Classification rationale
1
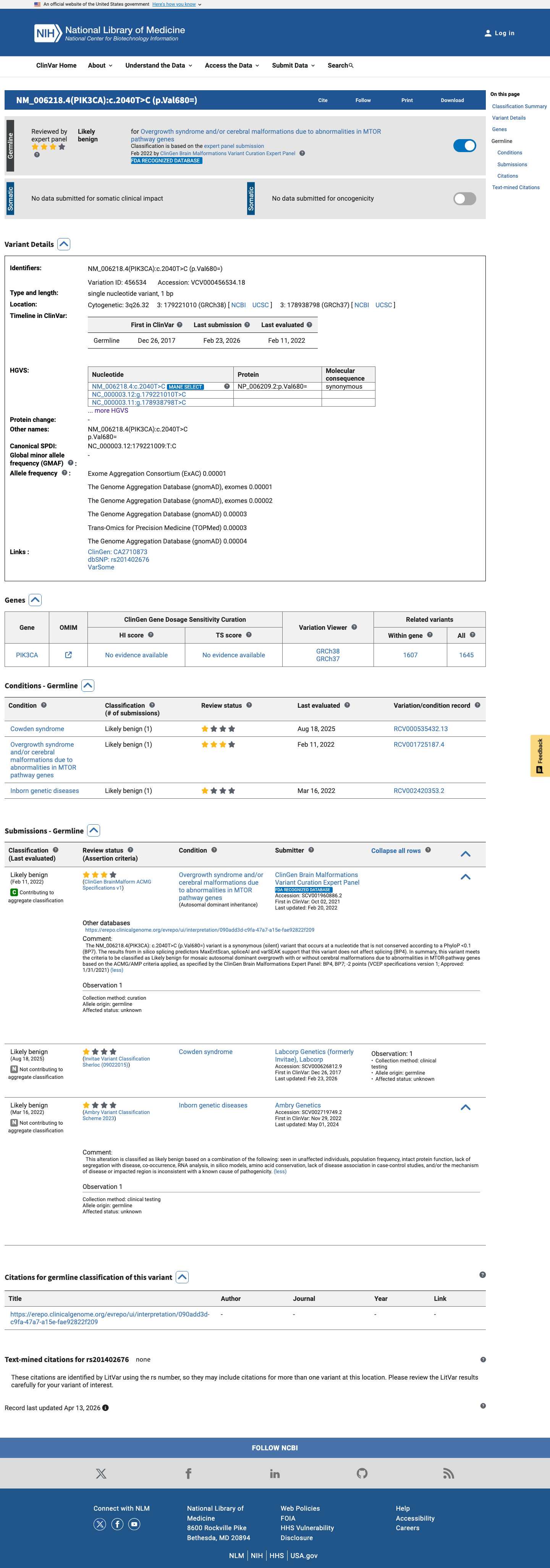
The PIK3CA c.2040T>C (p.Val680=) variant has been reported in ClinVar and is classified as likely benign by the ClinGen Brain Malformations Variant Curation Expert Panel.
clinvar ↗2
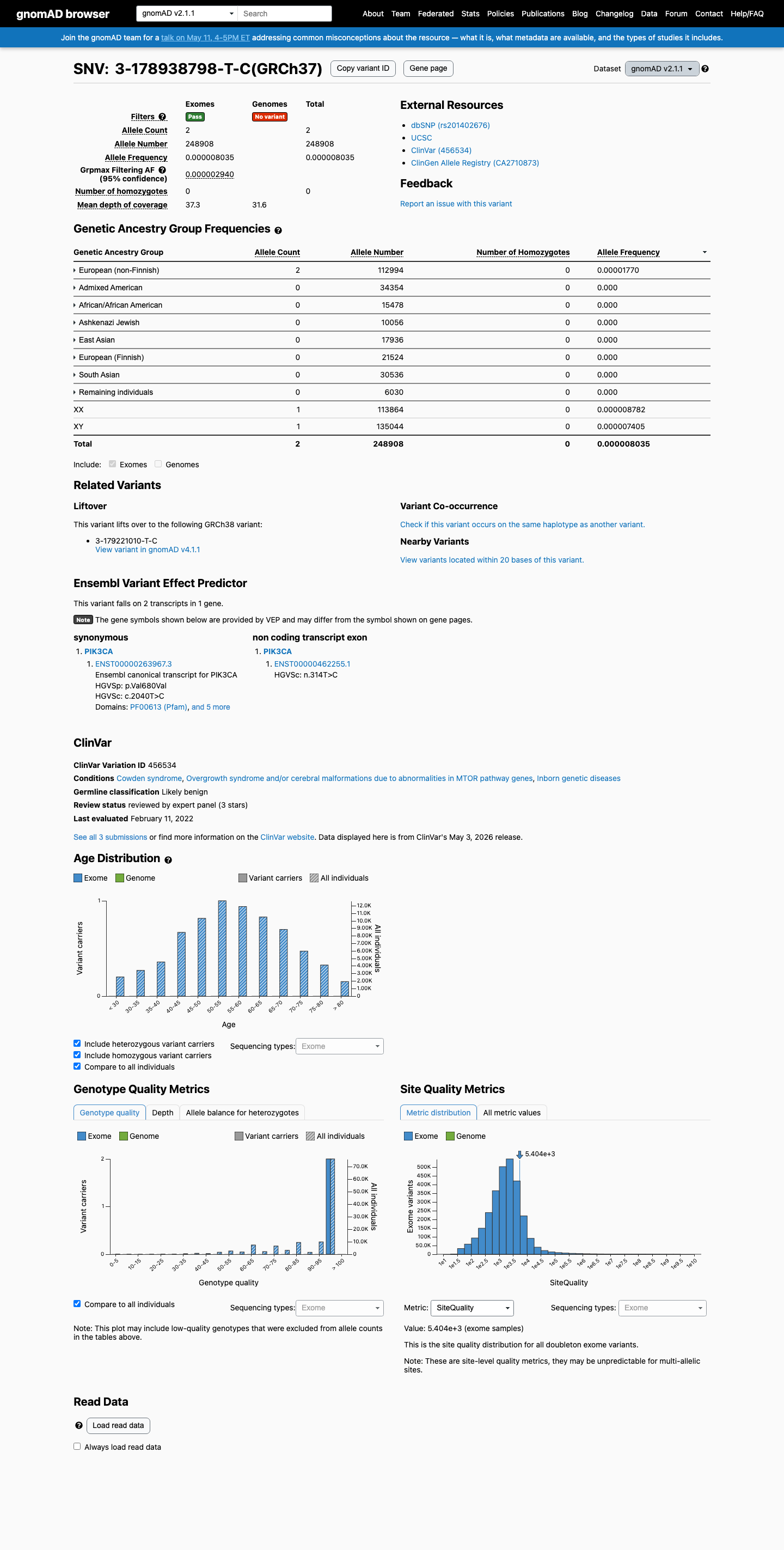
