Classification rationale
1
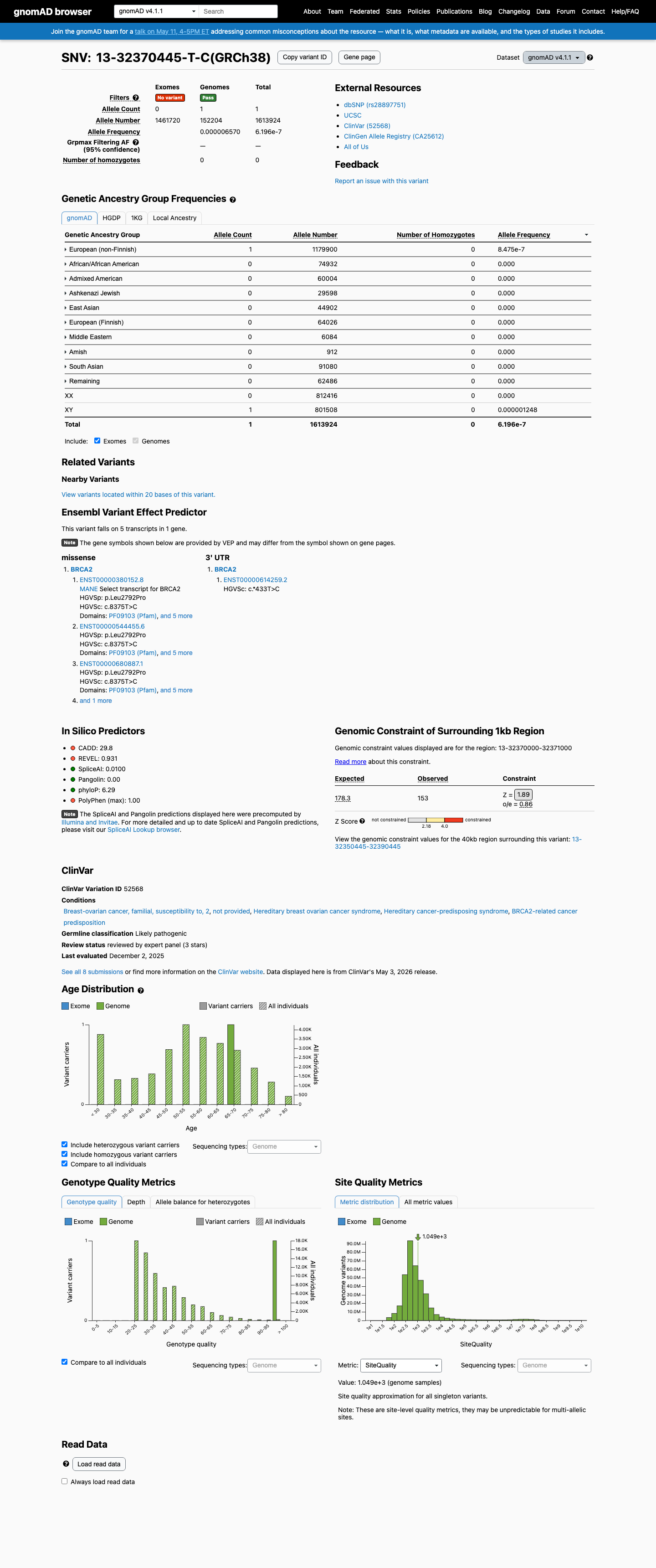
The BRCA2 c.8375T>C (p.Leu2792Pro) variant has been reported in ClinVar, including a Likely Pathogenic expert-panel classification from ClinGen ENIGMA.
clinvar ↗2
This variant is absent from gnomAD v2.1 and is present once in gnomAD v4.1 (1/1613924 alleles; AF 6.19608e-07), supporting rarity but not strict absence across available population datasets.
gnomad_v2 ↗ gnomad_v4 ↗3
In the ENIGMA BRCA2 curated functional dataset, two calibrated studies showed damaging protein function for this exact variant, and Table 9 assigns PS3 at strong strength.
