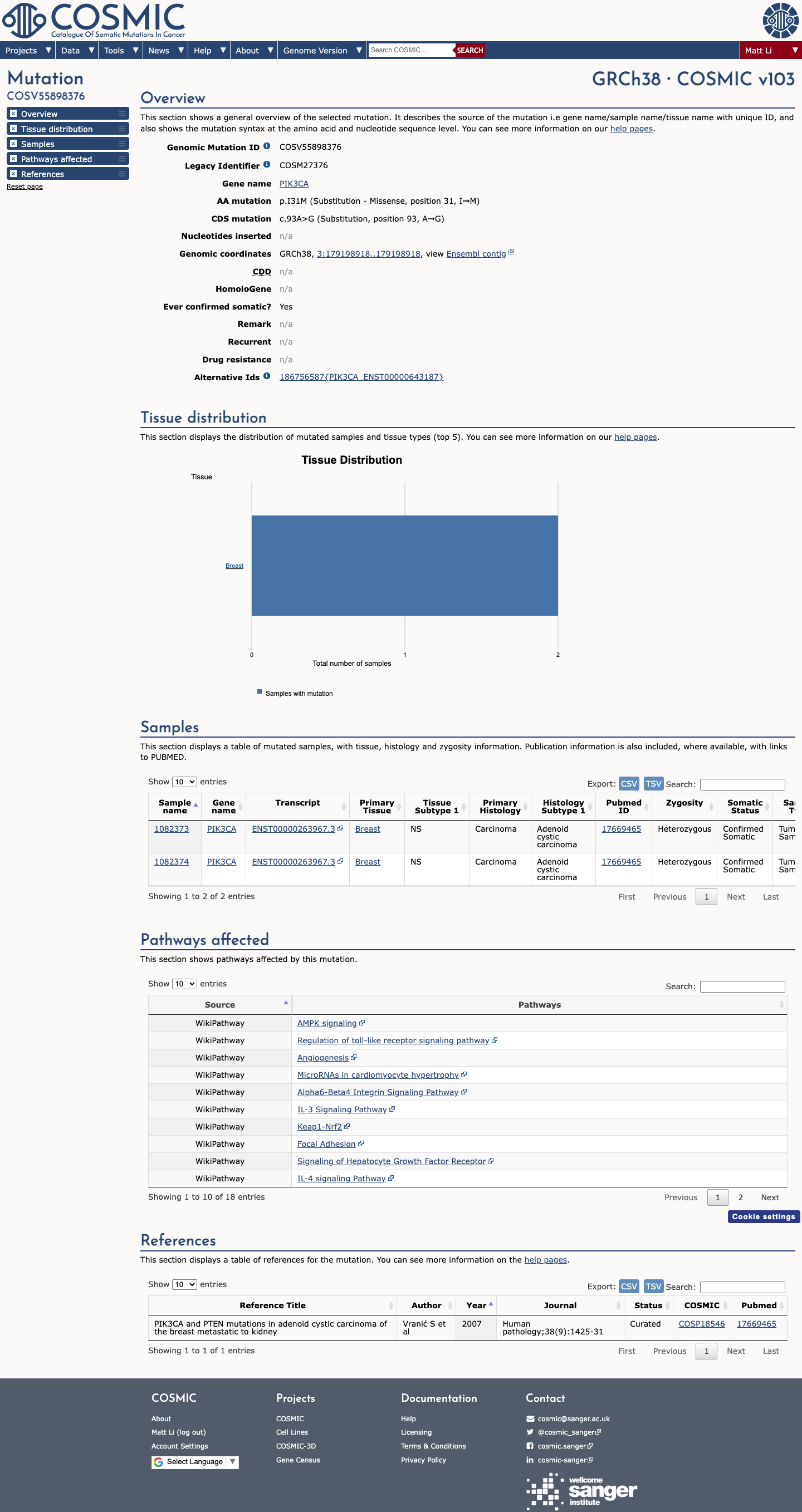
The PIK3CA c.93A>G (p.Ile31Met) variant has been observed in somatic cancers in COSMIC (COSV55898376, n=2) and has been reported in ClinVar as Uncertain Significance by the ClinGen Brain Malformations Variant Curation Expert Panel.
