Classification rationale
1
2
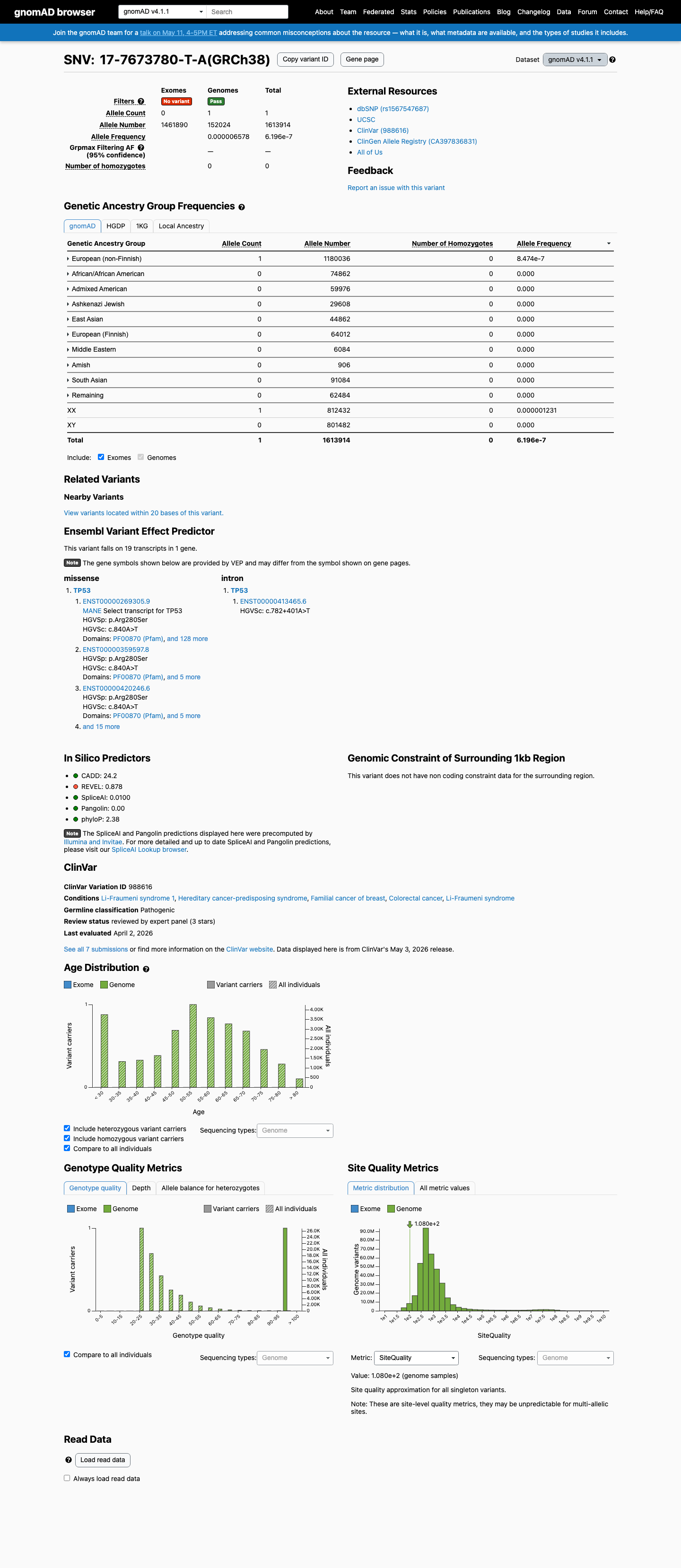
This variant is absent from gnomAD v2.1 and is present only once in gnomAD v4.1 (1/1,613,914 alleles; AF 6.19612e-07), which is below the TP53 PM2 threshold of 0.00003.
