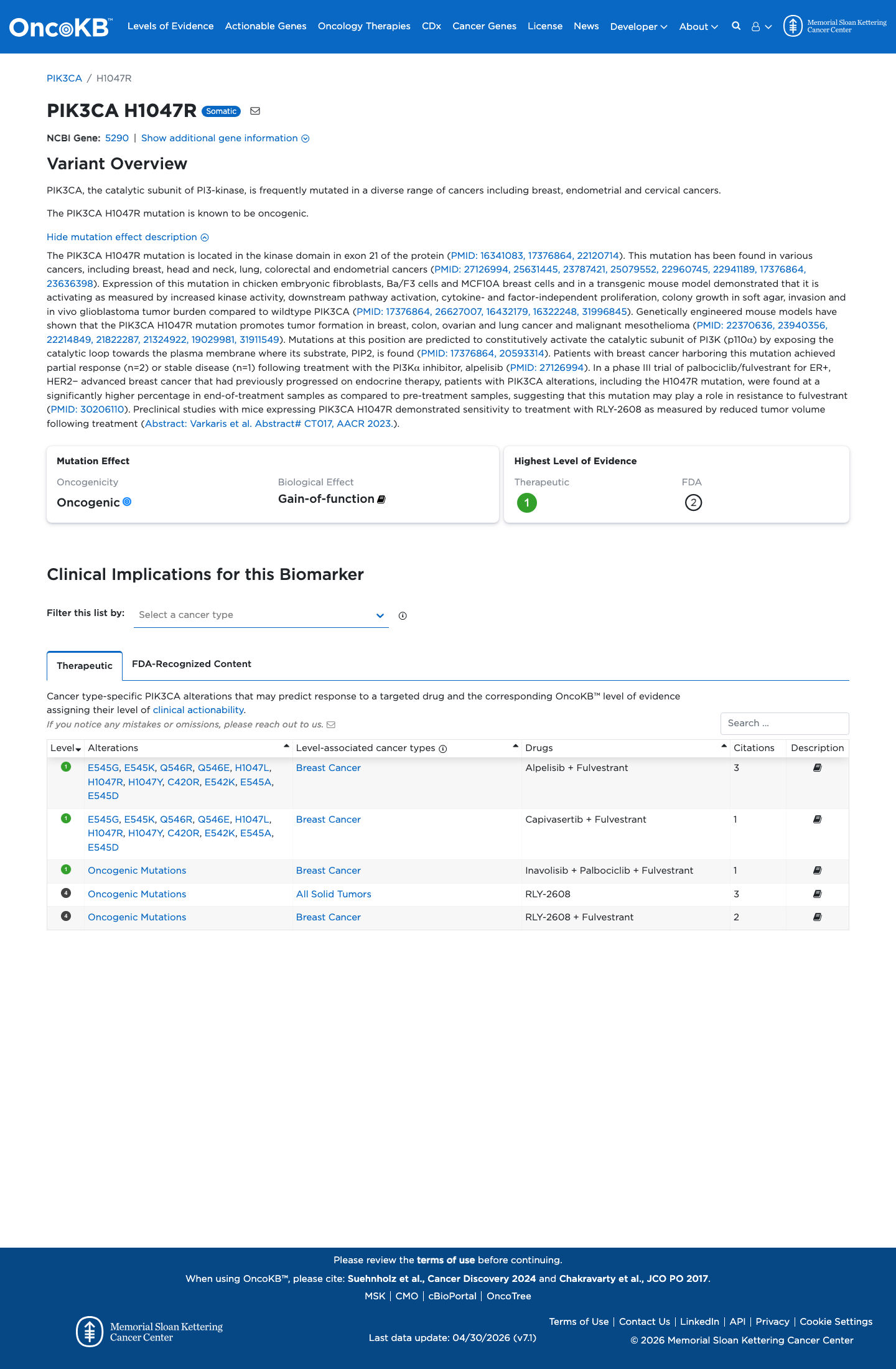
The PIK3CA c.3140A>G (p.His1047Arg) variant has been observed in somatic cancers and is reported in ClinVar, including a Pathogenic expert-panel classification by the ClinGen Brain Malformations Variant Curation Expert Panel.
clinvar ↗ PMID:16322248 ↗ PMID:16432179 ↗This variant is absent from gnomAD v4.1 and is present once in gnomAD v2.1 (1/248274 alleles; AF 0.00040%), which is below the VCEP PM2 range and well below the BS1 (>0.0185%) and BA1 (>0.0926%) population thresholds.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗Published functional studies showed increased PI3K activity, growth factor-independent and anchorage-independent proliferation, Akt-pathway activation, and tumor formation relative to wild type, consistent with a gain-of-function effect.