PMID:29884841 ↗ cspec ↗4
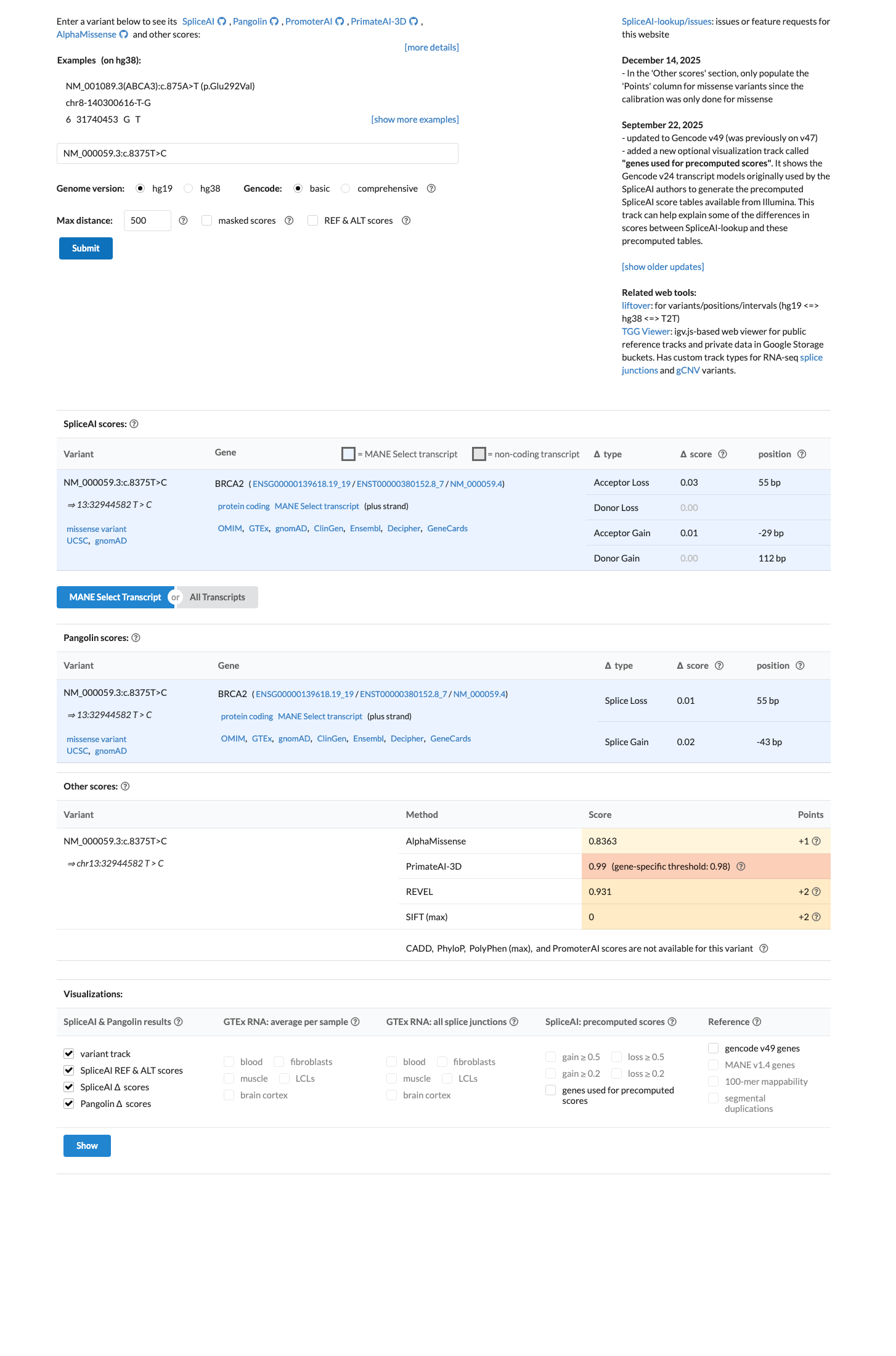
In silico data support a damaging protein effect within the BRCA2 DNA-binding domain, with BayesDel 0.54225 and REVEL 0.931, while SpliceAI predicts no significant splice impact (max delta 0.03).
spliceai ↗ cspec ↗