Classification rationale
1
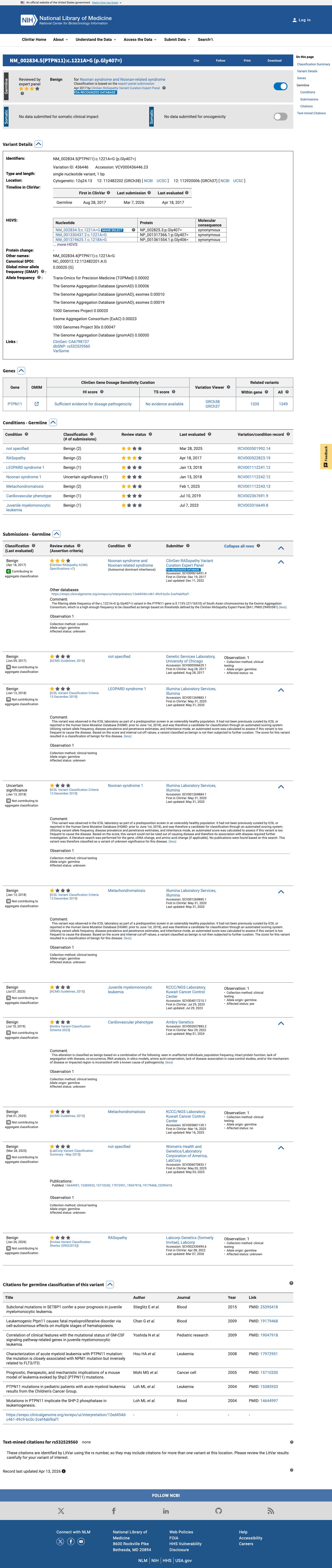
The PTPN11 c.1221A>G (p.Gly407=) variant has not been identified in a statistically significant somatic hotspot and has been reported in ClinVar as Benign, including a benign expert-panel assertion from the ClinGen RASopathy Variant Curation Expert Panel.
