Classification rationale
1
The SPG11 c.6062G>A (p.Arg2021Gln) variant has been reported in ClinVar with one likely benign submission and one uncertain significance submission.
clinvar ↗2
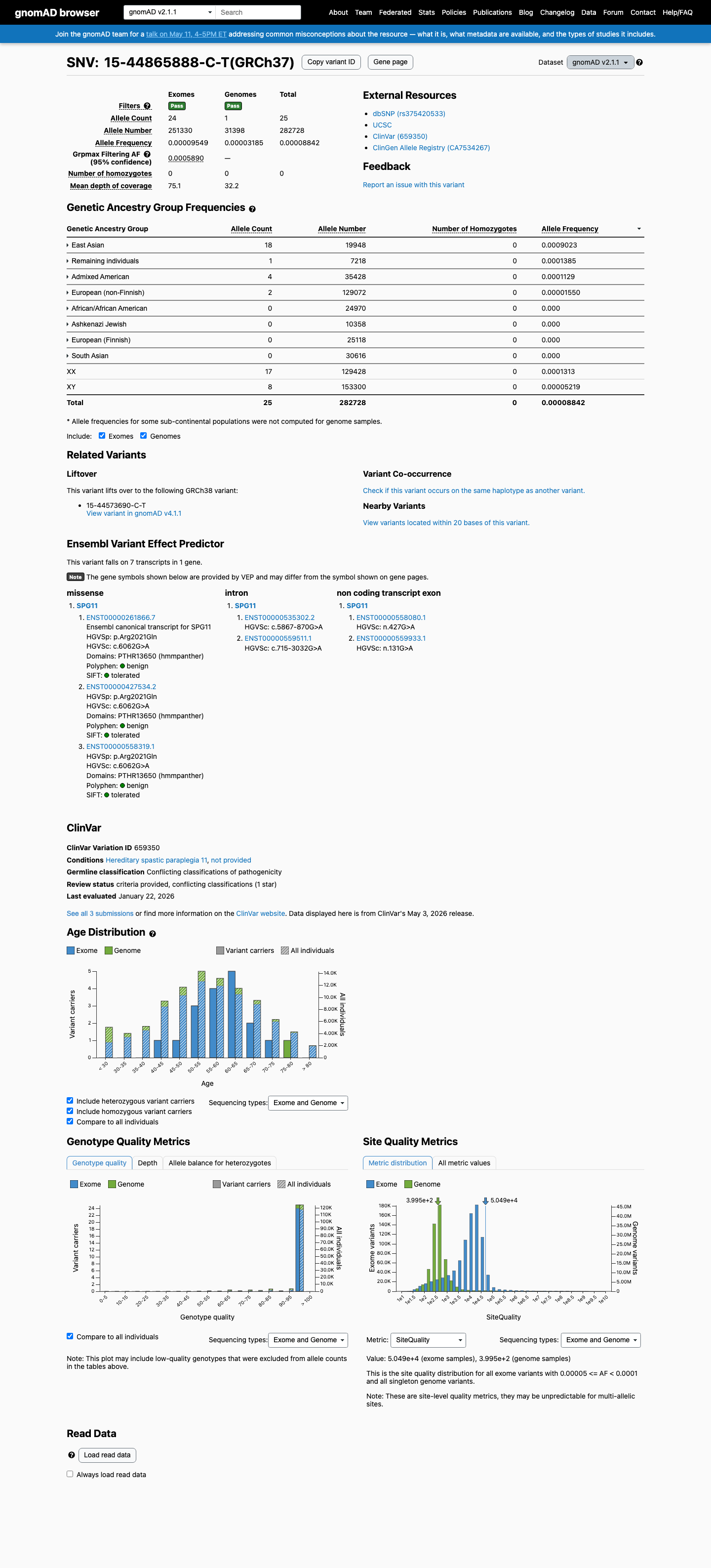
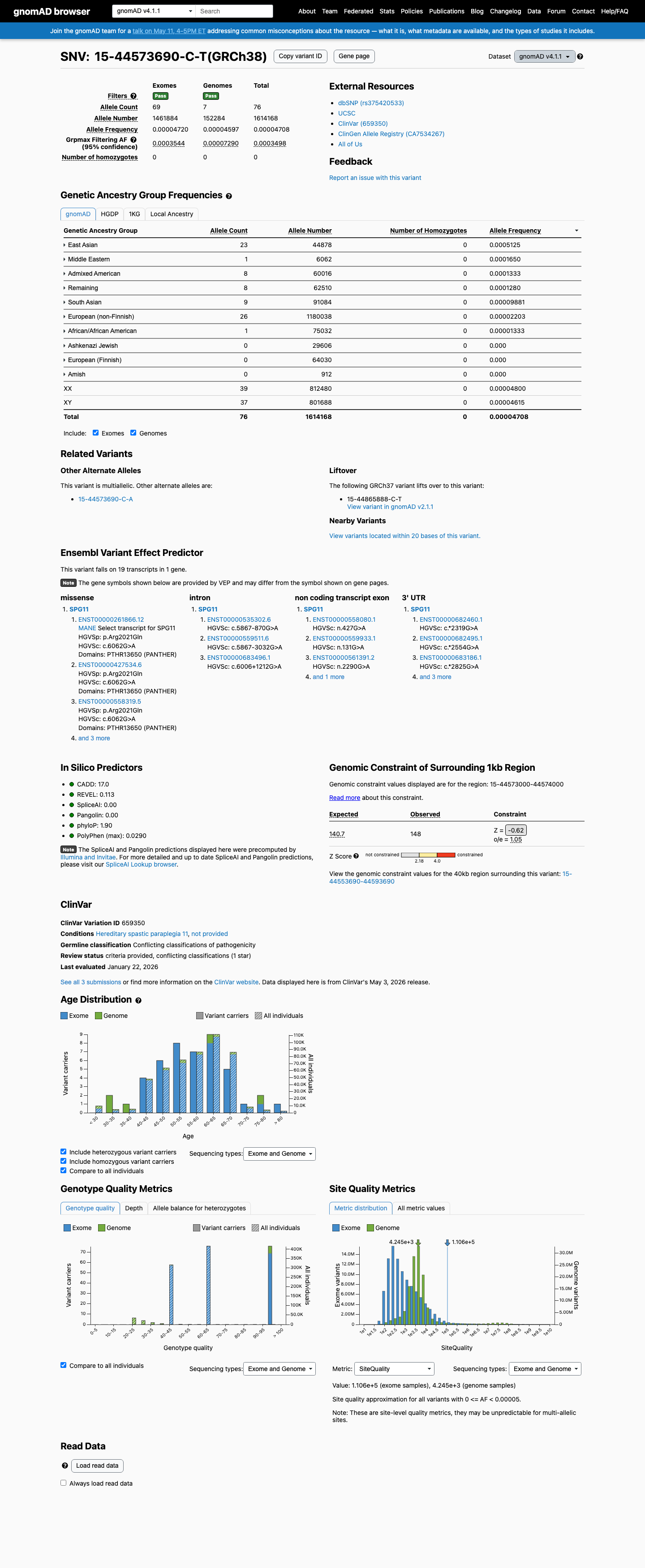
This variant is rare in population databases, with the highest observed frequency in East Asian individuals of 0.09023% in gnomAD v2.1 and 0.05125% in gnomAD v4.1, both below the 0.1% PM2 threshold used for this generic non-VCEP review.
