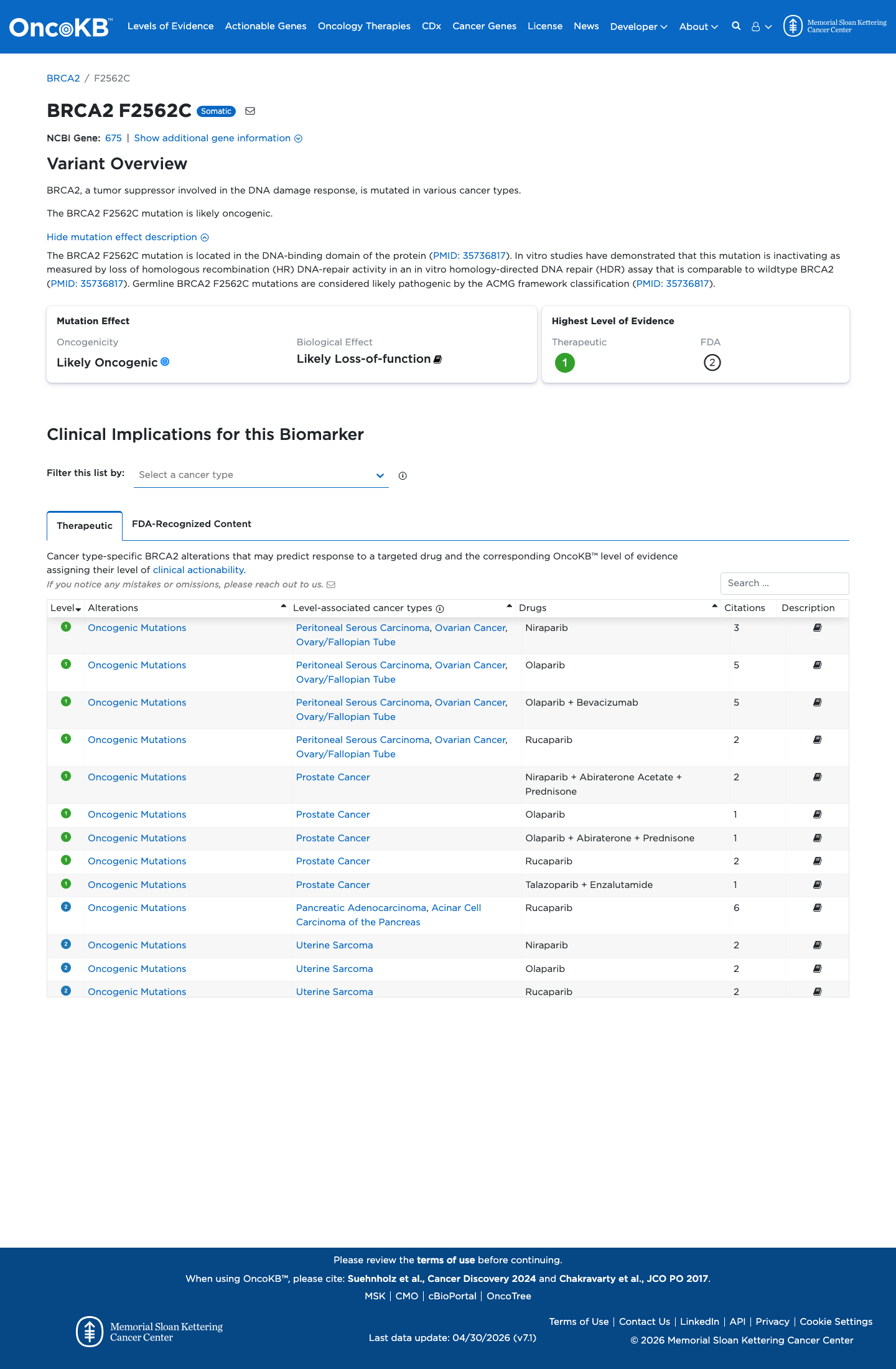
The BRCA2 c.7685T>G (p.Phe2562Cys; p.F2562C) variant has not been observed in COSMIC and has been reported in ClinVar, where the ClinGen ENIGMA BRCA1/2 expert panel classifies it as likely pathogenic.
clinvar ↗This variant is absent from gnomAD v2.1 and gnomAD v4.1, supporting rarity in population controls.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗In a calibrated BRCA2 functional study summarized by the ENIGMA functional assay table, this variant showed protein function similar to pathogenic control variants, supporting PS3_Strong.