PMID:16322248 ↗ PMID:16432179 ↗ PMID:17376864 ↗ PMID:20593314 ↗ PMID:22370636 ↗SpliceAI predicts no significant splice impact (max delta score 0.04), and REVEL and BayesDel scores are 0.455 and 0.247638, respectively; however, this VCEP does not use PP3 or BP4 for PIK3CA gain-of-function missense variants.
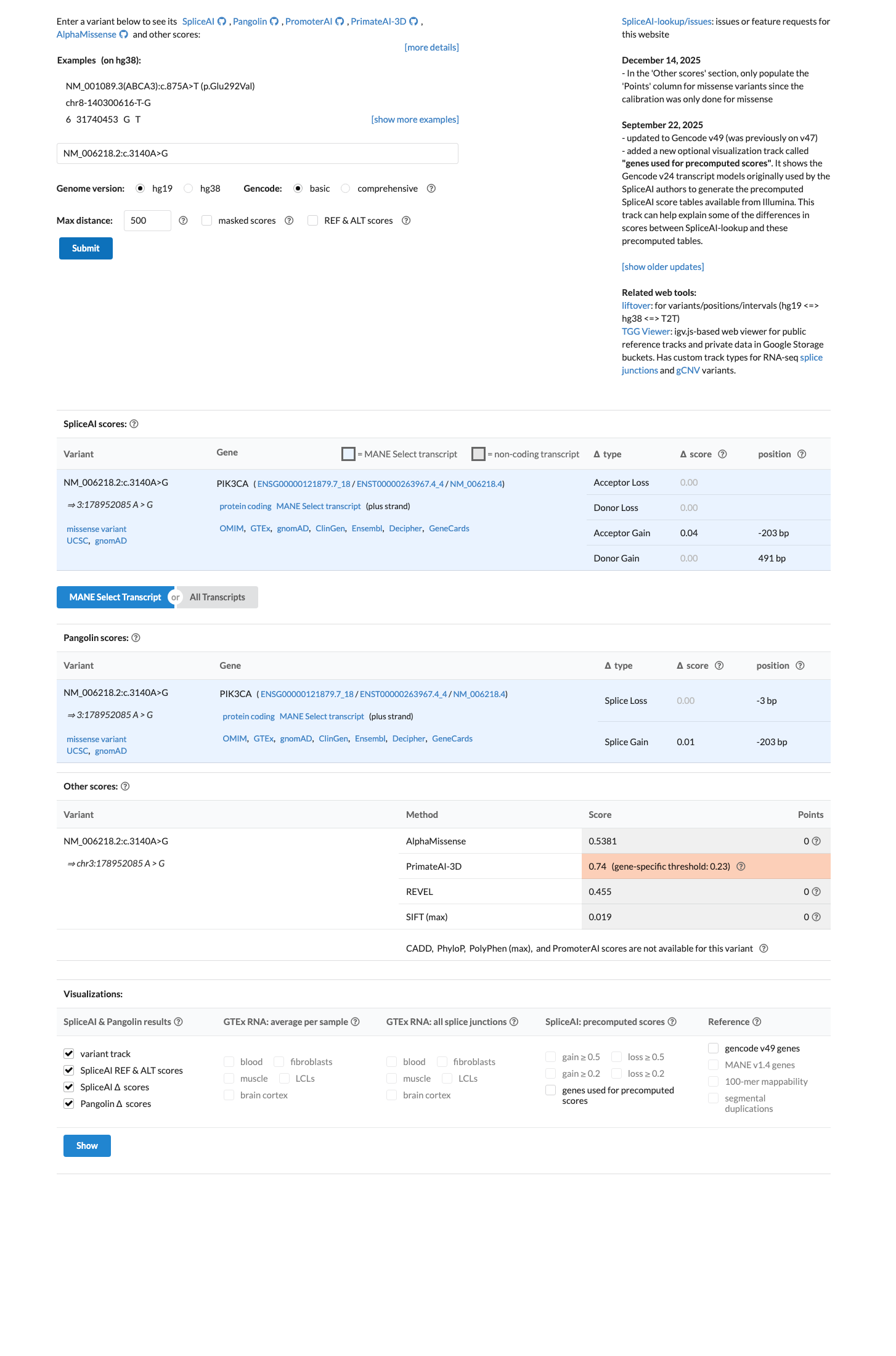
spliceai ↗ cspec ↗