Classification rationale
1
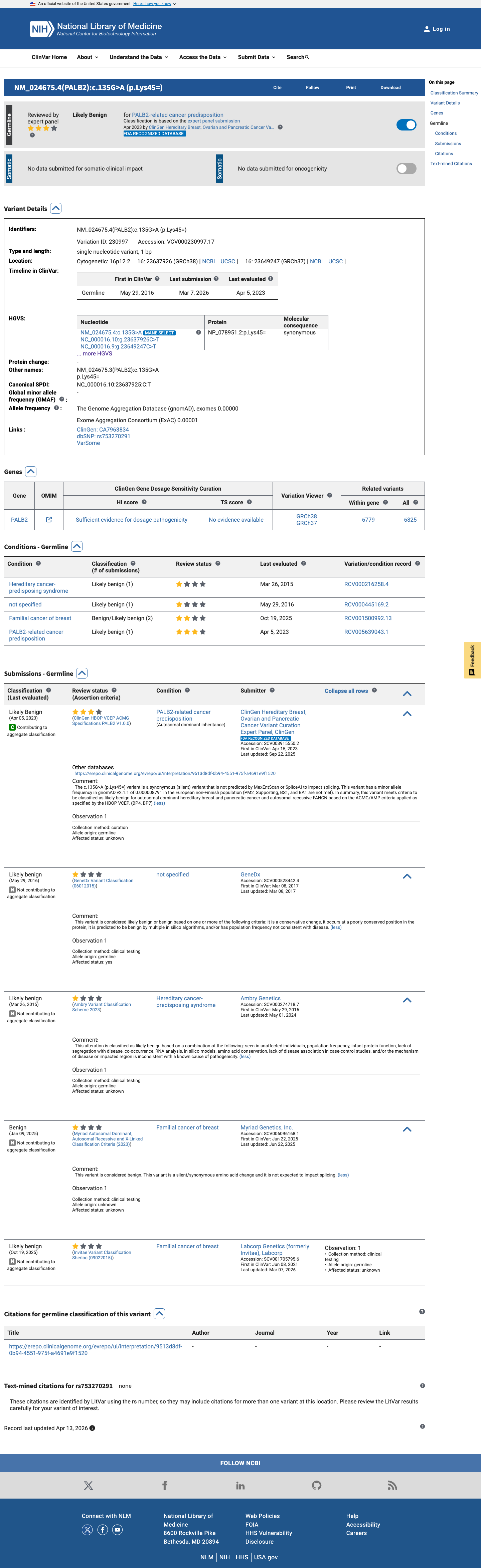
The PALB2 c.135G>A (p.Lys45=) variant has been reported in ClinVar with an expert panel classification of likely benign.
clinvar ↗2
This variant is present in gnomAD v4.1 at 2/1,614,022 alleles (AF 0.00012%) with a highest observed population frequency of 0.00017% and grpmax FAF 2.8e-07, which is below the PALB2 PM2_Supporting threshold of 0.000333% and below the BS1 and BA1 benign frequency thresholds.
gnomad_v4 ↗ cspec ↗3
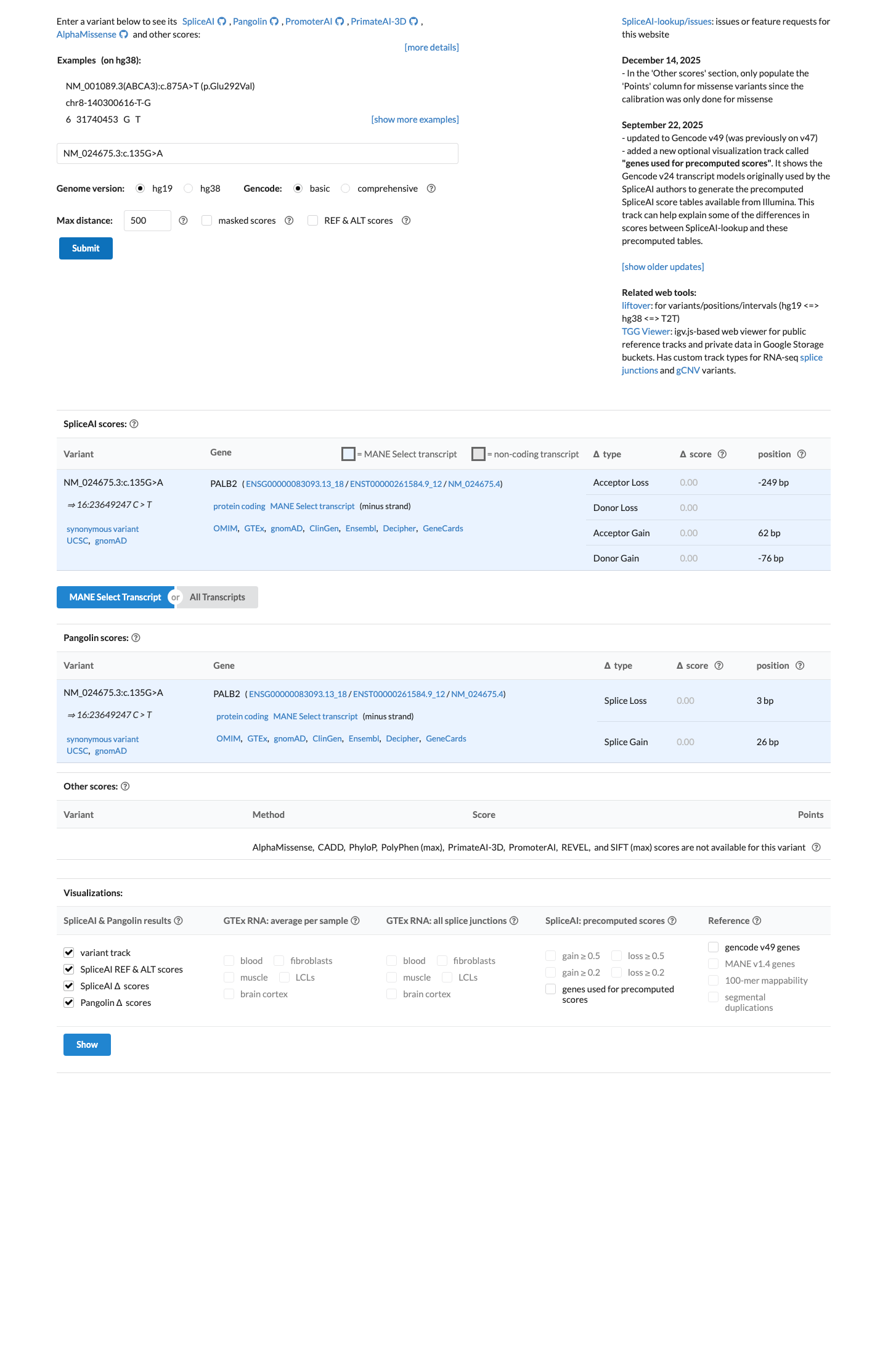
SpliceAI predicts no significant splice impact for this synonymous variant, with a maximum delta score of 0.00, which supports BP4 and argues against PP3 or a splice-based loss-of-function interpretation.
spliceai ↗ cspec ↗