Classification rationale
1
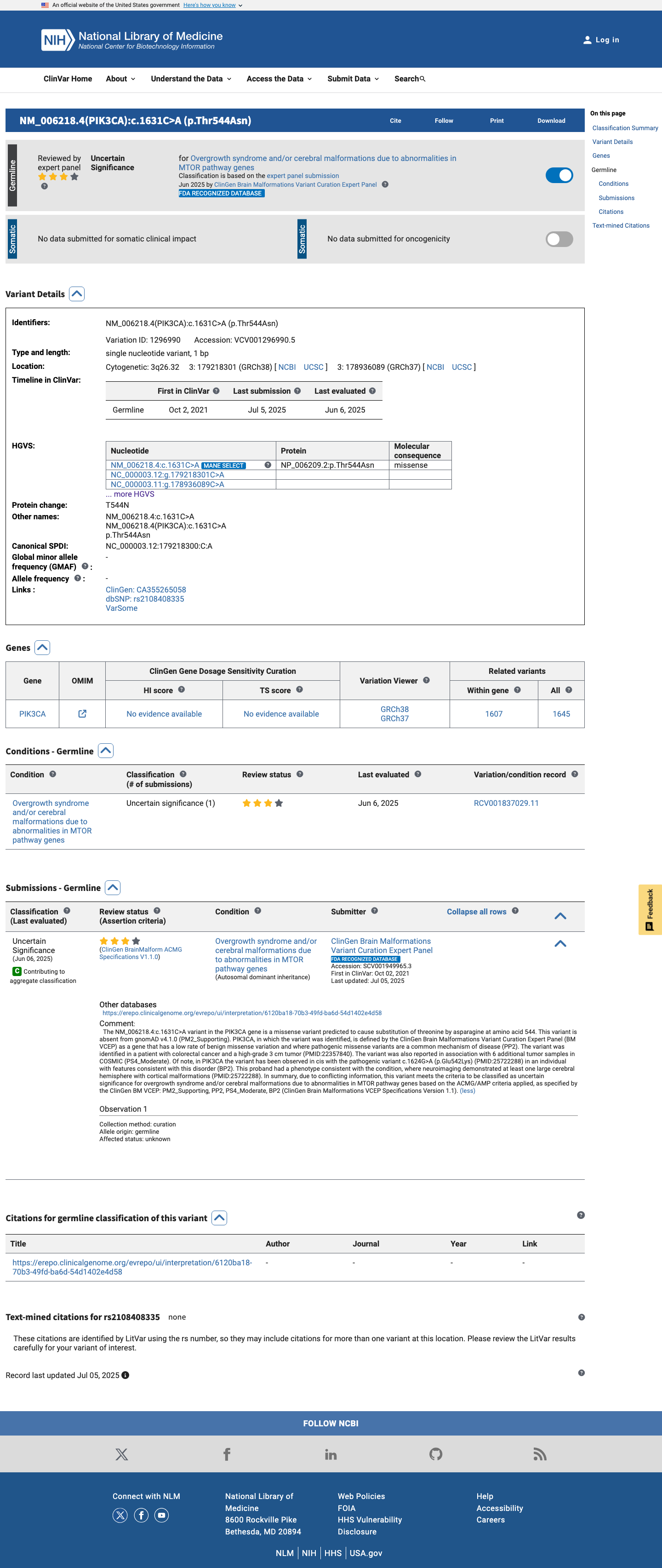
The PIK3CA c.1631C>A (p.Thr544Asn, T544N) variant has been reported in ClinVar and is currently classified as uncertain significance by the ClinGen Brain Malformations Variant Curation Expert Panel.
clinvar ↗2
This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting PM2 at Supporting strength under the Brain Malformations VCEP specification.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
This variant lies outside the VCEP-approved PIK3CA PM1 domains at amino acids 322-483 and 797-1068, so PM1 is not met.
