Classification rationale
1
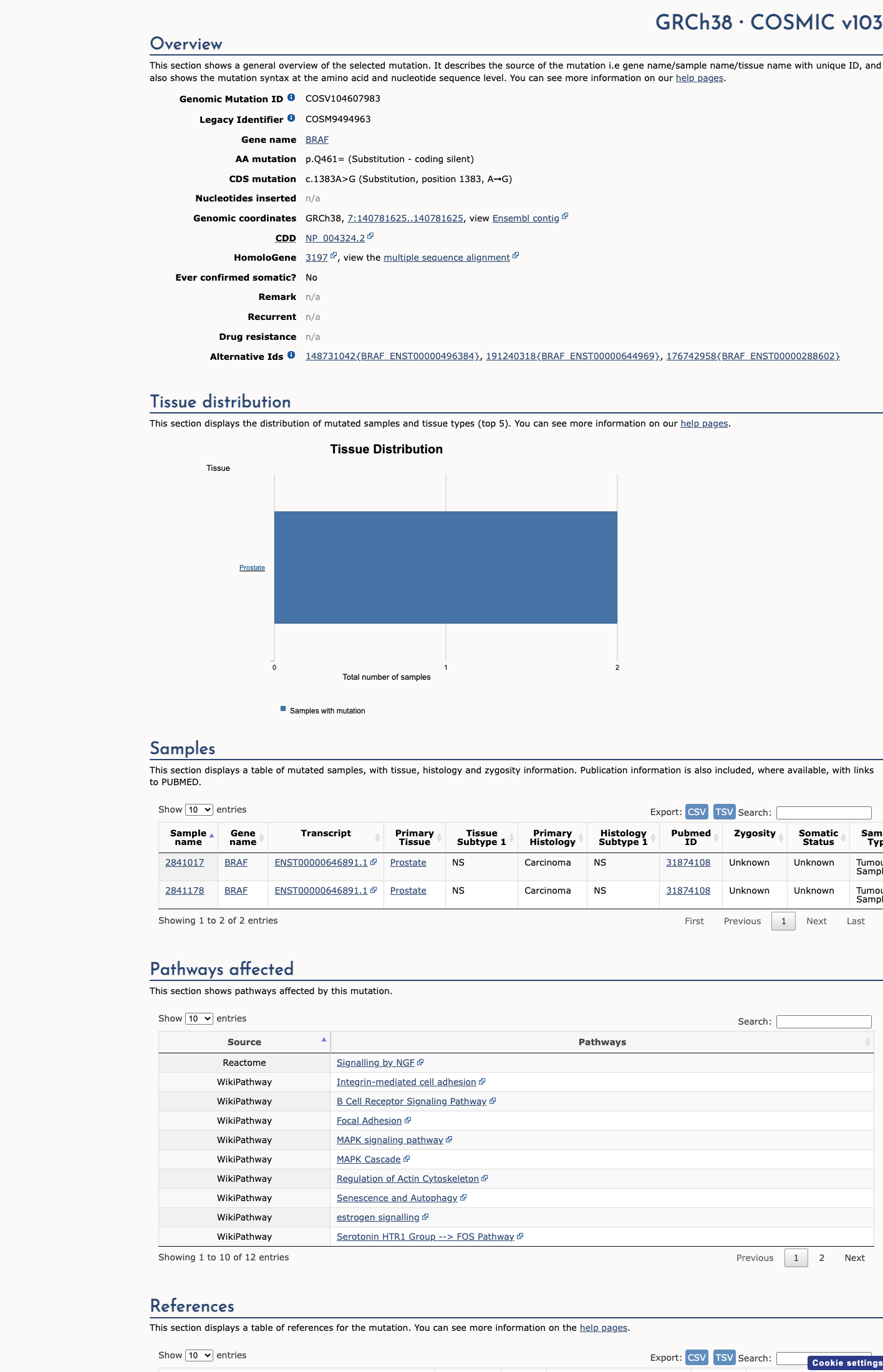
The BRAF c.1383A>G (p.Gln461=, p.Q461=) variant has been reported in ClinVar as benign, including a benign expert-panel assertion from the ClinGen RASopathy Variant Curation Expert Panel, and it has not been established as a statistically significant hotspot at codon 461.