hotspots ↗ clinvar ↗2
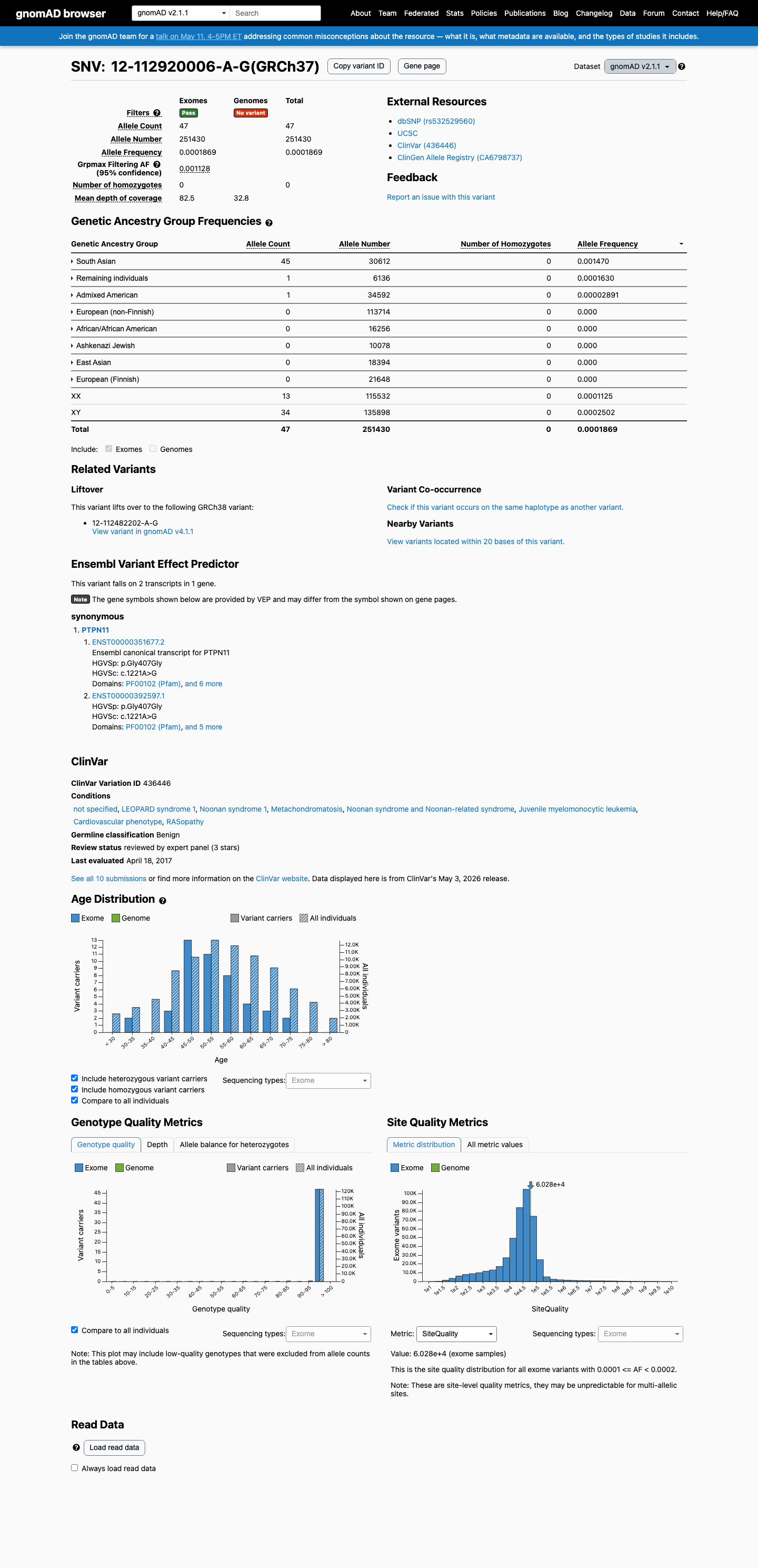
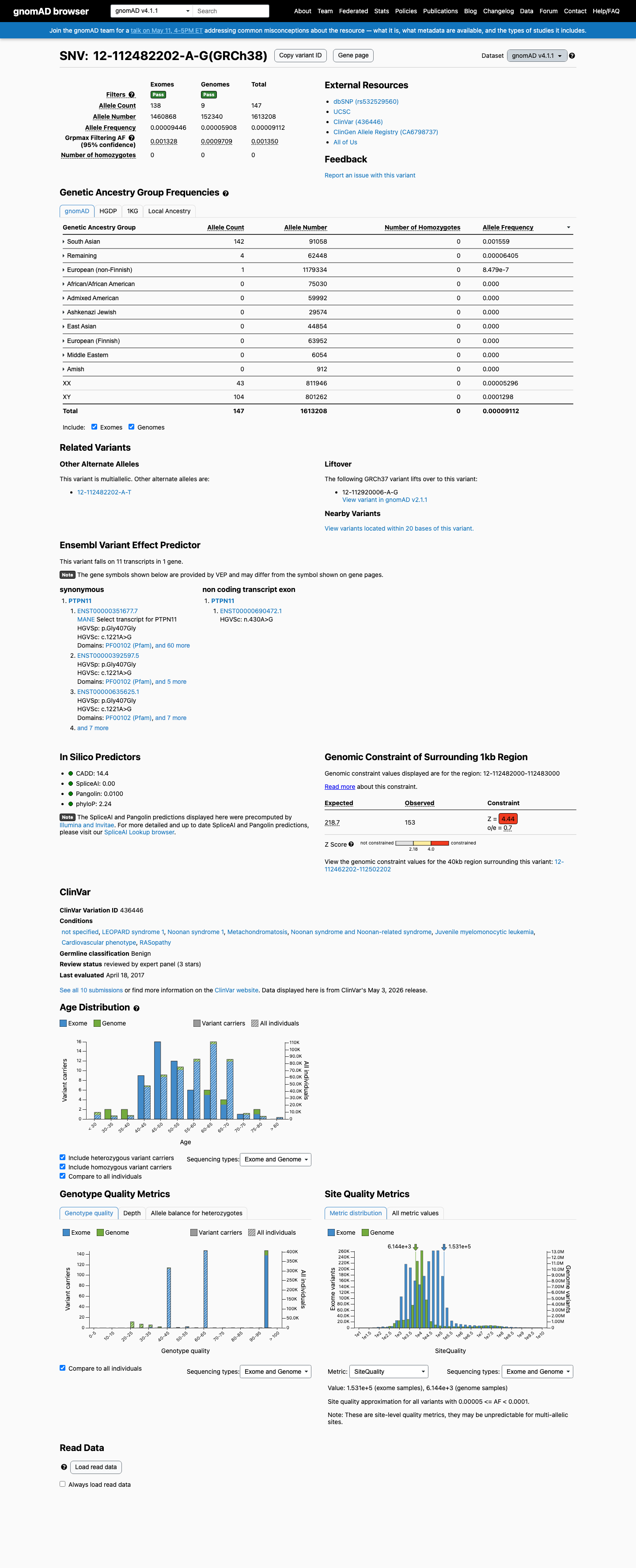
In gnomAD, this variant exceeds the PTPN11 RASopathy benign population thresholds, with grpmax filtering allele frequencies of 0.11284% in v2.1 and 0.13502% in v4.1, both above the BA1 