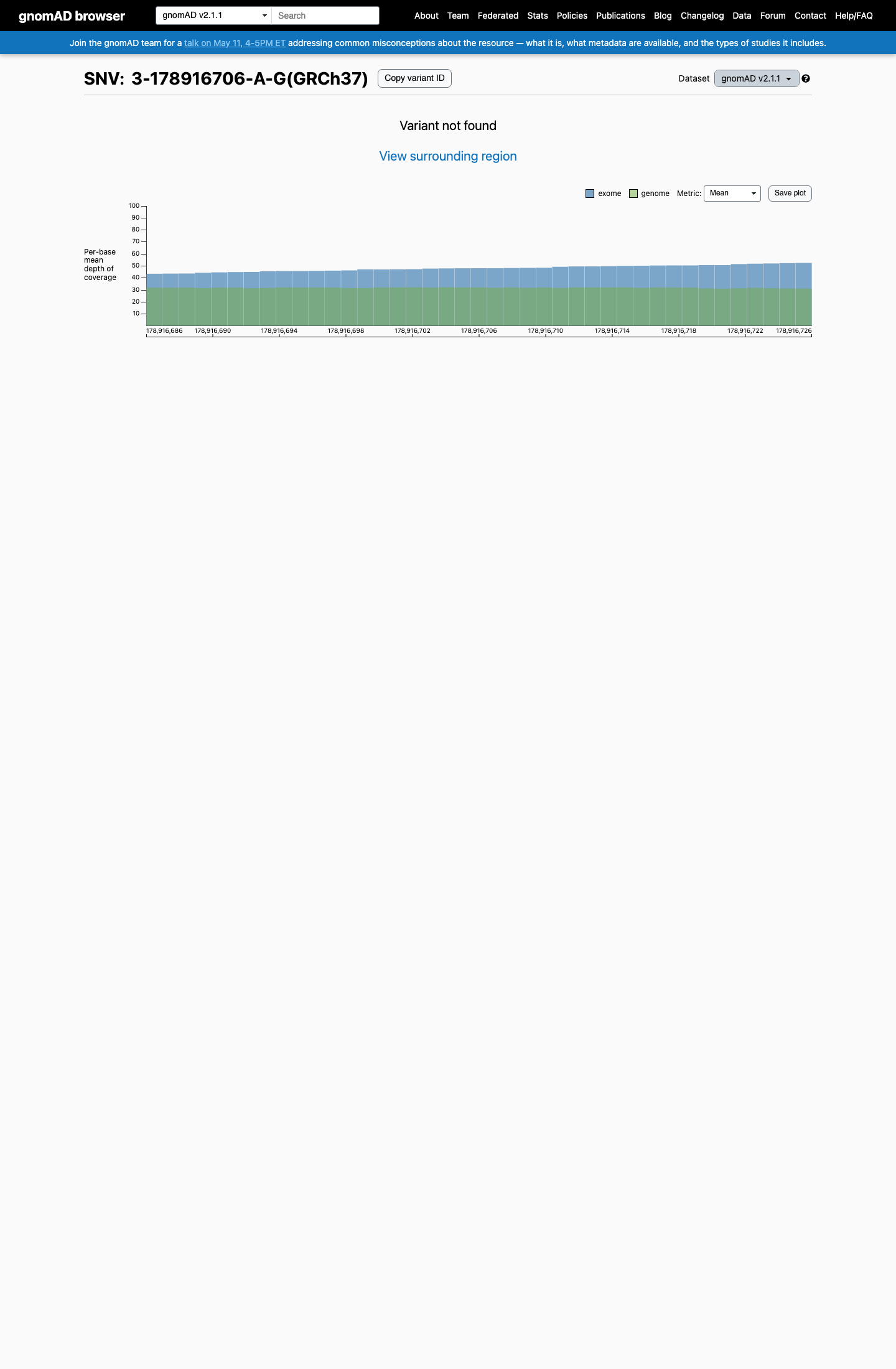
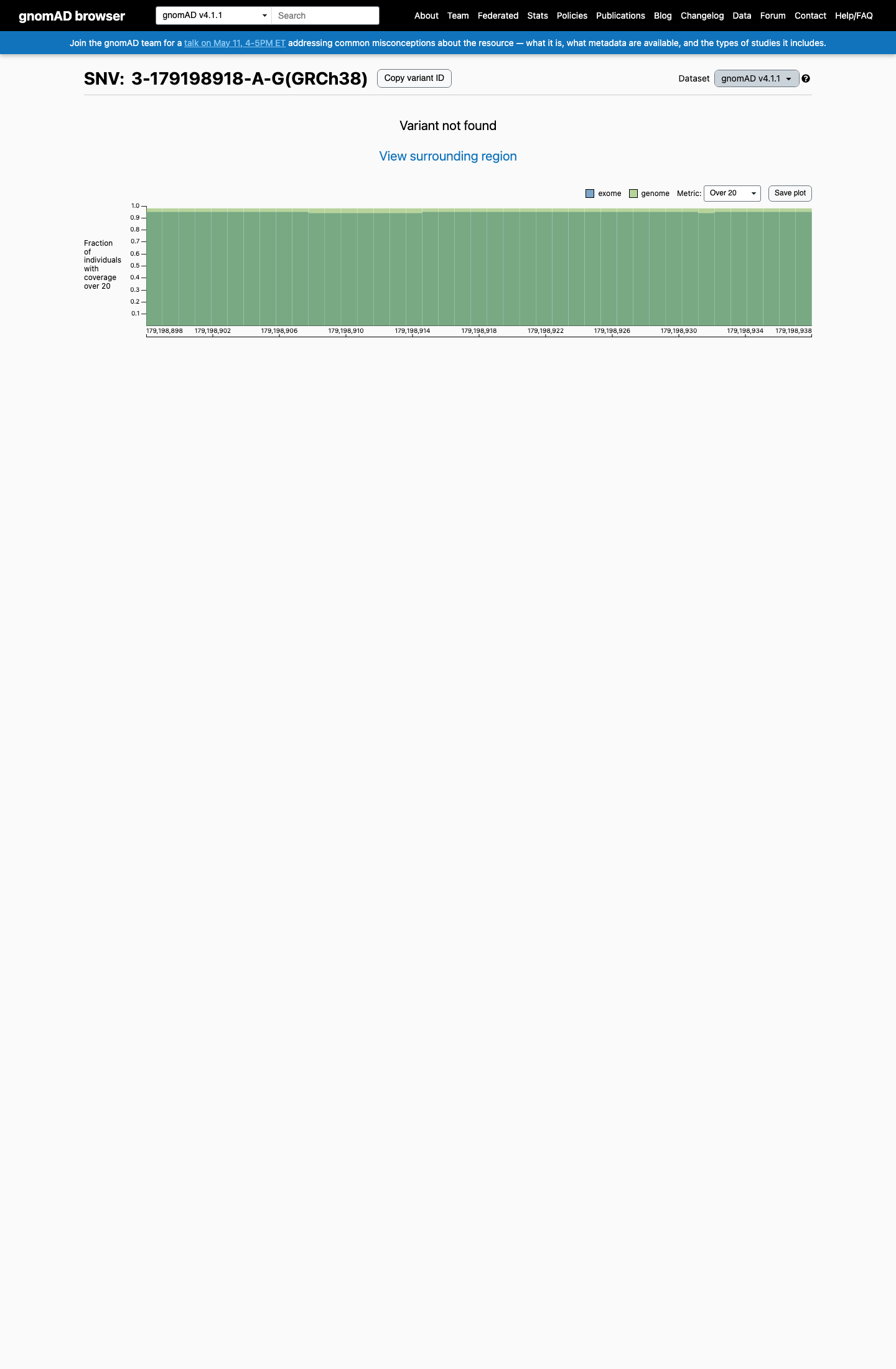
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population databases and meeting PM2 at supporting strength under the Brain Malformations VCEP framework.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗The missense change affects codon 31, which is outside the PIK3CA Table 4 kinase-domain intervals used for PM1 and was not identified as a statistically significant hotspot in the retrieved hotspot review.