clinvar ↗ hotspots ↗2
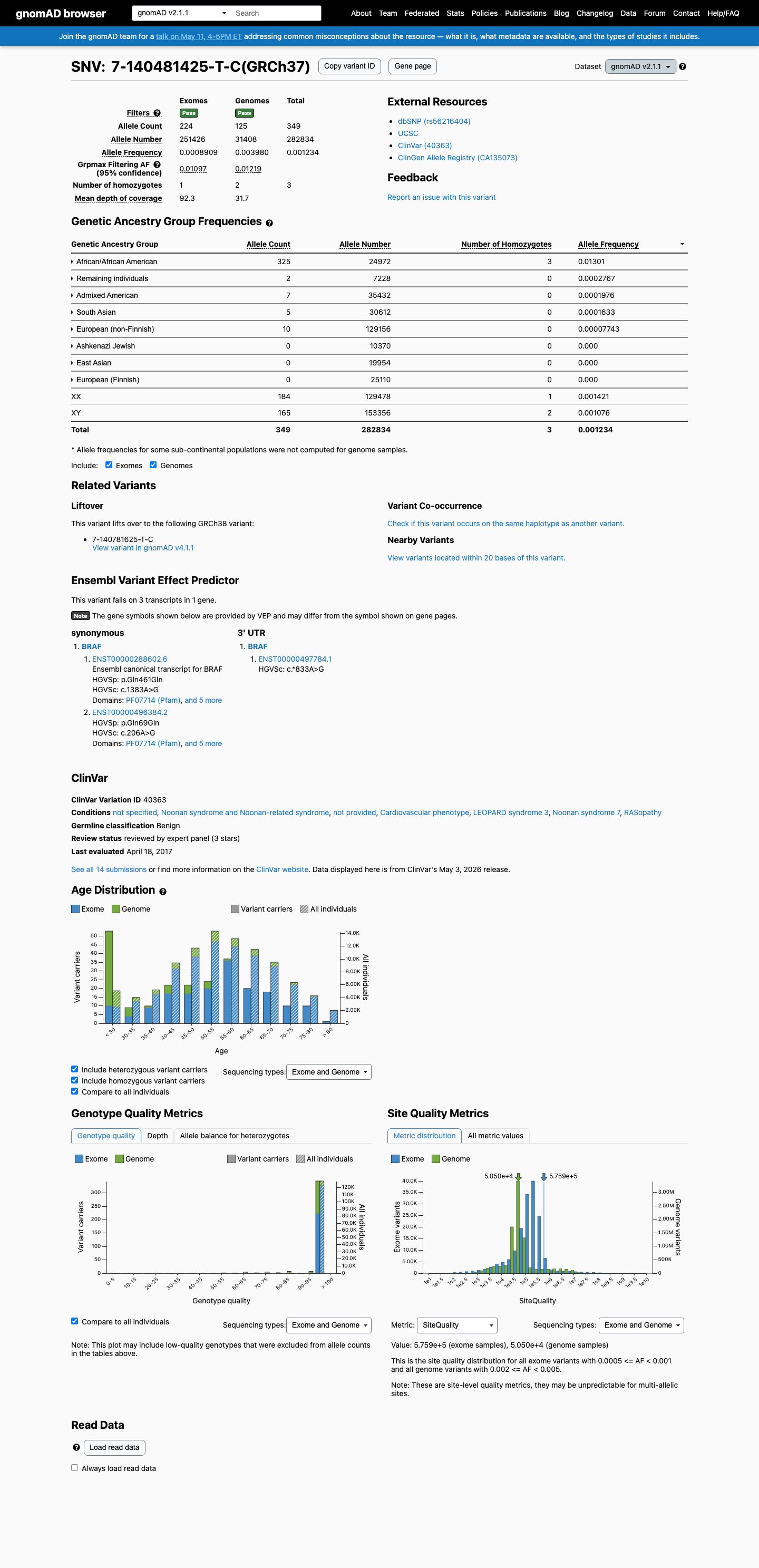
This variant is present in gnomAD v2.1 at 0.12339% overall (349/282834 alleles; 3 homozygotes), with an African/African American frequency of 1.30146% and grpmax filtering allele frequency of 1.2192%, which is above the BRAF RASopathy BA1 threshold of 0.05%.