gnomad_v2 ↗ gnomad_v4 ↗ cspec ↗3
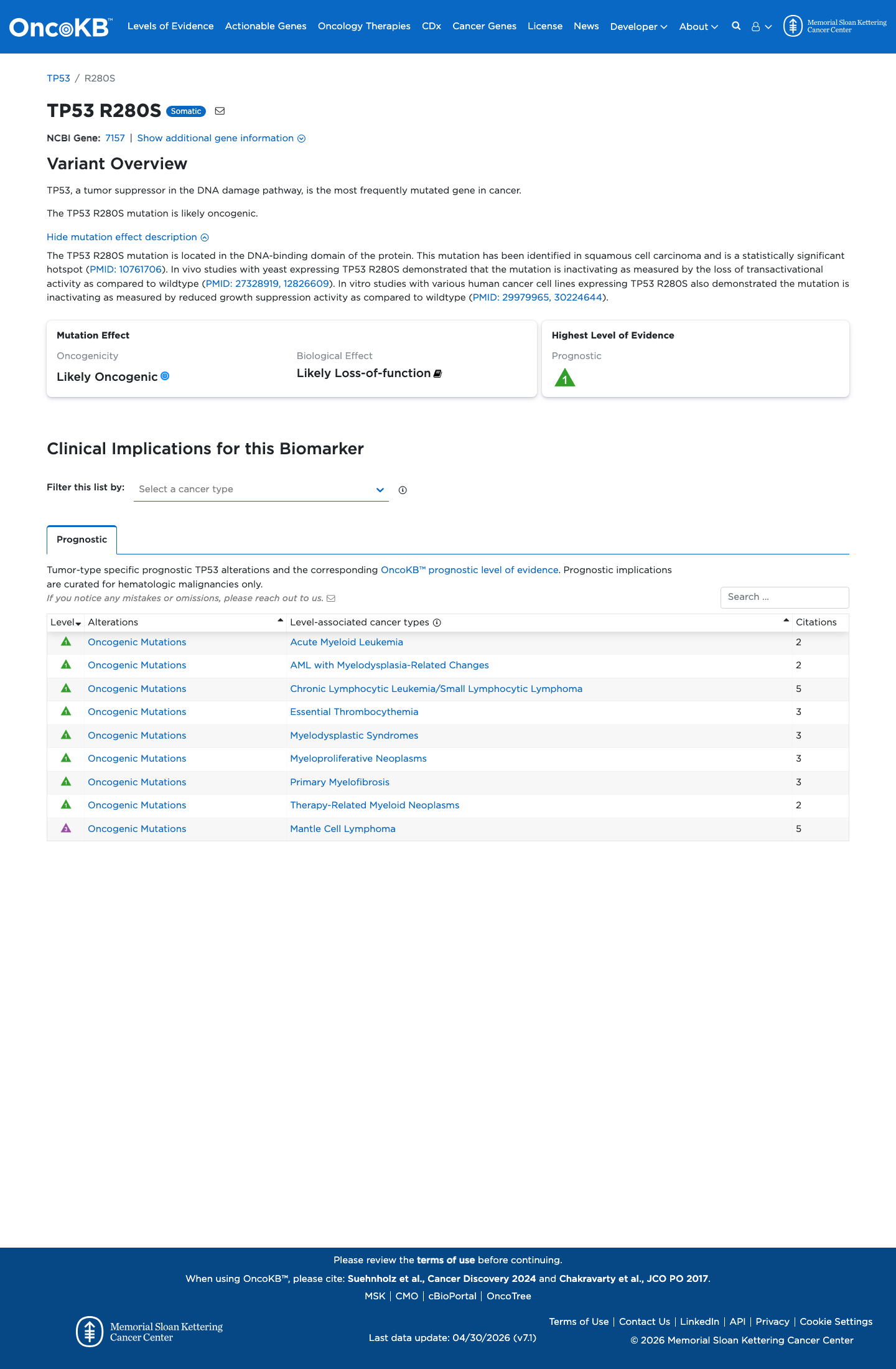
In the TP53 VCEP functional worksheet, p.Arg280Ser is non-functional in the Kato assay and shows loss of function across additional eligible assays, supporting PS3.
