Classification rationale
1
2
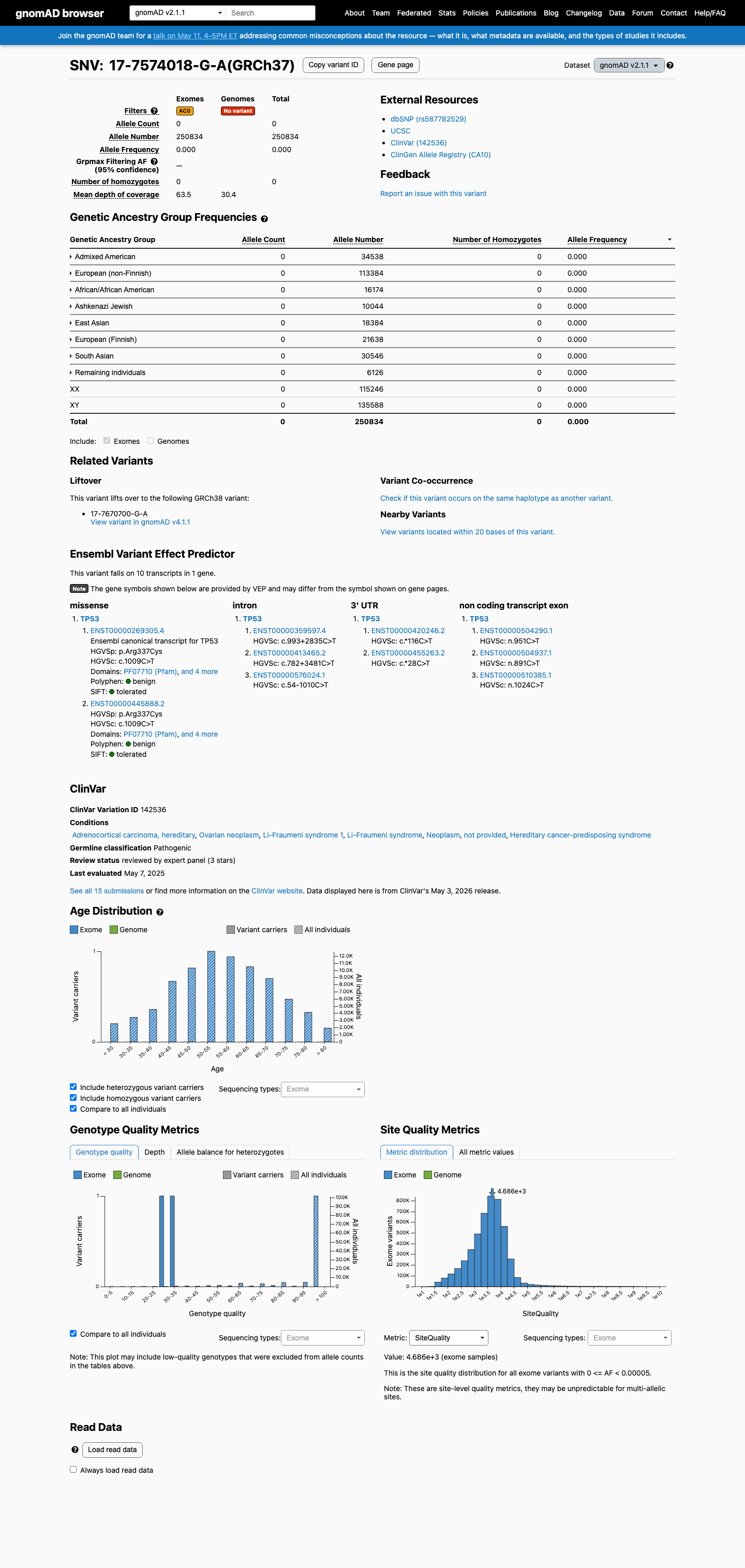
This variant is absent from gnomAD v2.1 and is not observed in gnomAD v4.1 (0/1613822 alleles), which supports rarity under the