gnomad_v2 ↗ cspec ↗3
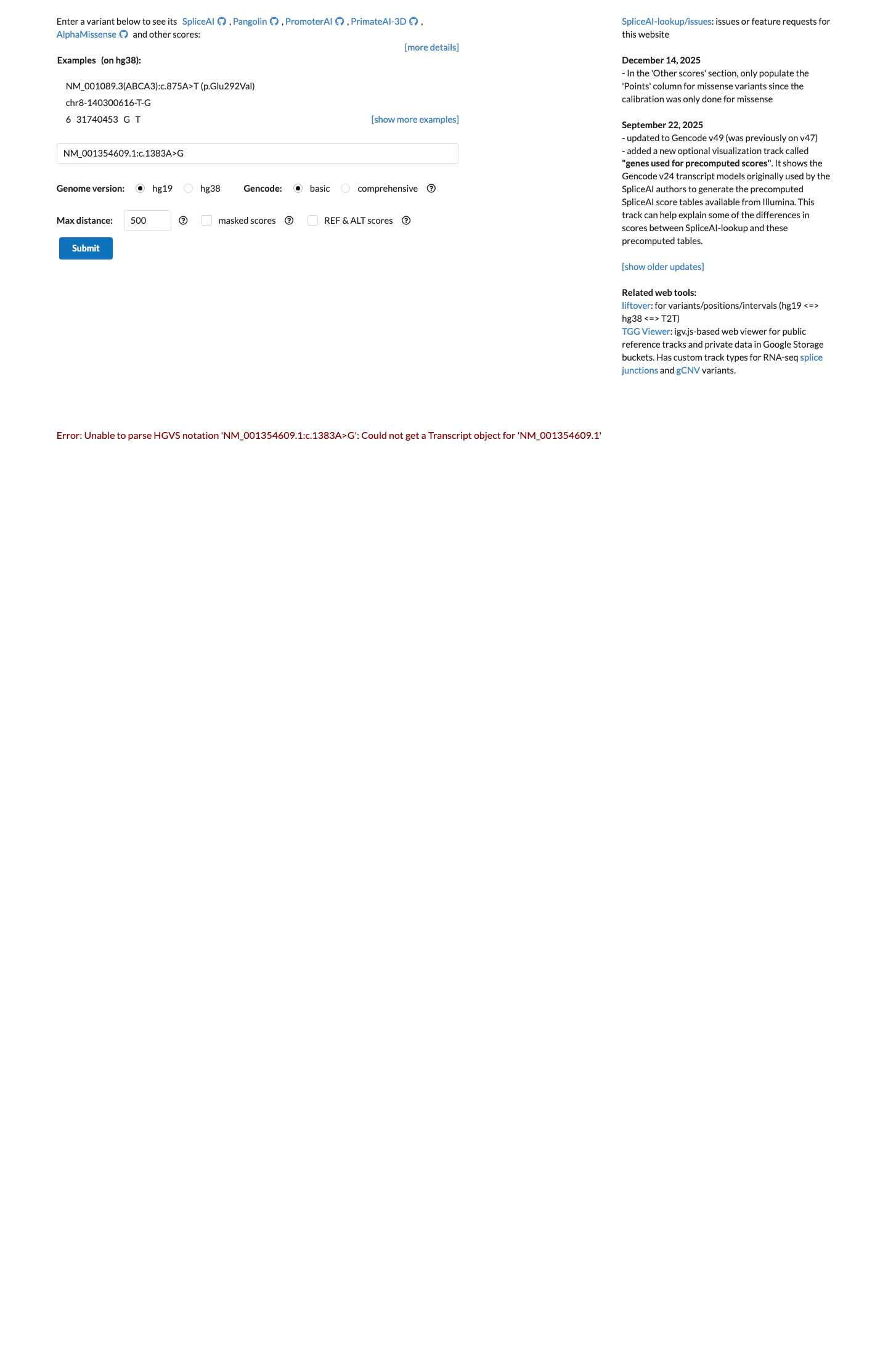
Computational data do not suggest a harmful effect: SpliceAI predicts no significant splice impact with a maximum delta score of 0.01, and the available REVEL score is 0.084.
spliceai ↗