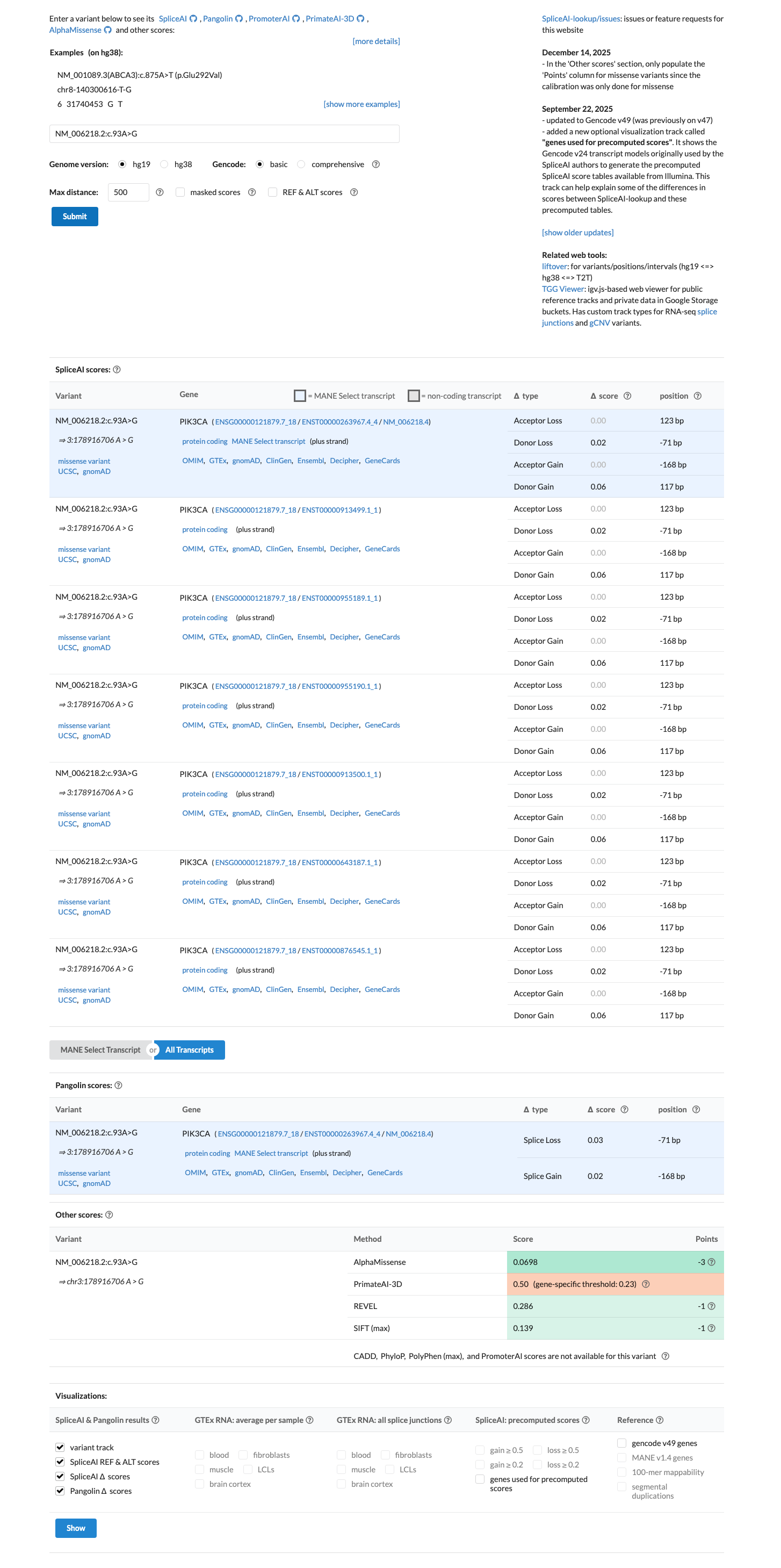
cspec ↗ hotspots ↗Computational review showed REVEL 0.286, BayesDel -0.0727233, and SpliceAI max delta 0.06, but the Brain Malformations VCEP does not apply PP3 to PIK3CA gain-of-function missense variants and restricts BP4 to synonymous, intronic, or UTR variants.
cspec ↗ spliceai ↗