cspec ↗4
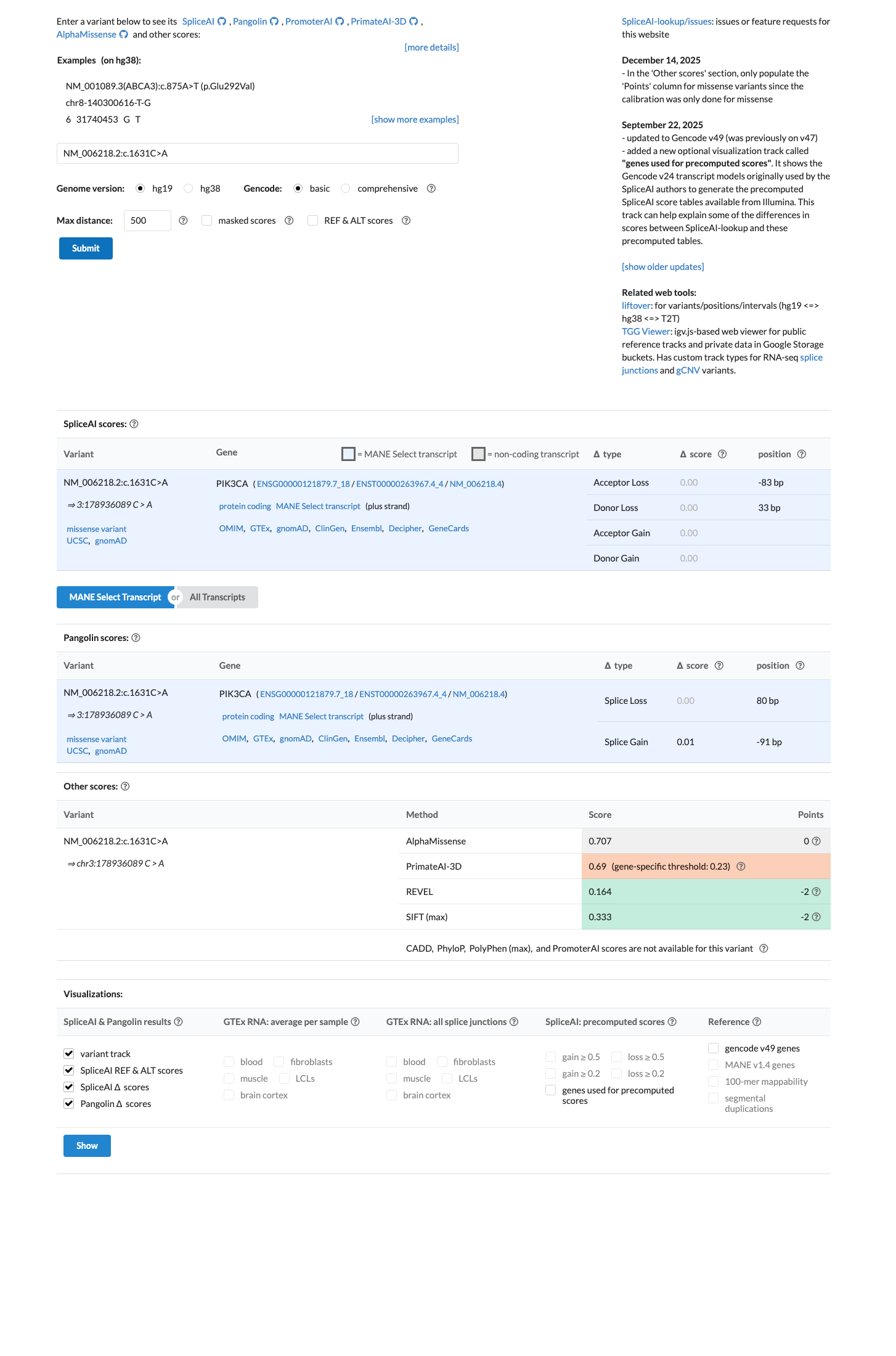
SpliceAI predicts no significant splice impact (max delta score 0.00), REVEL is 0.164, and BayesDel is -0.174013; however, the Brain Malformations VCEP does not apply PP3 or BP4 missense computational criteria to gain-of-function PIK3CA variants.
spliceai ↗