PMID:12826609 ↗ PMID:29979965 ↗ PMID:30224644 ↗4
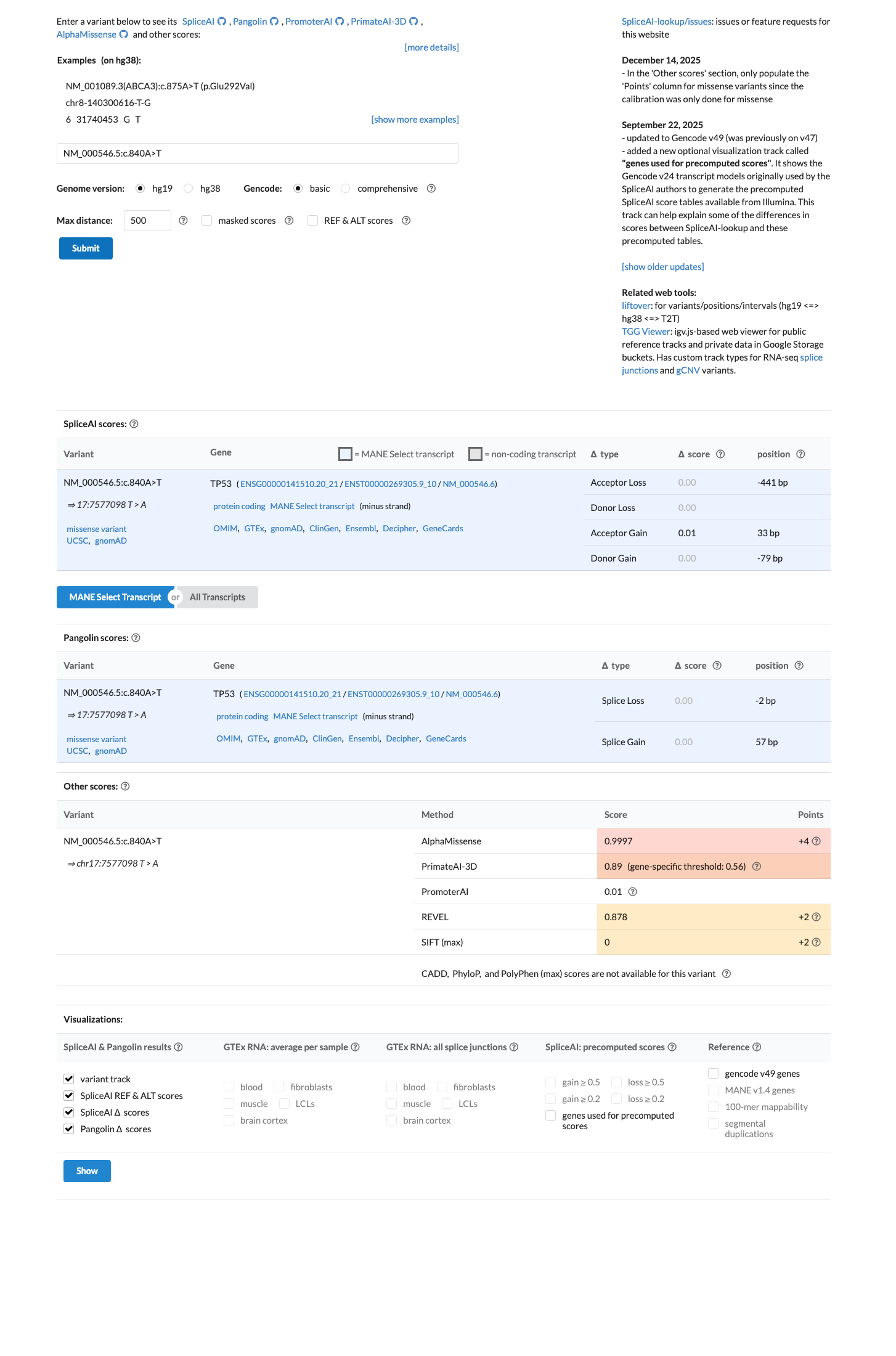
TP53 VCEP in silico resources assign PP3_Moderate for c.840A>T based on aGVGD Class C65 and BayesDel 0.519515; local computational data are concordant with BayesDel 0.471463 and REVEL 0.878, while SpliceAI predicts no significant splice impact (max delta score 0.01).
spliceai ↗