threshold of 0.05% and the BS1 threshold of 0.025%.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
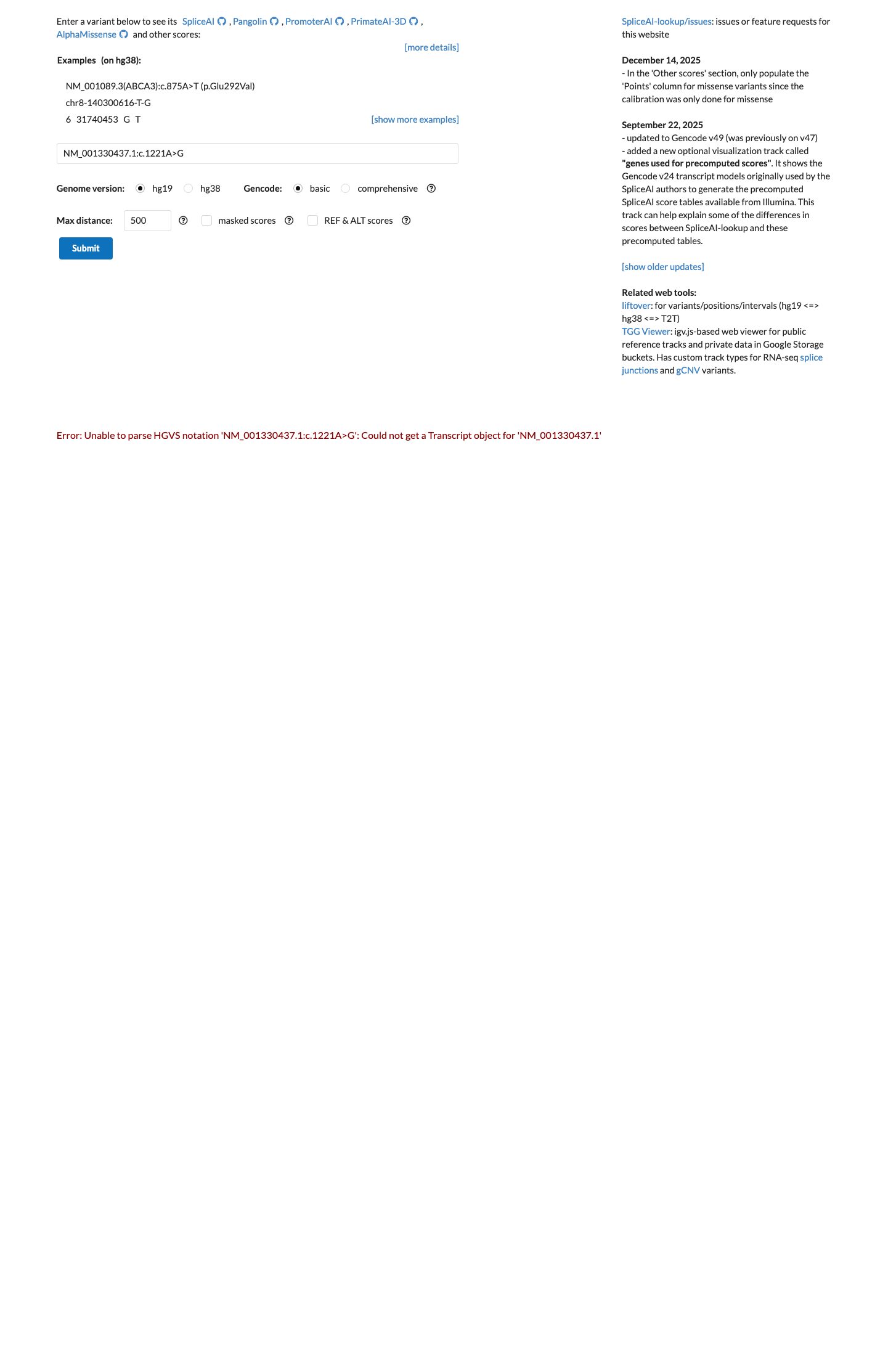
This synonymous variant does not change the encoded amino acid, and SpliceAI predicts no significant splice impact, with a maximum delta score of 0.02.
spliceai ↗