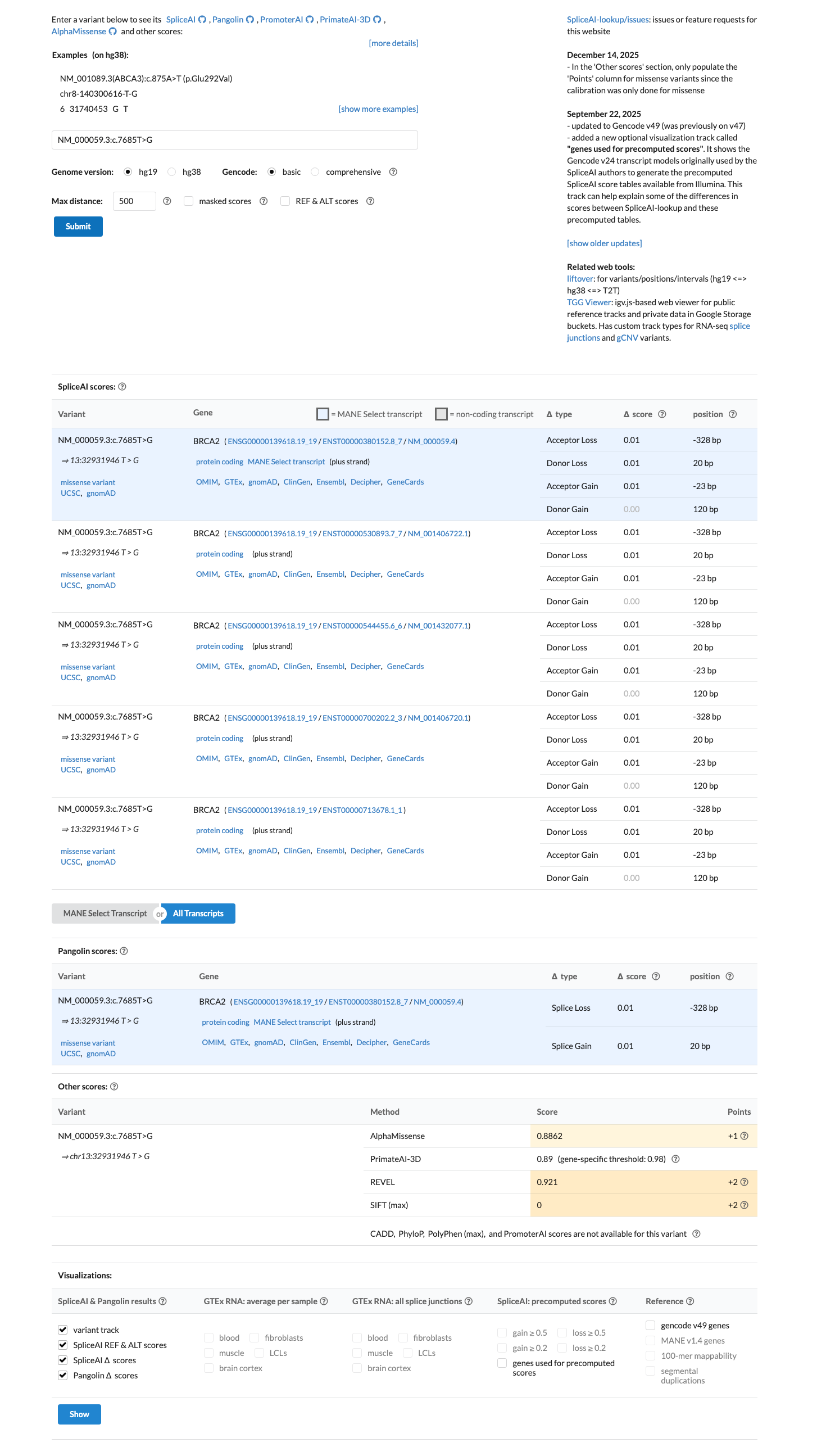
PMID:33609447 ↗ cspec ↗This missense change lies in the BRCA2 DNA-binding domain; BayesDel no-AF is 0.421539, which is above the ENIGMA PP3 threshold of 0.30, REVEL is 0.921, and SpliceAI predicts little splice effect with a maximum delta score of 0.01.
spliceai ↗ cspec ↗In the BRCA2 clinical-history likelihood-ratio dataset, this variant has an LR of 0.9916 in 1 proband, which is within the neutral zone and does not support PP4 or BP5.
PMID:31853058 ↗ cspec ↗