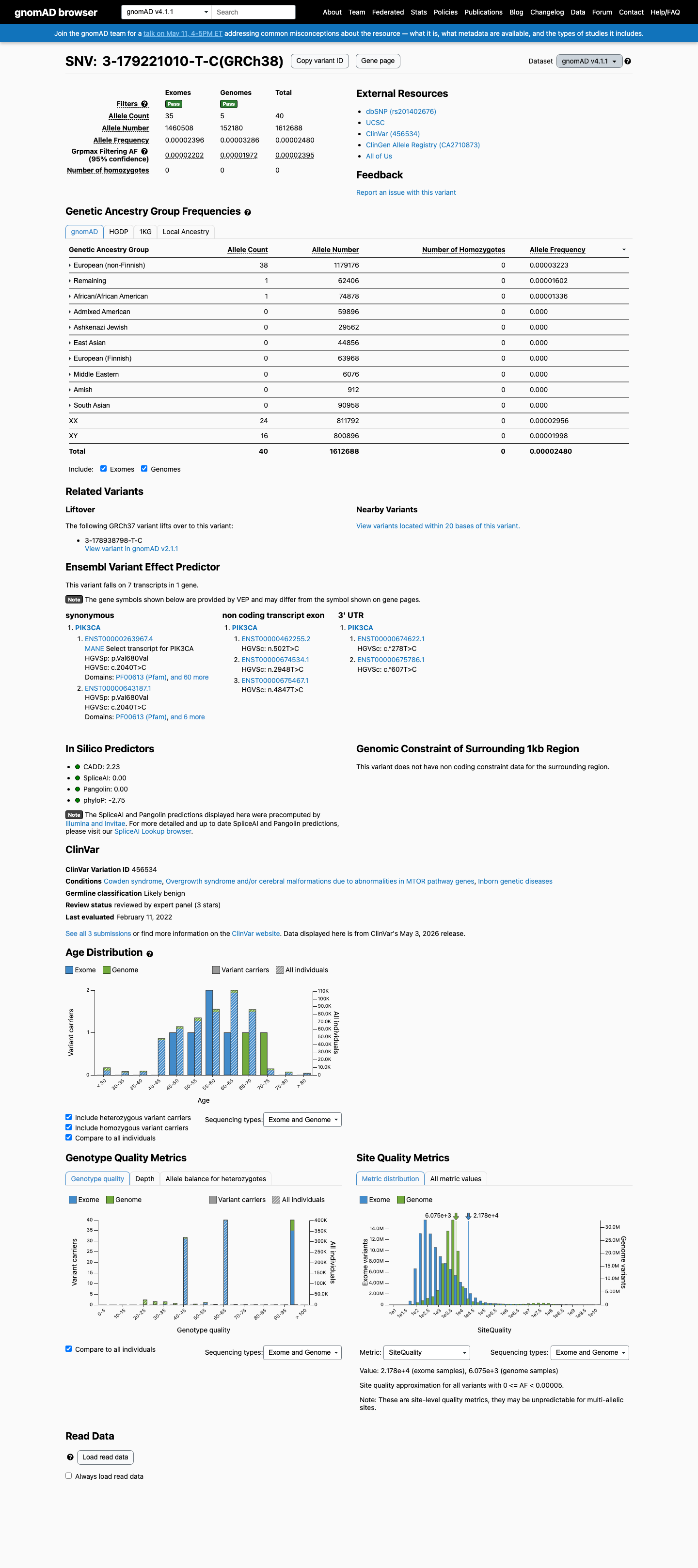
This variant is present in gnomAD at 0.00080% in v2.1 (2/248908 alleles) and 0.00248% in v4.1 (40/1612688 alleles), which is below the VCEP BS1 threshold of 0.0185% and BA1 threshold of 0.0926% but exceeds the VCEP PM2 one-person maximum.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