gnomad_v2 ↗ gnomad_v4 ↗3
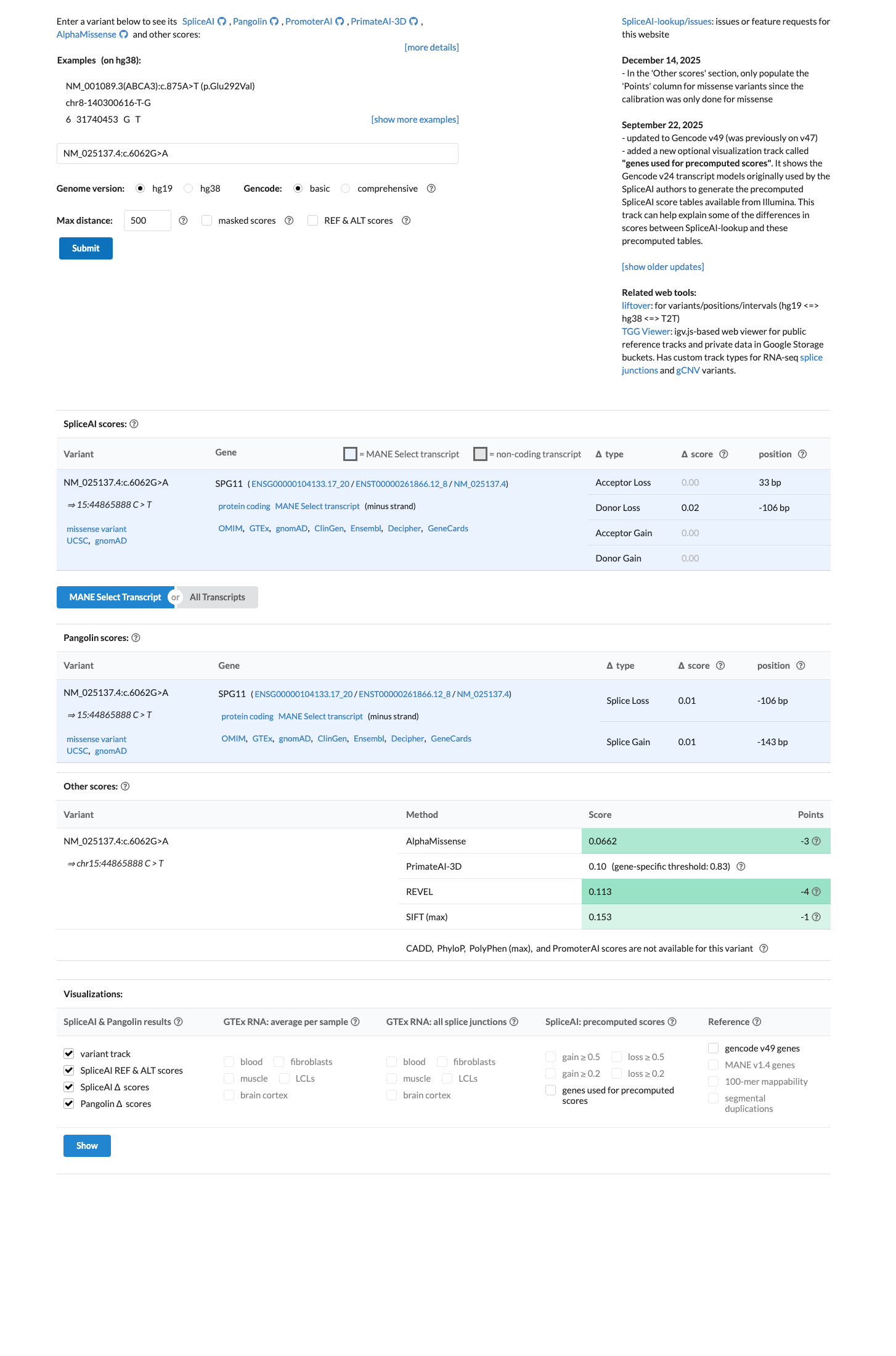
Computational data argue against a damaging effect, with REVEL 0.113, BayesDel -0.445237, and SpliceAI showing no significant predicted splice impact with a maximum delta score of 0.02.
spliceai ↗