TP53 PM2_Supporting threshold.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
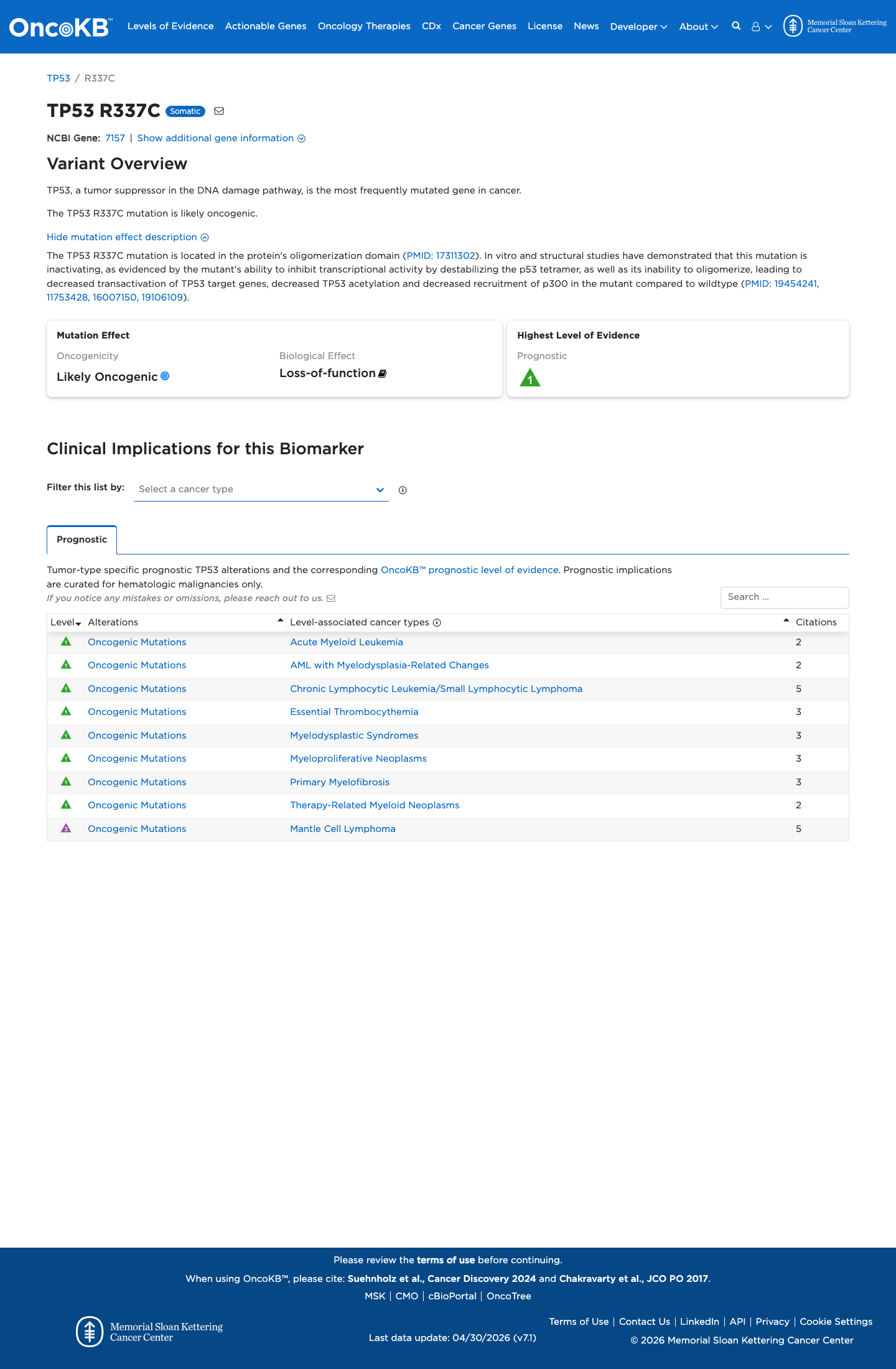
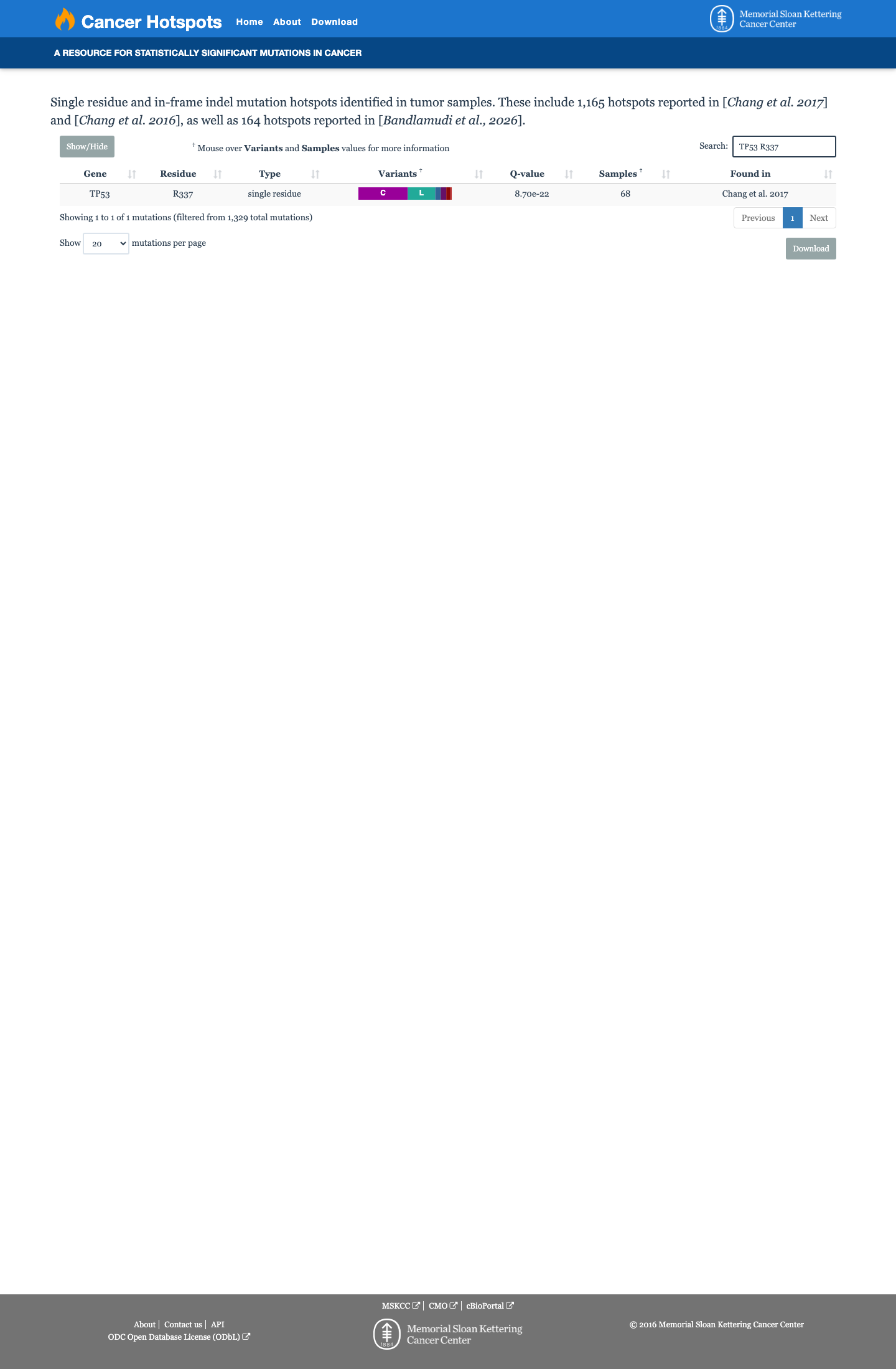
In the TP53 VCEP functional framework, published assay evidence compiled in the expert-panel worksheet shows p.Arg337Cys is non-functional or loss-of-function, with a PS3 assignment supporting a damaging effect on p53 activity.