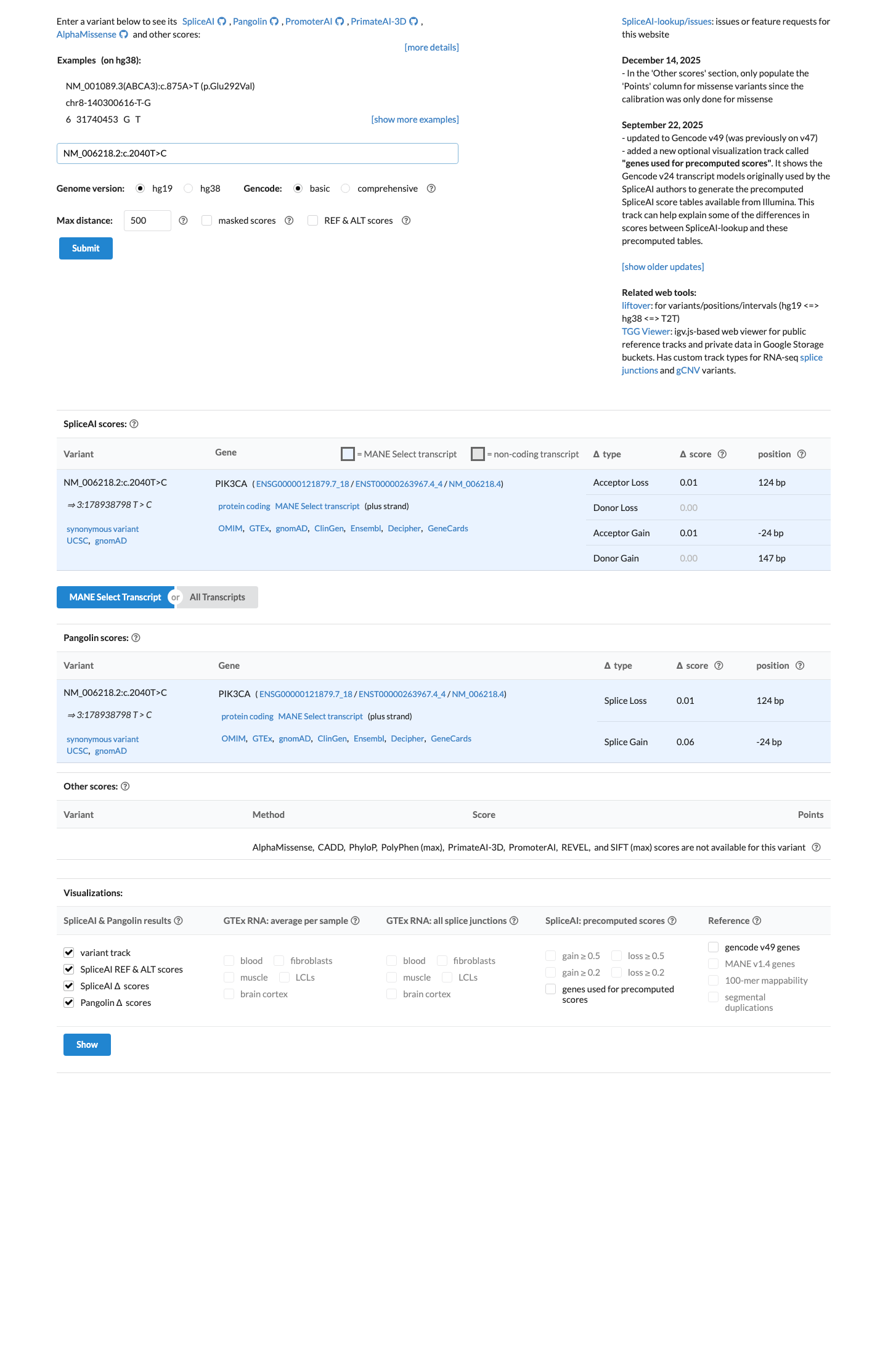
SpliceAI predicts no significant splice impact for this variant, with a maximum delta score of 0.01, although additional VCEP-specified splicing predictors and a conservation score were not identified for BP4 and BP7 assessment.
spliceai ↗ cspec ↗