PMID:16007150 ↗4
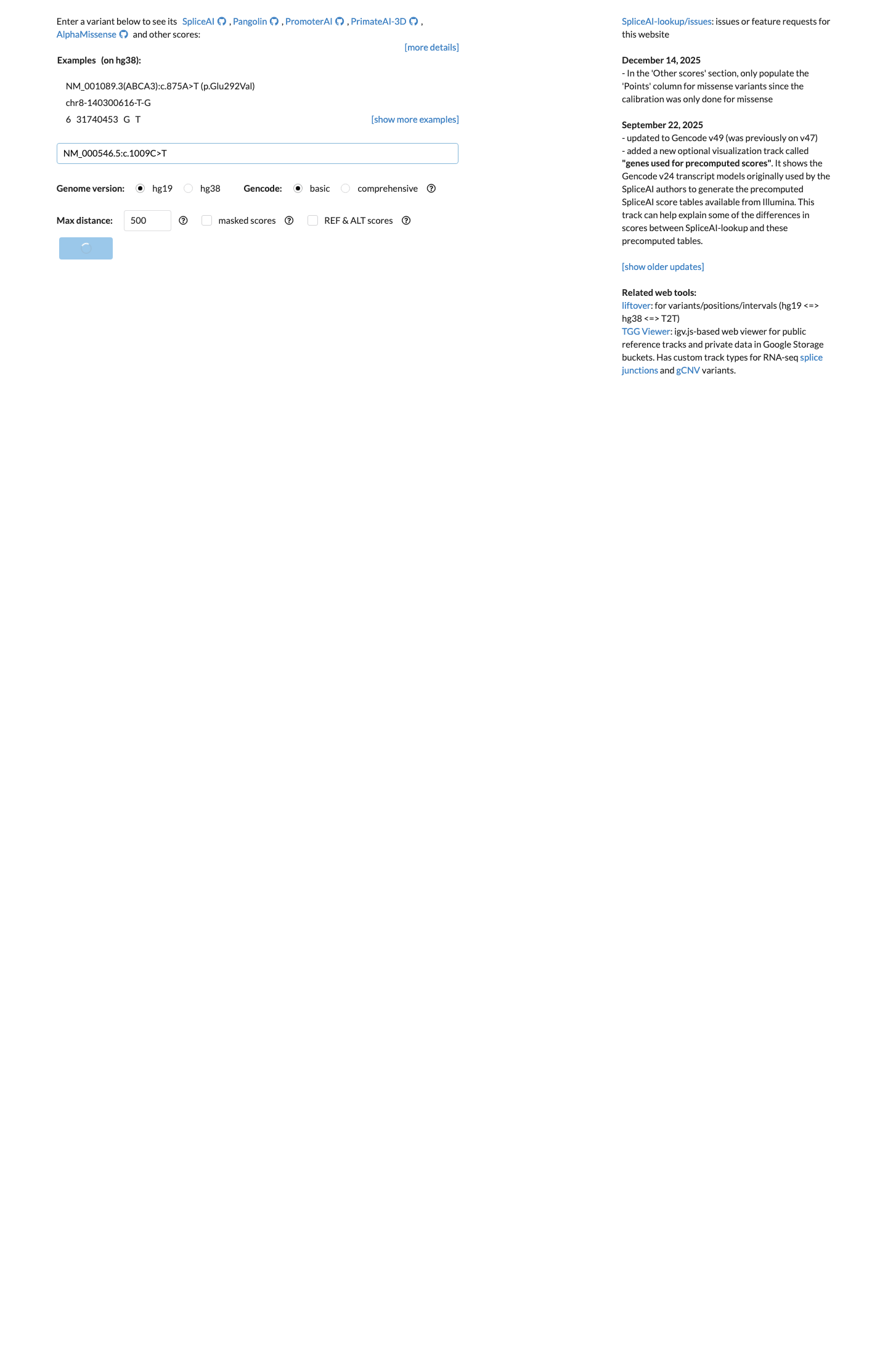
For in silico evaluation, the TP53 VCEP bioinformatic worksheet assigns PP3 to this missense change; BayesDel is 0.316468, REVEL is 0.715, and SpliceAI shows no predicted splice impact (max delta score 0.00).
spliceai ↗